+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30766 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mouse radial spoke complex | |||||||||
 Map data Map data | Mouse radial spoke head complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cilia / Complex / Signal transducer. / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmaintenance of ciliary planar beating movement pattern / radial spoke head 1 / radial spoke head 2 / radial spoke head 3 / radial spoke assembly / radial spoke head / mating / axonemal central apparatus assembly / radial spoke / outer dense fiber ...maintenance of ciliary planar beating movement pattern / radial spoke head 1 / radial spoke head 2 / radial spoke head 3 / radial spoke assembly / radial spoke head / mating / axonemal central apparatus assembly / radial spoke / outer dense fiber / epithelial cilium movement involved in extracellular fluid movement / cilium movement involved in cell motility / 9+2 motile cilium / axoneme assembly / cilium movement / kinocilium / flagellated sperm motility / meiotic spindle / motile cilium / axoneme / spermatid development / sperm flagellum / condensed nuclear chromosome / meiotic cell cycle / establishment of localization in cell / extracellular region / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
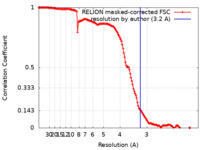
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Zheng W / Cong Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2021 Journal: Proc Natl Acad Sci U S A / Year: 2021Title: Distinct architecture and composition of mouse axonemal radial spoke head revealed by cryo-EM. Authors: Wei Zheng / Fan Li / Zhanyu Ding / Hao Liu / Lei Zhu / Cong Xu / Jiawei Li / Qi Gao / Yanxing Wang / Zhenglin Fu / Chao Peng / Xiumin Yan / Xueliang Zhu / Yao Cong /  Abstract: The radial spoke (RS) heads of motile cilia and flagella contact projections of the central pair (CP) apparatus to coordinate motility, but the morphology is distinct for protozoa and metazoa. Here ...The radial spoke (RS) heads of motile cilia and flagella contact projections of the central pair (CP) apparatus to coordinate motility, but the morphology is distinct for protozoa and metazoa. Here we show the murine RS head is compositionally distinct from that of Our reconstituted murine RS head core complex consists of Rsph1, Rsph3b, Rsph4a, and Rsph9, lacking Rsph6a and Rsph10b, whose orthologs exist in the protozoan RS head. We resolve its cryo-electron microscopy (cryo-EM) structure at 3.2-Å resolution. Our atomic model further reveals a twofold symmetric brake pad-shaped structure, in which Rsph4a and Rsph9 form a compact body extended laterally with two long arms of twisted Rsph1 β-sheets and potentially connected dorsally via Rsph3b to the RS stalk. Furthermore, our modeling suggests that the core complex contacts the periodic CP projections either rigidly through its tooth-shaped Rsph4a regions or elastically through both arms for optimized RS-CP interactions and mechanosignal transduction. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30766.map.gz emd_30766.map.gz | 14.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30766-v30.xml emd-30766-v30.xml emd-30766.xml emd-30766.xml | 12.3 KB 12.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30766_fsc.xml emd_30766_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_30766.png emd_30766.png | 71 KB | ||
| Filedesc metadata |  emd-30766.cif.gz emd-30766.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30766 http://ftp.pdbj.org/pub/emdb/structures/EMD-30766 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30766 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30766 | HTTPS FTP |
-Validation report
| Summary document |  emd_30766_validation.pdf.gz emd_30766_validation.pdf.gz | 353.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_30766_full_validation.pdf.gz emd_30766_full_validation.pdf.gz | 352.9 KB | Display | |
| Data in XML |  emd_30766_validation.xml.gz emd_30766_validation.xml.gz | 13.8 KB | Display | |
| Data in CIF |  emd_30766_validation.cif.gz emd_30766_validation.cif.gz | 18.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30766 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30766 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30766 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30766 | HTTPS FTP |
-Related structure data
| Related structure data |  7dmpMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_30766.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30766.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mouse radial spoke head complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.02 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mouse radial spoke head complex
| Entire | Name: Mouse radial spoke head complex |
|---|---|
| Components |
|
-Supramolecule #1: Mouse radial spoke head complex
| Supramolecule | Name: Mouse radial spoke head complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Radial spoke head protein 4 homolog A
| Macromolecule | Name: Radial spoke head protein 4 homolog A / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 80.214102 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MENSTSLKQE KENQEPGEAE RLWQGESDVS PQEPGPPSPE YREEEQRTDT EPAPRMSPSW SHQSRVSLST GDLTAGPEVS SSPPPPPLQ FHSTPLNTET TQDPVAASPT EKTANGIADT GTPYSDPWES SSAAKQSTSH YTSHAEESTF PQSQTPQPDL C GLRDASRN ...String: MENSTSLKQE KENQEPGEAE RLWQGESDVS PQEPGPPSPE YREEEQRTDT EPAPRMSPSW SHQSRVSLST GDLTAGPEVS SSPPPPPLQ FHSTPLNTET TQDPVAASPT EKTANGIADT GTPYSDPWES SSAAKQSTSH YTSHAEESTF PQSQTPQPDL C GLRDASRN KSKHKGLRFD LLQEEGSDSN CDPDQPEVGA SEAAQSMLEV AIQNAKAYLL STSSKSGLNL YDHLSKVLTK IL DERPADA VDIIENISQD VKMAHFNKKL DTLHNEYEML PAYEIAETQK ALFLQGHLEG ADSELEEEMA ESSLPNVMES AYY FEQAGV GLGTDETYRV FLALKQLTDT HPIQRCRFWG KILGLEMNYI VAEVEFRDGE DEEEVEEEGI AEERDNGGSE AGEE EEEEL PKSLYKAPQV IPKEESRTGA NKYVYFVCNV PGRPWVRLPS VTPAQIVTAR KIKKFFTGRL DAAVISYPPF PGNES NYLR AQIARISAGT HVSPLGFYQF GEEEGEEEEV EGGRDSYEEN PDFEGIQVID LVESLSNWVH HVQYILPQGR CNWFNP IQK DEDEEEEEEE DEEKGEEPDY IEQEVGPPLL TPISEDLGIQ NIPSWTTQLS SNLIPQYAIA VLRSNLWPGA YAFSNGK KF ENFYIGWGHK YCVENYTPPS PPPVYQEYPS GPEITEMNDP SVEEEQAFRM TQEPVALSTE ENEGTEDEDE DDED UniProtKB: Radial spoke head protein 4 homolog A |
-Macromolecule #2: Radial spoke head 1 homolog
| Macromolecule | Name: Radial spoke head 1 homolog / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 34.218254 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSDLGSEELE EEGENDLGEY EGERNEVGER HGHGKARLPN GDTYEGSYEF GKRHGQGTYK FKNGARYTGD YVKNKKHGQG TFIYPDGSR YEGEWADDQR HGQGVYYYVN NDTYTGEWFN HQRHGQGTYL YAETGSKYVG TWVHGQQEGA AELIHLNHRY Q GKFMNKNP ...String: MSDLGSEELE EEGENDLGEY EGERNEVGER HGHGKARLPN GDTYEGSYEF GKRHGQGTYK FKNGARYTGD YVKNKKHGQG TFIYPDGSR YEGEWADDQR HGQGVYYYVN NDTYTGEWFN HQRHGQGTYL YAETGSKYVG TWVHGQQEGA AELIHLNHRY Q GKFMNKNP VGPGKYVFDI GCEQHGEYRL TDTERGEEEE EEETLVNIVP KWKALNITEL ALWTPTLSEE QPPPEGQGQE EP QGLTGVG DPSEDIQAEG FEGELEPRGA DEDVDTFRQE SQENSYDIDQ GNLNFDEEPS DLQD UniProtKB: Radial spoke head 1 homolog |
-Macromolecule #3: Radial spoke head protein 9 homolog
| Macromolecule | Name: Radial spoke head protein 9 homolog / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 31.365902 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDADSLLLSL ELASGSGQGL SPDRRASLLT SLMLVKRDYR FARVLFWGRI LGLVADYYIA QGLSEDQLAP RKTLYSLNCT EWSLLPPAT EEMAMQISVV SGRFMGDPSH EYEHTELQKV NEGEKVFDEE VVVQIKEETR LVSIIDQIDK AVAIIPRGAL F KTPFGVTH ...String: MDADSLLLSL ELASGSGQGL SPDRRASLLT SLMLVKRDYR FARVLFWGRI LGLVADYYIA QGLSEDQLAP RKTLYSLNCT EWSLLPPAT EEMAMQISVV SGRFMGDPSH EYEHTELQKV NEGEKVFDEE VVVQIKEETR LVSIIDQIDK AVAIIPRGAL F KTPFGVTH VNRTFEGLPL SEVRKLSSYF HFREAIDLKN KTLLEKSDLE PSLDFLDSLE YDIPRGSWSI QMERGNALVV LR SLLWPGL TFYHAPRTKN YGYIYVGTGE KNMDLPFML UniProtKB: Radial spoke head protein 9 homolog |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)