[English] 日本語
 Yorodumi
Yorodumi- EMDB-3643: Complex of CMG helicase with polymerase epsilon lacking the catal... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3643 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Complex of CMG helicase with polymerase epsilon lacking the catalytic domain of Pol2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 29.6 Å | |||||||||
 Authors Authors | Zhou JC / Janska A / Goswami P / Renault L / Abid Ali F / Kotecha A / Diffley JFX / Costa A | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2017 Journal: Proc Natl Acad Sci U S A / Year: 2017Title: CMG-Pol epsilon dynamics suggests a mechanism for the establishment of leading-strand synthesis in the eukaryotic replisome. Authors: Jin Chuan Zhou / Agnieszka Janska / Panchali Goswami / Ludovic Renault / Ferdos Abid Ali / Abhay Kotecha / John F X Diffley / Alessandro Costa /  Abstract: The replisome unwinds and synthesizes DNA for genome duplication. In eukaryotes, the Cdc45-MCM-GINS (CMG) helicase and the leading-strand polymerase, Pol epsilon, form a stable assembly. The ...The replisome unwinds and synthesizes DNA for genome duplication. In eukaryotes, the Cdc45-MCM-GINS (CMG) helicase and the leading-strand polymerase, Pol epsilon, form a stable assembly. The mechanism for coupling DNA unwinding with synthesis is starting to be elucidated, however the architecture and dynamics of the replication fork remain only partially understood, preventing a molecular understanding of chromosome replication. To address this issue, we conducted a systematic single-particle EM study on multiple permutations of the reconstituted CMG-Pol epsilon assembly. Pol epsilon contains two flexibly tethered lobes. The noncatalytic lobe is anchored to the motor of the helicase, whereas the polymerization domain extends toward the side of the helicase. We observe two alternate configurations of the DNA synthesis domain in the CMG-bound Pol epsilon. We propose that this conformational switch might control DNA template engagement and release, modulating replisome progression. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3643.map.gz emd_3643.map.gz | 446.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3643-v30.xml emd-3643-v30.xml emd-3643.xml emd-3643.xml | 7.9 KB 7.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3643.png emd_3643.png | 153.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3643 http://ftp.pdbj.org/pub/emdb/structures/EMD-3643 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3643 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3643 | HTTPS FTP |
-Validation report
| Summary document |  emd_3643_validation.pdf.gz emd_3643_validation.pdf.gz | 189.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3643_full_validation.pdf.gz emd_3643_full_validation.pdf.gz | 188.3 KB | Display | |
| Data in XML |  emd_3643_validation.xml.gz emd_3643_validation.xml.gz | 5.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3643 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3643 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3643 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3643 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3643.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3643.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.45 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
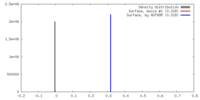
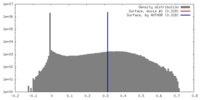
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Complex of CMG helicase with polymerase epsilon where the catalyt...
| Entire | Name: Complex of CMG helicase with polymerase epsilon where the catalytic domain has been deleted |
|---|---|
| Components |
|
-Supramolecule #1: Complex of CMG helicase with polymerase epsilon where the catalyt...
| Supramolecule | Name: Complex of CMG helicase with polymerase epsilon where the catalytic domain has been deleted type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Staining | Type: NEGATIVE / Material: uranyl formate |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 1000 (2k x 2k) / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: OTHER |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 29.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 15526 |
|---|---|
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera

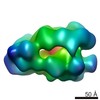








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)