[English] 日本語
 Yorodumi
Yorodumi- EMDB-34592: Cryo-EM structure of the p300 catalytic core bound to the H4K12ac... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 2 (4.8 angstrom resolution) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Acetyl taransferase / Complex / Nucleosome / TRANSFERASE / TRANSFERASE-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationbehavioral defense response / negative regulation of protein oligomerization / peptidyl-lysine propionylation / histone lactyltransferase (CoA-dependent) activity / peptidyl-lysine crotonylation / peptidyl-lysine butyrylation / histone butyryltransferase activity / histone isonicotinyltransferase activity / histone H1K75 acetyltransferase activity / swimming ...behavioral defense response / negative regulation of protein oligomerization / peptidyl-lysine propionylation / histone lactyltransferase (CoA-dependent) activity / peptidyl-lysine crotonylation / peptidyl-lysine butyrylation / histone butyryltransferase activity / histone isonicotinyltransferase activity / histone H1K75 acetyltransferase activity / swimming / histone H3K122 acetyltransferase activity / internal protein amino acid acetylation / peptide butyryltransferase activity / regulation of tubulin deacetylation / histone H2B acetyltransferase activity / peptide 2-hydroxyisobutyryltransferase activity / histone crotonyltransferase activity / NOTCH2 intracellular domain regulates transcription / protein propionyltransferase activity / thigmotaxis / L-lysine N6-acetyltransferase activity, acting on acetyl phosphate as donor / positive regulation of TORC2 signaling / internal peptidyl-lysine acetylation / histone H4 acetyltransferase activity / negative regulation of chromosome condensation / histone H3 acetyltransferase activity / NFE2L2 regulating ER-stress associated genes / cellular response to L-leucine / acetylation-dependent protein binding / NFE2L2 regulating inflammation associated genes / Activation of the TFAP2 (AP-2) family of transcription factors / NGF-stimulated transcription / histone H3K18 acetyltransferase activity / N-terminal peptidyl-lysine acetylation / histone H3K27 acetyltransferase activity / LRR FLII-interacting protein 1 (LRRFIP1) activates type I IFN production / NFE2L2 regulates pentose phosphate pathway genes / NFE2L2 regulating MDR associated enzymes / STAT3 nuclear events downstream of ALK signaling / negative regulation of brown fat cell differentiation / Polo-like kinase mediated events / regulation of mitochondrion organization / regulation of androgen receptor signaling pathway / host-mediated activation of viral transcription / Regulation of FOXO transcriptional activity by acetylation / TGFBR3 expression / Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / RUNX3 regulates NOTCH signaling / NOTCH4 Intracellular Domain Regulates Transcription / Regulation of NFE2L2 gene expression / Regulation of gene expression by Hypoxia-inducible Factor / Nuclear events mediated by NFE2L2 / face morphogenesis / platelet formation / NOTCH3 Intracellular Domain Regulates Transcription / TRAF6 mediated IRF7 activation / regulation of glycolytic process / NFE2L2 regulating tumorigenic genes / NFE2L2 regulating anti-oxidant/detoxification enzymes / protein acetylation / megakaryocyte development / nuclear androgen receptor binding / acyltransferase activity / STAT family protein binding / Formation of paraxial mesoderm / acetyltransferase activity / histone acetyltransferase activity / FOXO-mediated transcription of cell death genes / stimulatory C-type lectin receptor signaling pathway / positive regulation of transforming growth factor beta receptor signaling pathway / Zygotic genome activation (ZGA) / DNA repair-dependent chromatin remodeling / PI5P Regulates TP53 Acetylation / RUNX1 interacts with co-factors whose precise effect on RUNX1 targets is not known / histone acetyltransferase complex / : / RUNX3 regulates p14-ARF / intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / negative regulation of gluconeogenesis / pre-mRNA intronic binding / Attenuation phase / canonical Wnt signaling pathway / protein-lysine-acetyltransferase activity / positive regulation of T-helper 17 cell lineage commitment / negative regulation of tumor necrosis factor-mediated signaling pathway / somitogenesis / NF-kappaB binding / fat cell differentiation / histone acetyltransferase / skeletal muscle tissue development / negative regulation of protein-containing complex assembly / NR1H3 & NR1H2 regulate gene expression linked to cholesterol transport and efflux / SARS-CoV-1 targets host intracellular signalling and regulatory pathways / regulation of cellular response to heat / Regulation of TP53 Activity through Acetylation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / canonical NF-kappaB signal transduction Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
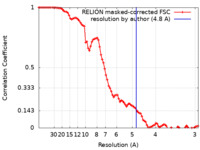
| Method | single particle reconstruction / cryo EM / Resolution: 4.8 Å | |||||||||
 Authors Authors | Kikuchi M / Morita S / Wakamori M / Shin S / Uchikubo-Kamo T / Shirouzu M / Umehara T | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Epigenetic mechanisms to propagate histone acetylation by p300/CBP. Authors: Masaki Kikuchi / Satoshi Morita / Masatoshi Wakamori / Shin Sato / Tomomi Uchikubo-Kamo / Takehiro Suzuki / Naoshi Dohmae / Mikako Shirouzu / Takashi Umehara /  Abstract: Histone acetylation is important for the activation of gene transcription but little is known about its direct read/write mechanisms. Here, we report cryogenic electron microscopy structures in which ...Histone acetylation is important for the activation of gene transcription but little is known about its direct read/write mechanisms. Here, we report cryogenic electron microscopy structures in which a p300/CREB-binding protein (CBP) multidomain monomer recognizes histone H4 N-terminal tail (NT) acetylation (ac) in a nucleosome and acetylates non-H4 histone NTs within the same nucleosome. p300/CBP not only recognized H4NTac via the bromodomain pocket responsible for reading, but also interacted with the DNA minor grooves via the outside of that pocket. This directed the catalytic center of p300/CBP to one of the non-H4 histone NTs. The primary target that p300 writes by reading H4NTac was H2BNT, and H2BNTac promoted H2A-H2B dissociation from the nucleosome. We propose a model in which p300/CBP replicates histone N-terminal tail acetylation within the H3-H4 tetramer to inherit epigenetic storage, and transcribes it from the H3-H4 tetramer to the H2B-H2A dimers to activate context-dependent gene transcription through local nucleosome destabilization. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34592.map.gz emd_34592.map.gz | 55.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34592-v30.xml emd-34592-v30.xml emd-34592.xml emd-34592.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34592_fsc.xml emd_34592_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_34592.png emd_34592.png | 72.3 KB | ||
| Masks |  emd_34592_msk_1.map emd_34592_msk_1.map | 59.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-34592.cif.gz emd-34592.cif.gz | 6.8 KB | ||
| Others |  emd_34592_half_map_1.map.gz emd_34592_half_map_1.map.gz emd_34592_half_map_2.map.gz emd_34592_half_map_2.map.gz | 46 MB 46 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34592 http://ftp.pdbj.org/pub/emdb/structures/EMD-34592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34592 | HTTPS FTP |
-Related structure data
| Related structure data |  8hajMC  8hagC  8hahC  8haiC  8hakC  8halC  8hamC  8hanC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34592.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34592.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.47 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_34592_msk_1.map emd_34592_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_34592_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34592_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : the p300 catalytic core bound to the H4K12acK16ac nucleosome
| Entire | Name: the p300 catalytic core bound to the H4K12acK16ac nucleosome |
|---|---|
| Components |
|
-Supramolecule #1: the p300 catalytic core bound to the H4K12acK16ac nucleosome
| Supramolecule | Name: the p300 catalytic core bound to the H4K12acK16ac nucleosome type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 / Details: p300 was expressed in insect cells. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Histone H3.1
| Macromolecule | Name: Histone H3.1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 15.305969 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ARTKQTARKS TGGKAPRKQL ATKAARKSAP ATGGVKKPHR YRPGTVALRE IRRYQKSTEL LIRKLPFQRL VREIAQDFKT DLRFQSSAV MALQEACEAY LVGLFEDTNL CAIHAKRVTI MPKDIQLARR IRGERA UniProtKB: Histone H3.1 |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.345289 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SGRGKGGKGL G(ALY)GGA(ALY)RHRK VLRDNIQGIT KPAIRRLARR GGVKRISGLI YEETRGVLKV FLENVIRDAV TY TEHAKRK TVTAMDVVYA LKRQGRTLYG FGG |
-Macromolecule #3: Histone H2A type 1-B/E
| Macromolecule | Name: Histone H2A type 1-B/E / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.034355 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SGRGKQGGKA RAKAKTRSSR AGLQFPVGRV HRLLRKGNYS ERVGAGAPVY LAAVLEYLTA EILELAGNAA RDNKKTRIIP RHLQLAIRN DEELNKLLGR VTIAQGGVLP NIQAVLLPKK TESHHKAKGK UniProtKB: Histone H2A type 1-B/E |
-Macromolecule #4: Histone H2B type 1-J
| Macromolecule | Name: Histone H2B type 1-J / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.804045 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PEPAKSAPAP KKGSKKAVTK AQKKDGKKRK RSRKESYSIY VYKVLKQVHP DTGISSKAMG IMNSFVNDIF ERIAGEASRL AHYNKRSTI TSREIQTAVR LLLPGELAKH AVSEGTKAVT KYTSAK UniProtKB: Histone H2B type 1-J |
-Macromolecule #6: Histone acetyltransferase p300
| Macromolecule | Name: Histone acetyltransferase p300 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO / EC number: histone acetyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 92.314391 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: GSSGSSGIFK PEELRQALMP TLEALYRQDP ESLPFRQPVD PQLLGIPDYF DIVKSPMDLS TIKRKLDTGQ YQEPWQYVDD IWLMFNNAW LYNRKTSRVY KYCSKLSEVF EQEIDPVMQS LGYCCGRKLE FSPQTLCCYG KQLCTIPRDA TYYSYQNRYH F CEKCFNEI ...String: GSSGSSGIFK PEELRQALMP TLEALYRQDP ESLPFRQPVD PQLLGIPDYF DIVKSPMDLS TIKRKLDTGQ YQEPWQYVDD IWLMFNNAW LYNRKTSRVY KYCSKLSEVF EQEIDPVMQS LGYCCGRKLE FSPQTLCCYG KQLCTIPRDA TYYSYQNRYH F CEKCFNEI QGESVSLGDD PSQPQTTINK EQFSKRKNDT LDPELFVECT ECGRKMHQIC VLHHEIIWPA GFVCDGCLKK SA RTRKENK FSAKRLPSTR LGTFLENRVN DFLRRQNHPE SGEVTVRVVH ASDKTVEVKP GMKARFVDSG EMAESFPYRT KAL FAFEEI DGVDLCFFGM HVQEYGSDCP PPNQRRVYIS YLDSVHFFRP KCLRTAVYHE ILIGYLEYVK KLGYTTGHIW ACPP SEGDD YIFHCHPPDQ KIPKPKRLQE WYKKMLDKAV SERIVHDYKD IFKQATEDRL TSAKELPYFE GDFWPNVLEE SIKEL EQEE EERKREENTS NESTDVTKGD SKNAKKKNNK KTSKNKSSLS RGNKKKPGMP NVSNDLSQKL YATMEKHKEV FFVIRL IAG PAANSLPPIV DPDPLIPCDL MDGRDAFLTL ARDKHLEFSS LRRAQWSTMC MLVELHTQSQ DRFVYTCNEC KHHVETR WH CTVCEDYDLC ITCYNTKNHD HKMEKLGLGL DDESNNQQAA ATQSPGDSRR LSIQRCIQSL VHACQCRNAN CSLPSCQK M KRVVQHTKGC KRKTNGGCPI CKQLIALCCY HAKHCQENKC PVPFCLNIKQ KLRQQQLQHR LQQAQMLRRR MASMQ UniProtKB: Histone acetyltransferase p300 |
-Macromolecule #5: DNA (180-mer)
| Macromolecule | Name: DNA (180-mer) / type: dna / ID: 5 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.560527 KDa |
| Sequence | String: (DA)(DT)(DC)(DC)(DG)(DT)(DC)(DC)(DG)(DT) (DT)(DA)(DC)(DC)(DG)(DC)(DC)(DA)(DT)(DC) (DA)(DA)(DT)(DA)(DT)(DC)(DC)(DA)(DC) (DC)(DT)(DG)(DC)(DA)(DG)(DA)(DT)(DT)(DC) (DT) (DA)(DC)(DC)(DA)(DA)(DA) ...String: (DA)(DT)(DC)(DC)(DG)(DT)(DC)(DC)(DG)(DT) (DT)(DA)(DC)(DC)(DG)(DC)(DC)(DA)(DT)(DC) (DA)(DA)(DT)(DA)(DT)(DC)(DC)(DA)(DC) (DC)(DT)(DG)(DC)(DA)(DG)(DA)(DT)(DT)(DC) (DT) (DA)(DC)(DC)(DA)(DA)(DA)(DA)(DG) (DT)(DG)(DT)(DA)(DT)(DT)(DT)(DG)(DG)(DA) (DA)(DA) (DC)(DT)(DG)(DC)(DT)(DC)(DC) (DA)(DT)(DC)(DA)(DA)(DA)(DA)(DG)(DG)(DC) (DA)(DT)(DG) (DT)(DT)(DC)(DA)(DG)(DC) (DT)(DG)(DA)(DA)(DT)(DT)(DC)(DA)(DG)(DC) (DT)(DG)(DA)(DA) (DC)(DA)(DT)(DG)(DC) (DC)(DT)(DT)(DT)(DT)(DG)(DA)(DT)(DG)(DG) (DA)(DG)(DC)(DA)(DG) (DT)(DT)(DT)(DC) (DC)(DA)(DA)(DA)(DT)(DA)(DC)(DA)(DC)(DT) (DT)(DT)(DT)(DG)(DG)(DT) (DA)(DG)(DA) (DA)(DT)(DC)(DT)(DG)(DC)(DA)(DG)(DG)(DT) (DG)(DG)(DA)(DT)(DA)(DT)(DT) (DG)(DA) (DT)(DG)(DG)(DC)(DG)(DG)(DT)(DA)(DA)(DC) (DG)(DG)(DA)(DC)(DG)(DG)(DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.7 µm / Nominal defocus min: 0.9 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)