+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30437 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human CLCN7-OSTM1 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / Chloride channel / lysosome / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationtransepithelial chloride transport / chloride:proton antiporter activity / response to pH / chloride transmembrane transporter activity / chloride channel activity / chloride channel complex / osteoclast differentiation / Stimuli-sensing channels / lysosomal membrane / intracellular membrane-bounded organelle ...transepithelial chloride transport / chloride:proton antiporter activity / response to pH / chloride transmembrane transporter activity / chloride channel activity / chloride channel complex / osteoclast differentiation / Stimuli-sensing channels / lysosomal membrane / intracellular membrane-bounded organelle / ATP binding / membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Yang GH / Lu YM | |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of the human CLCN7-OSTM1 complex Authors: Yang GH / Lu YM / Zhang YM / Jia YT / Li XM / Lei JL | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30437.map.gz emd_30437.map.gz | 77.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30437-v30.xml emd-30437-v30.xml emd-30437.xml emd-30437.xml | 12.3 KB 12.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30437.png emd_30437.png | 173.2 KB | ||
| Filedesc metadata |  emd-30437.cif.gz emd-30437.cif.gz | 6.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30437 http://ftp.pdbj.org/pub/emdb/structures/EMD-30437 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30437 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30437 | HTTPS FTP |
-Validation report
| Summary document |  emd_30437_validation.pdf.gz emd_30437_validation.pdf.gz | 550.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_30437_full_validation.pdf.gz emd_30437_full_validation.pdf.gz | 549.8 KB | Display | |
| Data in XML |  emd_30437_validation.xml.gz emd_30437_validation.xml.gz | 6.3 KB | Display | |
| Data in CIF |  emd_30437_validation.cif.gz emd_30437_validation.cif.gz | 7.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30437 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30437 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30437 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30437 | HTTPS FTP |
-Related structure data
| Related structure data |  7cq6MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30437.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30437.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.091 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
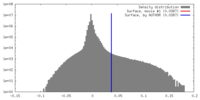
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human CLCN7-OSTM1 complex
| Entire | Name: human CLCN7-OSTM1 complex |
|---|---|
| Components |
|
-Supramolecule #1: human CLCN7-OSTM1 complex
| Supramolecule | Name: human CLCN7-OSTM1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Osteopetrosis-associated transmembrane protein 1
| Macromolecule | Name: Osteopetrosis-associated transmembrane protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.638 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEPGPTAAQR RCSLPPWLPL GLLLWSGLAL GALPFGSSPH RVFHDLLSEQ QLLEVEDLSL SLLQGGGLGP LSLPPDLPDL DPECRELLL DFANSSAELT GCLVRSARPV RLCQTCYPLF QQVVSKMDNI SRAAGNTSES QSCARSLLMA DRMQIVVILS E FFNTTWQE ...String: MEPGPTAAQR RCSLPPWLPL GLLLWSGLAL GALPFGSSPH RVFHDLLSEQ QLLEVEDLSL SLLQGGGLGP LSLPPDLPDL DPECRELLL DFANSSAELT GCLVRSARPV RLCQTCYPLF QQVVSKMDNI SRAAGNTSES QSCARSLLMA DRMQIVVILS E FFNTTWQE ANCANCLTNN SEELSNSTVY FLNLFNHTLT CFEHNLQGNA HSLLQTKNYS EVCKNCREAY KTLSSLYSEM QK MNELENK AEPGTHLCID VEDAMNITRK LWSRTFNCSV PCSDTVPVIA VSVFILFLPV VFYLSSFLHS EQKKRKLILP KRL KSSTSF ANIQENSNLE HHHHHHHH UniProtKB: Osteopetrosis-associated transmembrane protein 1 |
-Macromolecule #2: H(+)/Cl(-) exchange transporter 7
| Macromolecule | Name: H(+)/Cl(-) exchange transporter 7 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 90.906008 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MASDYKDDDD KASDEVDAGT MANVSKKVSW SGRDRDDEEA APLLRRTARP GGGTPLLNGA GPGAARQSPR SALFRVGHMS SVELDDELL DPDMDPPHPF PKEIPHNEKL LSLKYESLDY DNSENQLFLE EERRINHTAF RTVEIKRWVI CALIGILTGL V ACFIDIVV ...String: MASDYKDDDD KASDEVDAGT MANVSKKVSW SGRDRDDEEA APLLRRTARP GGGTPLLNGA GPGAARQSPR SALFRVGHMS SVELDDELL DPDMDPPHPF PKEIPHNEKL LSLKYESLDY DNSENQLFLE EERRINHTAF RTVEIKRWVI CALIGILTGL V ACFIDIVV ENLAGLKYRV IKGNIDKFTE KGGLSFSLLL WATLNAAFVL VGSVIVAFIE PVAAGSGIPQ IKCFLNGVKI PH VVRLKTL VIKVSGVILS VVGGLAVGKE GPMIHSGSVI AAGISQGRST SLKRDFKIFE YFRRDTEKRD FVSAGAAAGV SAA FGAPVG GVLFSLEEGA SFWNQFLTWR IFFASMISTF TLNFVLSIYH GNMWDLSSPG LINFGRFDSE KMAYTIHEIP VFIA MGVVG GVLGAVFNAL NYWLTMFRIR YIHRPCLQVI EAVLVAAVTA TVAFVLIYSS RDCQPLQGGS MSYPLQLFCA DGEYN SMAA AFFNTPEKSV VSLFHDPPGS YNPLTLGLFT LVYFFLACWT YGLTVSAGVF IPSLLIGAAW GRLFGISLSY LTGAAI WAD PGKYALMGAA AQLGGIVRMT LSLTVIMMEA TSNVTYGFPI MLVLMTAKIV GDVFIEGLYD MHIQLQSVPF LHWEAPV TS HSLTAREVMS TPVTCLRRRE KVGVIVDVLS DTASNHNGFP VVEHADDTQP ARLQGLILRS QLIVLLKHKV FVERSNLG L VQRRLRLKDF RDAYPRFPPI QSIHVSQDER ECTMDLSEFM NPSPYTVPQE ASLPRVFKLF RALGLRHLVV VDNRNQVVG LVTRKDLARY RLGKRGLEEL SLAQT UniProtKB: H(+)/Cl(-) exchange transporter 7 |
-Macromolecule #4: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 4 / Number of copies: 4 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.5625 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)