+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31148 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | human voltage-gated potassium channel KV1.3 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | potassium channel / complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpinceau fiber / methylglyoxal reductase (NADPH) (acetol producing) activity / : / regulation of protein localization to cell surface / voltage-gated monoatomic ion channel activity / delayed rectifier potassium channel activity / Voltage gated Potassium channels / Oxidoreductases; Acting on the CH-OH group of donors; With NAD+ or NADP+ as acceptor / juxtaparanode region of axon / regulation of potassium ion transmembrane transport ...pinceau fiber / methylglyoxal reductase (NADPH) (acetol producing) activity / : / regulation of protein localization to cell surface / voltage-gated monoatomic ion channel activity / delayed rectifier potassium channel activity / Voltage gated Potassium channels / Oxidoreductases; Acting on the CH-OH group of donors; With NAD+ or NADP+ as acceptor / juxtaparanode region of axon / regulation of potassium ion transmembrane transport / tertiary granule membrane / action potential / voltage-gated potassium channel activity / potassium channel regulator activity / specific granule membrane / voltage-gated potassium channel complex / potassium ion transmembrane transport / potassium ion transport / protein homooligomerization / cytoplasmic side of plasma membrane / transmembrane transporter binding / cytoskeleton / axon / Neutrophil degranulation / synapse / perinuclear region of cytoplasm / membrane / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Liu S / Zhao Y | |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2021 Journal: Cell Discov / Year: 2021Title: Structures of wild-type and H451N mutant human lymphocyte potassium channel K1.3. Authors: Sanling Liu / Yue Zhao / Hao Dong / Liang Xiao / Yong Zhang / Yuqin Yang / Seow Theng Ong / K George Chandy / Longhua Zhang / Changlin Tian /   | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31148.map.gz emd_31148.map.gz | 7.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31148-v30.xml emd-31148-v30.xml emd-31148.xml emd-31148.xml | 10.8 KB 10.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_31148_fsc.xml emd_31148_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_31148.png emd_31148.png | 98.5 KB | ||
| Filedesc metadata |  emd-31148.cif.gz emd-31148.cif.gz | 5.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31148 http://ftp.pdbj.org/pub/emdb/structures/EMD-31148 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31148 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31148 | HTTPS FTP |
-Related structure data
| Related structure data |  7ej1MC  7ej2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31148.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31148.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.014 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
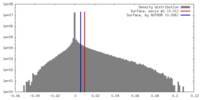
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human potassium channel KV1.3/KVb2.1 complex
| Entire | Name: Human potassium channel KV1.3/KVb2.1 complex |
|---|---|
| Components |
|
-Supramolecule #1: Human potassium channel KV1.3/KVb2.1 complex
| Supramolecule | Name: Human potassium channel KV1.3/KVb2.1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Voltage-gated potassium channel subunit beta-2
| Macromolecule | Name: Voltage-gated potassium channel subunit beta-2 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO EC number: Oxidoreductases; Acting on the CH-OH group of donors; With NAD+ or NADP+ as acceptor |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.049152 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MYPESTTGSP ARLSLRQTGS PGMIYSTRYG SPKRQLQFYR NLGKSGLRVS CLGLGTWVTF GGQITDEMAE QLMTLAYDNG INLFDTAEV YAAGKAEVVL GNIIKKKGWR RSSLVITTKI FWGGKAETER GLSRKHIIEG LKASLERLQL EYVDVVFANR P DPNTPMEE ...String: MYPESTTGSP ARLSLRQTGS PGMIYSTRYG SPKRQLQFYR NLGKSGLRVS CLGLGTWVTF GGQITDEMAE QLMTLAYDNG INLFDTAEV YAAGKAEVVL GNIIKKKGWR RSSLVITTKI FWGGKAETER GLSRKHIIEG LKASLERLQL EYVDVVFANR P DPNTPMEE TVRAMTHVIN QGMAMYWGTS RWSSMEIMEA YSVARQFNLT PPICEQAEYH MFQREKVEVQ LPELFHKIGV GA MTWSPLA CGIVSGKYDS GIPPYSRASL KGYQWLKDKI LSEEGRRQQA KLKELQAIAE RLGCTLPQLA IAWCLRNEGV SSV LLGASN ADQLMENIGA IQVLPKLSSS IIHEIDSILG NKPYSKKDYR S UniProtKB: Voltage-gated potassium channel subunit beta-2 |
-Macromolecule #2: Potassium voltage-gated channel subfamily A member 3
| Macromolecule | Name: Potassium voltage-gated channel subfamily A member 3 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 63.90648 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDERLSLLRS PPPPSARHRA HPPQRPASSG GAHTLVNHGY AEPAAGRELP PDMTVVPGDH LLEPEVADGG GAPPQGGCGG GGCDRYEPL PPSLPAAGEQ DCCGERVVIN ISGLRFETQL KTLCQFPETL LGDPKRRMRY FDPLRNEYFF DRNRPSFDAI L YYYQSGGR ...String: MDERLSLLRS PPPPSARHRA HPPQRPASSG GAHTLVNHGY AEPAAGRELP PDMTVVPGDH LLEPEVADGG GAPPQGGCGG GGCDRYEPL PPSLPAAGEQ DCCGERVVIN ISGLRFETQL KTLCQFPETL LGDPKRRMRY FDPLRNEYFF DRNRPSFDAI L YYYQSGGR IRRPVNVPID IFSEEIRFYQ LGEEAMEKFR EDEGFLREEE RPLPRRDFQR QVWLLFEYPE SSGPARGIAI VS VLVILIS IVIFCLETLP EFRDEKDYPA STSQDSFEAA GNSTSGSRAG ASSFSDPFFV VETLCIIWFS FELLVRFFAC PSK ATFSRN IMNLIDIVAI IPYFITLGTE LAERQGNGQQ AMSLAILRVI RLVRVFRIFK LSRHSKGLQI LGQTLKASMR ELGL LIFFL FIGVILFSSA VYFAEADDPT SGFSSIPDAF WWAVVTMTTV GYGDMHPVTI GGKIVGSLCA IAGVLTIALP VPVIV SNFN YFYHRETEGE EQSQYMHVGS CQHLSSSAEE LRKARSNSTL SKSEYMVIEE GGMNHSAFPQ TPFKTGNSTA TCTTNN NPN SCVNIKKIFT DV UniProtKB: Potassium voltage-gated channel subfamily A member 3 |
-Macromolecule #3: NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
| Macromolecule | Name: NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: NAP |
|---|---|
| Molecular weight | Theoretical: 743.405 Da |
| Chemical component information |  ChemComp-NAP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 62.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)