+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Craspase-NTR | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | IMMUNE SYSTEM-RNA complex | |||||||||
| Function / homology | CHAT domain / CHAT domain / : / CRISPR type III-associated protein / RAMP superfamily / defense response to virus / RAMP superfamily protein / CHAT domain protein Function and homology information Function and homology information | |||||||||
| Biological species |  Candidatus Scalindua brodae (bacteria) Candidatus Scalindua brodae (bacteria) | |||||||||
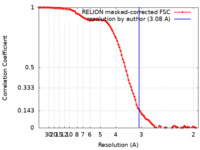
| Method | single particle reconstruction / cryo EM / Resolution: 3.08 Å | |||||||||
 Authors Authors | Feng Y / Zhang L | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2022 Journal: Mol Cell / Year: 2022Title: Target RNA activates the protease activity of Craspase to confer antiviral defense. Authors: Xi Liu / Laixing Zhang / Hao Wang / Yu Xiu / Ling Huang / Zhengyu Gao / Ningning Li / Feixue Li / Weijia Xiong / Teng Gao / Yi Zhang / Maojun Yang / Yue Feng /  Abstract: In the type III-E CRISPR-Cas system, a Cas effector (gRAMP) is associated with a TPR-CHAT to form Craspase (CRISPR-guided caspase). However, both the structural features of gRAMP and the immunity ...In the type III-E CRISPR-Cas system, a Cas effector (gRAMP) is associated with a TPR-CHAT to form Craspase (CRISPR-guided caspase). However, both the structural features of gRAMP and the immunity mechanism remain unknown for this system. Here, we report structures of gRAMP-crRNA and gRAMP:cRNA:target RNA as well as structures of Craspase and Craspase complexed with cognate target RNA (CTR) or non-cognate target RNA (NTR). Importantly, the 3' anti-tag region of NTR and CTR binds at two distinct channels in Craspase, and CTR with a non-complementary 3' anti-tag induces a marked conformational change of the TPR-CHAT, which allosterically activates its protease activity to cleave an ancillary protein Csx30. This cleavage then triggers an abortive infection as the antiviral strategy of the type III-E system. Together, our study provides crucial insights into both the catalytic mechanism of the gRAMP and the immunity mechanism of the type III-E system. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33439.map.gz emd_33439.map.gz | 69.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33439-v30.xml emd-33439-v30.xml emd-33439.xml emd-33439.xml | 25.8 KB 25.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33439_fsc.xml emd_33439_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_33439.png emd_33439.png | 27.5 KB | ||
| Masks |  emd_33439_msk_1.map emd_33439_msk_1.map | 83.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-33439.cif.gz emd-33439.cif.gz | 8 KB | ||
| Others |  emd_33439_additional_1.map.gz emd_33439_additional_1.map.gz emd_33439_half_map_1.map.gz emd_33439_half_map_1.map.gz emd_33439_half_map_2.map.gz emd_33439_half_map_2.map.gz | 6.4 MB 65.4 MB 65.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33439 http://ftp.pdbj.org/pub/emdb/structures/EMD-33439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33439 | HTTPS FTP |
-Related structure data
| Related structure data |  7xt4MC  7xsoC  7xspC  7xsqC  7xsrC  7xssC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33439.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33439.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.97 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_33439_msk_1.map emd_33439_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_33439_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33439_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_33439_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Craspase-NTR complex
| Entire | Name: Craspase-NTR complex |
|---|---|
| Components |
|
-Supramolecule #1: Craspase-NTR complex
| Supramolecule | Name: Craspase-NTR complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Candidatus Scalindua brodae (bacteria) Candidatus Scalindua brodae (bacteria) |
-Supramolecule #2: RAMP superfamily protein, CHAT domain protein
| Supramolecule | Name: RAMP superfamily protein, CHAT domain protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1, #3 |
|---|
-Supramolecule #3: RNA
| Supramolecule | Name: RNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Candidatus Scalindua brodae (bacteria) / Synthetically produced: Yes Candidatus Scalindua brodae (bacteria) / Synthetically produced: Yes |
-Supramolecule #4: RNA (34-MER)
| Supramolecule | Name: RNA (34-MER) / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4 |
|---|
-Macromolecule #1: CHAT domain protein
| Macromolecule | Name: CHAT domain protein / type: protein_or_peptide / ID: 1 Details: Using a different transcription start point, the protein has additional five residues in its N-terminus. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Candidatus Scalindua brodae (bacteria) Candidatus Scalindua brodae (bacteria) |
| Molecular weight | Theoretical: 82.549992 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNNTEENIDR IQEPTREDID RKEAERLLDE AFNPRTKPVD RKKIINSALK ILIGLYKEKK DDLTSASFIS IARAYYLVSI TILPKGTTI PEKKKEALRK GIEFIDRAIN KFNGSILDSQ RAFRIKSVLS IEFNRIDREK CDNIKLKNLL NEAVDKGCTD F DTYEWDIQ ...String: MNNTEENIDR IQEPTREDID RKEAERLLDE AFNPRTKPVD RKKIINSALK ILIGLYKEKK DDLTSASFIS IARAYYLVSI TILPKGTTI PEKKKEALRK GIEFIDRAIN KFNGSILDSQ RAFRIKSVLS IEFNRIDREK CDNIKLKNLL NEAVDKGCTD F DTYEWDIQ IAIRLCELGV DMEGHFDNLI KSNKANDLQK AKAYYFIKKD DHKAKEHMDK CTASLKYTPC SHRLWDETVG FI ERLKGDS STLWRDFAIK TYRSCRVQEK ETGTLRLRWY WSRHRVLYDM AFLAVKEQAD DEEPDVNVKQ AKIKKLAEIS DSL KSRFSL RLSDMEKMPK SDDESNHEFK KFLDKCVTAY QDGYVINRSE DKEGQGENKS TTSKQPEPRP QAKLLELTQV PEGW VVVHF YLNKLEGMGN AIVFDKCANS WQYKEFQYKE LFEVFLTWQA NYNLYKENAA EHLVTLCKKI GETMPFLFCD NFIPN GKDV LFVPHDFLHR LPLHGSIENK TNGKLFLENH SCCYLPAWSF ASEKEASTSD EYVLLKNFDQ GHFETLQNNQ IWGTQS VKD GASSDDLENI RNNPRLLTIL CHGEANMSNP FRSMLKLANG GITYLEILNS VKGLKGSQVI LGACETDLVP PLSDVMD EH YSVATALLLI GAAGVVGTMW KVRSNKTKSL IEWKLENIEY KLNEWQKETG GAAYKDHPPT FYRSIAFRSI GFPL UniProtKB: CHAT domain protein |
-Macromolecule #3: RAMP superfamily protein
| Macromolecule | Name: RAMP superfamily protein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Candidatus Scalindua brodae (bacteria) Candidatus Scalindua brodae (bacteria) |
| Molecular weight | Theoretical: 197.823797 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKSNDMNITV ELTFFEPYRL VEWFDWDARK KSHSAMRGQA FAQWTWKGKG RTAGKSFITG TLVRSAVIKA VEELLSLNNG KWEGVPCCN GSFQTDESKG KKPSFLRKRH TLQWQANNKN ICDKEEACPF CILLGRFDNA GKVHERNKDY DIHFSNFDLD H KQEKNDLR ...String: MKSNDMNITV ELTFFEPYRL VEWFDWDARK KSHSAMRGQA FAQWTWKGKG RTAGKSFITG TLVRSAVIKA VEELLSLNNG KWEGVPCCN GSFQTDESKG KKPSFLRKRH TLQWQANNKN ICDKEEACPF CILLGRFDNA GKVHERNKDY DIHFSNFDLD H KQEKNDLR LVDIASGRIL NRVDFDTGKA KDYFRTWEAD YETYGTYTGR ITLRNEHAKK LLLASLGFVD KLCGALCRIE VI KKSESPL PSDTKEQSYT KDDTVEVLSE DHNDELRKQA EVIVEAFKQN DKLEKIRILA DAIRTLRLHG EGVIEKDELP DGK EERDKG HHLWDIKVQG TALRTKLKEL WQSNKDIGWR KFTEMLGSNL YLIYKKETGG VSTRFRILGD TEYYSKAHDS EGSD LFIPV TPPEGIETKE WIIVGRLKAA TPFYFGVQQP SDSIPGKEKK SEDSLVINEH TSFNILLDKE NRYRIPRSAL RGALR RDLR TAFGSGCNVS LGGQILCNCK VCIEMRRITL KDSVSDFSEP PEIRYRIAKN PGTATVEDGS LFDIEVGPEG LTFPFV LRY RGHKFPEQLS SVIRYWEEND GKNGMAWLGG LDSTGKGRFA LKDIKIFEWD LNQKINEYIK ERGMRGKEKE LLEMGES SL PDGLIPYKFF EERECLFPYK ENLKPQWSEV QYTIEVGSPL LTADTISALT EPGNRDAIAY KKRVYNDGNN AIEPEPRF A VKSETHRGIF RTAVGRRTGD LGKEDHEDCT CDMCIIFGNE HESSKIRFED LELINGNEFE KLEKHIDHVA IDRFTGGAL DKAKFDTYPL AGSPKKPLKL KGRFWIKKGF SGDHKLLITT ALSDIRDGLY PLGSKGGVGY GWVAGISIDD NVPDDFKEMI NKTEMPLPE EVEESNNGPI NNDYVHPGHQ SPKQDHKNKN IYYPHYFLDS GSKVYREKDI ITHEEFTEEL LSGKINCKLE T LTPLIIPD TSDENGLKLQ GNKPGHKNYK FFNINGELMI PGSELRGMLR THFEALTKSC FAIFGEDSTL SWRMNADEKD YK IDSNSIR KMESQRNPKY RIPDELQKEL RNSGNGLFNR LYTSERRFWS DVSNKFENSI DYKREILRCA GRPKNYKGGI IRQ RKDSLM AEELKVHRLP LYDNFDIPDS AYKANDHCRK SATCSTSRGC RERFTCGIKV RDKNRVFLNA ANNNRQYLNN IKKS NHDLY LQYLKGEKKI RFNSKVITGS ERSPIDVIAE LNERGRQTGF IKLSGLNNSN KSQGNTGTTF NSGWDRFELN ILLDD LETR PSKSDYPRPR LLFTKDQYEY NITKRCERVF EIDKGNKTGY PVDDQIKKNY EDILDSYDGI KDQEVAERFD TFTRGS KLK VGDLVYFHID GDNKIDSLIP VRISRKCASK TLGGKLDKAL HPCTGLSDGL CPGCHLFGTT DYKGRVKFGF AKYENGP EW LITRGNNPER SLTLGVLESP RPAFSIPDDE SEIPGRKFYL HHNGWRIIRQ KQLEIRETVQ PERNVTTEVM DKGNVFSF D VRFENLREWE LGLLLQSLDP GKNIAHKLGK GKPYGFGSVK IKIDSLHTFK INSNNDKIKR VPQSDIREYI NKGYQKLIE WSGNNSIQKG NVLPQWHVIP HIDKLYKLLW VPFLNDSKLE PDVRYPVLNE ESKGYIEGSD YTYKKLGDKD NLPYKTRVKG LTTPWSPWN PFQVIAEHEE QEVNVTGSRP SVTDKIERDG KMV UniProtKB: RAMP superfamily protein |
-Macromolecule #2: RNA (5'-R(P*GP*GP*GP*GP*CP*AP*GP*AP*AP*AP*AP*UP*UP*GP*GP*GP*UP*AP...
| Macromolecule | Name: RNA (5'-R(P*GP*GP*GP*GP*CP*AP*GP*AP*AP*AP*AP*UP*UP*GP*GP*GP*UP*AP*CP*CP*G)-3') type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Candidatus Scalindua brodae (bacteria) Candidatus Scalindua brodae (bacteria) |
| Molecular weight | Theoretical: 14.906905 KDa |
| Sequence | String: CUCUAGUAAC AGCCGUGGAG UCCGGGGCAG AAAAUUGGGU ACCGUG |
-Macromolecule #4: RNA (34-MER)
| Macromolecule | Name: RNA (34-MER) / type: rna / ID: 4 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Candidatus Scalindua brodae (bacteria) Candidatus Scalindua brodae (bacteria) |
| Molecular weight | Theoretical: 17.828559 KDa |
| Sequence | String: GACUUAAUGU CACGGUACCC AAUUUUCUGC CCCGGACUCC ACGGCUGUUA CUAGAG |
-Macromolecule #5: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 5 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: DARK FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.7 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)