+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3302 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the Salipro-POT complex at 6.48 A | |||||||||
 Map data Map data | Reconstruction of Salipro-POT | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | POT / salipro / lipid / disk / membrane / protein / transporter / peptider | |||||||||
| Function / homology |  Function and homology information Function and homology informationdipeptide transmembrane transport / tripeptide transmembrane transport / tripeptide transmembrane transporter activity / peptide:proton symporter activity / dipeptide transmembrane transporter activity / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Shewanella oneidensis (bacteria) Shewanella oneidensis (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.48 Å | |||||||||
 Authors Authors | Frauenfeld J / Loeving R / Armache JP / Sonnen A / Guettou F / Moberg P / Zhu L / Jegerschoeld C / Flayhan A / Briggs J ...Frauenfeld J / Loeving R / Armache JP / Sonnen A / Guettou F / Moberg P / Zhu L / Jegerschoeld C / Flayhan A / Briggs J / Garoff H / Loew C / Cheng Y / Nordlund P | |||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2016 Journal: Nat Methods / Year: 2016Title: A saposin-lipoprotein nanoparticle system for membrane proteins. Authors: Jens Frauenfeld / Robin Löving / Jean-Paul Armache / Andreas F-P Sonnen / Fatma Guettou / Per Moberg / Lin Zhu / Caroline Jegerschöld / Ali Flayhan / John A G Briggs / Henrik Garoff / ...Authors: Jens Frauenfeld / Robin Löving / Jean-Paul Armache / Andreas F-P Sonnen / Fatma Guettou / Per Moberg / Lin Zhu / Caroline Jegerschöld / Ali Flayhan / John A G Briggs / Henrik Garoff / Christian Löw / Yifan Cheng / Pär Nordlund /    Abstract: A limiting factor in membrane protein research is the ability to solubilize and stabilize such proteins. Detergents are used most often for solubilizing membrane proteins, but they are associated ...A limiting factor in membrane protein research is the ability to solubilize and stabilize such proteins. Detergents are used most often for solubilizing membrane proteins, but they are associated with protein instability and poor compatibility with structural and biophysical studies. Here we present a saposin-lipoprotein nanoparticle system, Salipro, which allows for the reconstitution of membrane proteins in a lipid environment that is stabilized by a scaffold of saposin proteins. We demonstrate the applicability of the method on two purified membrane protein complexes as well as by the direct solubilization and nanoparticle incorporation of a viral membrane protein complex from the virus membrane. Our approach facilitated high-resolution structural studies of the bacterial peptide transporter PeptTSo2 by single-particle cryo-electron microscopy (cryo-EM) and allowed us to stabilize the HIV envelope glycoprotein in a functional state. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3302.map.gz emd_3302.map.gz | 45.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3302-v30.xml emd-3302-v30.xml emd-3302.xml emd-3302.xml | 9.1 KB 9.1 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-3302.png EMD-3302.png | 763.3 KB | ||
| Others |  emd_3302_additional_1.map.gz emd_3302_additional_1.map.gz emd_3302_half_map_1.map.gz emd_3302_half_map_1.map.gz emd_3302_half_map_2.map.gz emd_3302_half_map_2.map.gz | 2.6 MB 45.8 MB 45.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3302 http://ftp.pdbj.org/pub/emdb/structures/EMD-3302 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3302 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3302 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3302.map.gz / Format: CCP4 / Size: 62.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3302.map.gz / Format: CCP4 / Size: 62.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of Salipro-POT | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.2156 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 3302 additional 1.map
| File | emd_3302_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
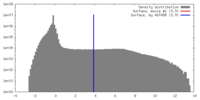
| Density Histograms |
-Supplemental map: emd 3302 half map 1.map
| File | emd_3302_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Supplemental map: emd 3302 half map 2.map
| File | emd_3302_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Salipro-POT
| Entire | Name: Salipro-POT |
|---|---|
| Components |
|
-Supramolecule #1000: Salipro-POT
| Supramolecule | Name: Salipro-POT / type: sample / ID: 1000 / Oligomeric state: One tetramer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 224 MDa / Theoretical: 250 MDa |
-Macromolecule #1: Proton-coupled oligopeptide transporter
| Macromolecule | Name: Proton-coupled oligopeptide transporter / type: protein_or_peptide / ID: 1 / Name.synonym: PeptTSo2, / Number of copies: 1 / Oligomeric state: Tetramer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Shewanella oneidensis (bacteria) Shewanella oneidensis (bacteria) |
| Molecular weight | Experimental: 224 MDa / Theoretical: 250 MDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Proton:oligopeptide symporter POT family |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 120 K / Instrument: FEI VITROBOT MARK I / Method: Blot for 6 before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Date | Sep 15, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 288 / Average electron dose: 10 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 41132 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 0.0026 µm / Nominal defocus min: 0.0018 µm / Nominal magnification: 31000 |
| Sample stage | Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Processed with Relion 1.3 |
|---|---|
| CTF correction | Details: each particle |
| Final reconstruction | Applied symmetry - Point group: C4 (4 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 6.48 Å / Resolution method: OTHER / Software - Name: Relion / Number images used: 9913 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)