[English] 日本語
 Yorodumi
Yorodumi- EMDB-31032: Cryo-EM structure of Gi-bound metabotropic glutamate receptor mGlu4 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31032 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
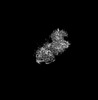
| Title | Cryo-EM structure of Gi-bound metabotropic glutamate receptor mGlu4 | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | mGlu4 / GPCR / Cryo-EM / complex / MEMBRANE PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationadenylate cyclase-inhibiting G protein-coupled glutamate receptor signaling pathway / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / negative regulation of adenylate cyclase activity / G protein-coupled glutamate receptor signaling pathway / neurotransmitter secretion / Class C/3 (Metabotropic glutamate/pheromone receptors) / GTP metabolic process / glutamate receptor activity / positive regulation of macroautophagy / regulation of synaptic transmission, glutamatergic ...adenylate cyclase-inhibiting G protein-coupled glutamate receptor signaling pathway / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / negative regulation of adenylate cyclase activity / G protein-coupled glutamate receptor signaling pathway / neurotransmitter secretion / Class C/3 (Metabotropic glutamate/pheromone receptors) / GTP metabolic process / glutamate receptor activity / positive regulation of macroautophagy / regulation of synaptic transmission, glutamatergic / Adenylate cyclase inhibitory pathway / regulation of neuron apoptotic process / G protein-coupled receptor binding / G protein-coupled receptor activity / G-protein beta/gamma-subunit complex binding / centriolar satellite / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G beta:gamma signalling through BTK / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / GDP binding / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / sperm principal piece / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / presynapse / retina development in camera-type eye / GTPase binding / fibroblast proliferation / cytoplasmic vesicle / midbody / Ca2+ pathway / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / chemical synaptic transmission / Ras protein signal transduction / positive regulation of MAPK cascade / Extra-nuclear estrogen signaling / cell population proliferation / ciliary basal body / G protein-coupled receptor signaling pathway / cell division / lysosomal membrane / GTPase activity / centrosome / synapse / GTP binding / protein-containing complex binding / nucleolus / Golgi apparatus / signal transduction / extracellular exosome / nucleoplasm / membrane / metal ion binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | ||||||||||||||||||
 Authors Authors | Lin S / Han S | ||||||||||||||||||
| Funding support |  China, 5 items China, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: Structures of G-bound metabotropic glutamate receptors mGlu2 and mGlu4. Authors: Shuling Lin / Shuo Han / Xiaoqing Cai / Qiuxiang Tan / Kexiu Zhou / Dejian Wang / Xinwei Wang / Juan Du / Cuiying Yi / Xiaojing Chu / Antao Dai / Yan Zhou / Yan Chen / Yu Zhou / Hong Liu / ...Authors: Shuling Lin / Shuo Han / Xiaoqing Cai / Qiuxiang Tan / Kexiu Zhou / Dejian Wang / Xinwei Wang / Juan Du / Cuiying Yi / Xiaojing Chu / Antao Dai / Yan Zhou / Yan Chen / Yu Zhou / Hong Liu / Jianfeng Liu / Dehua Yang / Ming-Wei Wang / Qiang Zhao / Beili Wu /  Abstract: The metabotropic glutamate receptors (mGlus) have key roles in modulating cell excitability and synaptic transmission in response to glutamate (the main excitatory neurotransmitter in the central ...The metabotropic glutamate receptors (mGlus) have key roles in modulating cell excitability and synaptic transmission in response to glutamate (the main excitatory neurotransmitter in the central nervous system). It has previously been suggested that only one receptor subunit within an mGlu homodimer is responsible for coupling to G protein during receptor activation. However, the molecular mechanism that underlies the asymmetric signalling of mGlus remains unknown. Here we report two cryo-electron microscopy structures of human mGlu2 and mGlu4 bound to heterotrimeric G protein. The structures reveal a G-protein-binding site formed by three intracellular loops and helices III and IV that is distinct from the corresponding binding site in all of the other G-protein-coupled receptor (GPCR) structures. Furthermore, we observed an asymmetric dimer interface of the transmembrane domain of the receptor in the two mGlu-G structures. We confirmed that the asymmetric dimerization is crucial for receptor activation, which was supported by functional data; this dimerization may provide a molecular basis for the asymmetric signal transduction of mGlus. These findings offer insights into receptor signalling of class C GPCRs. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31032.map.gz emd_31032.map.gz | 158.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31032-v30.xml emd-31032-v30.xml emd-31032.xml emd-31032.xml | 16.7 KB 16.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31032.png emd_31032.png | 33.8 KB | ||
| Filedesc metadata |  emd-31032.cif.gz emd-31032.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31032 http://ftp.pdbj.org/pub/emdb/structures/EMD-31032 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31032 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31032 | HTTPS FTP |
-Related structure data
| Related structure data |  7e9hMC  7e9gC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31032.map.gz / Format: CCP4 / Size: 189.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31032.map.gz / Format: CCP4 / Size: 189.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
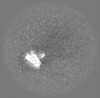
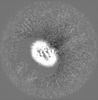
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
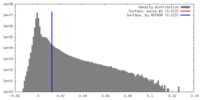
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Complex of mGlu4 homodimer with heterotrimeric Gi3 protein in pre...
| Entire | Name: Complex of mGlu4 homodimer with heterotrimeric Gi3 protein in presence of antibody |
|---|---|
| Components |
|
-Supramolecule #1: Complex of mGlu4 homodimer with heterotrimeric Gi3 protein in pre...
| Supramolecule | Name: Complex of mGlu4 homodimer with heterotrimeric Gi3 protein in presence of antibody type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Metabotropic glutamate receptor 4
| Macromolecule | Name: Metabotropic glutamate receptor 4 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 99.458359 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: DYKDDDDGAP KPKGHPHMNS IRIDGDITLG GLFPVHGRGS EGKPCGELKK EKGIHRLEAM LFALDRINND PDLLPNITLG ARILDTCSR DTHALEQSLT FVQALIEKDG TEVRCGSGGP PIITKPERVV GVIGASGSSV SIMVANILRL FKIPQISYAS T APDLSDNS ...String: DYKDDDDGAP KPKGHPHMNS IRIDGDITLG GLFPVHGRGS EGKPCGELKK EKGIHRLEAM LFALDRINND PDLLPNITLG ARILDTCSR DTHALEQSLT FVQALIEKDG TEVRCGSGGP PIITKPERVV GVIGASGSSV SIMVANILRL FKIPQISYAS T APDLSDNS RYDFFSRVVP SDTYQAQAMV DIVRALKWNY VSTVASEGSY GESGVEAFIQ KSREDGGVCI AQSVKIPREP KA GEFDKII RRLLETSNAR AVIIFANEDD IRRVLEAARR ANQTGHFFWM GSDSWGSKIA PVLHLEEVAE GAVTILPKRM SVR GFDRYF SSRTLDNNRR NIWFAEFWED NFHCKLSRHA LKKGSHVKKC TNRERIGQDS AYEQEGKVQF VIDAVYAMGH ALHA MHRDL CPGRVGLCPR MDPVDGTQLL KYIRNVNFSG IAGNPVTFNE NGDAPGRYDI YQYQLRNDSA EYKVIGSWTD HLHLR IERM HWPGSGQQLP RSICSLPCQP GERKKTVKGM PCCWHCEPCT GYQYQVDRYT CKTCPYDMRP TENRTGCRPI PIIKLE WGS PWAVLPLFLA VVGIAATLFV VITFVRYNDT PIVKASGREL SYVLLAGIFL CYATTFLMIA EPDLGTCSLR RIFLGLG MS ISYAALLTKT NRIYRIFEQG KRSVSAPRFI SPASQLAITF SLISLQLLGI CVWFVVDPSH SVVDFQDQRT LDPRFARG V LKCDISDLSL ICLLGYSMLL MVTCTVYAIK TRGVPETFNE AKPIGFTMYT TCIVWLAFIP IFFGTSQSAD KLYIQTTTL TVSVSLSASV SLGMLYMPKV YIILFHPEQN VPKRKRSLKA VVTAATMSNK FTQKGNFRPN GEAKSELCEN LEAPALATKQ TYVTYTNHA I UniProtKB: Metabotropic glutamate receptor 4 |
-Macromolecule #2: Guanine nucleotide-binding protein G(i) subunit alpha-3
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-3 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.55207 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AKEVKLLLLG AGESGKNTIV KQMKIIHEDG YSEDECKQYK VVVYSNTIQS IIAIIRAMG RLKIDFGEAA RADDARQLFV LAGSAEEGVM TPELAGVIKR LWRDGGVQAC FSRSREYQLN DSASYYLNDL D RISQSNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AKEVKLLLLG AGESGKNTIV KQMKIIHEDG YSEDECKQYK VVVYSNTIQS IIAIIRAMG RLKIDFGEAA RADDARQLFV LAGSAEEGVM TPELAGVIKR LWRDGGVQAC FSRSREYQLN DSASYYLNDL D RISQSNYI PTQQDVLRTR VKTTGIVETH FTDKDLYFKM FDVGAQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHASM KLFDSICNNK WFTETSIILF LNKKDLFEEK IKRSPLTICY PEYTGSNTYE EAAAYIQCQF EDLNRRKDTK EIY THFTCS TDTKNVQFVF DAVTDVIIKN NLKECGLY UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-3 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.744371 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI ...String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI VTSSGDTTCA LWDIETGQQT TTFTGHTGDV MSLSLAPDTR LFVSGACDAS AKLWDVREGM CRQTFTGHES DI NAICFFP NGNAFATGSD DATCRLFDLR ADQELMTYSH DNIICGITSV SFSKSGRLLL AGYDDFNCNV WDALKADRAG VLA GHDNRV SCLGVTDDGM AVATGSWDSF LKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.409588 KDa |
| Recombinant expression | Organism:  Baculovirus expression vector pFastBac1-HM Baculovirus expression vector pFastBac1-HM |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAALEVLFQ |
-Macromolecule #6: PHOSPHOSERINE
| Macromolecule | Name: PHOSPHOSERINE / type: ligand / ID: 6 / Number of copies: 2 / Formula: SEP |
|---|---|
| Molecular weight | Theoretical: 185.072 Da |
| Chemical component information | 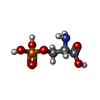 ChemComp-SEP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 617760 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)