[English] 日本語
 Yorodumi
Yorodumi- EMDB-29490: HIV-1 Vif in complex with human APOBEC3H, CBF-beta, ELOB, ELOC, C... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | HIV-1 Vif in complex with human APOBEC3H, CBF-beta, ELOB, ELOC, CUL5, and RBX2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | virus-host protein complex / ANTIVIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of focal adhesion disassembly / RUNX3 regulates RUNX1-mediated transcription / mRNA Editing: C to U Conversion / Formation of the Editosome / RUNX1 regulates transcription of genes involved in BCR signaling / RUNX1 regulates transcription of genes involved in interleukin signaling / RUNX2 regulates bone development / core-binding factor complex / RUNX1 regulates expression of components of tight junctions / positive regulation of CD8-positive, alpha-beta T cell differentiation ...negative regulation of focal adhesion disassembly / RUNX3 regulates RUNX1-mediated transcription / mRNA Editing: C to U Conversion / Formation of the Editosome / RUNX1 regulates transcription of genes involved in BCR signaling / RUNX1 regulates transcription of genes involved in interleukin signaling / RUNX2 regulates bone development / core-binding factor complex / RUNX1 regulates expression of components of tight junctions / positive regulation of CD8-positive, alpha-beta T cell differentiation / RUNX2 regulates chondrocyte maturation / single-stranded DNA cytosine deaminase / negative regulation of CD4-positive, alpha-beta T cell differentiation / negative regulation of single stranded viral RNA replication via double stranded DNA intermediate / DNA cytosine deamination / cytidine to uridine editing / host-mediated suppression of viral genome replication / clearance of foreign intracellular DNA / RUNX1 and FOXP3 control the development of regulatory T lymphocytes (Tregs) / cytidine deaminase activity / ERBB2 signaling pathway / lymphocyte differentiation / RUNX2 regulates genes involved in cell migration / Transcriptional regulation by RUNX2 / negative regulation of focal adhesion assembly / RUNX2 regulates genes involved in differentiation of myeloid cells / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / transposable element silencing / reelin-mediated signaling pathway / RUNX3 Regulates Immune Response and Cell Migration / cullin-RING-type E3 NEDD8 transferase / NEDD8 transferase activity / myeloid cell differentiation / regulation of neuron migration / definitive hemopoiesis / target-directed miRNA degradation / elongin complex / protein K11-linked ubiquitination / RUNX1 regulates transcription of genes involved in differentiation of myeloid cells / protein neddylation / Regulation of RUNX1 Expression and Activity / NEDD8 ligase activity / response to redox state / VCB complex / Cul5-RING ubiquitin ligase complex / RUNX1 regulates transcription of genes involved in WNT signaling / RUNX1 regulates estrogen receptor mediated transcription / ubiquitin-dependent protein catabolic process via the C-end degron rule pathway / SCF ubiquitin ligase complex / Cul2-RING ubiquitin ligase complex / SCF-dependent proteasomal ubiquitin-dependent protein catabolic process / RUNX2 regulates osteoblast differentiation / ubiquitin ligase complex scaffold activity / RUNX1 interacts with co-factors whose precise effect on RUNX1 targets is not known / Pausing and recovery of Tat-mediated HIV elongation / Tat-mediated HIV elongation arrest and recovery / RUNX3 regulates p14-ARF / HIV elongation arrest and recovery / Pausing and recovery of HIV elongation / cullin family protein binding / Tat-mediated elongation of the HIV-1 transcript / Formation of HIV-1 elongation complex containing HIV-1 Tat / Formation of HIV elongation complex in the absence of HIV Tat / site of DNA damage / RNA Polymerase II Transcription Elongation / Formation of RNA Pol II elongation complex / viral life cycle / endoplasmic reticulum unfolded protein response / cell maturation / RNA Polymerase II Pre-transcription Events / post-translational protein modification / intrinsic apoptotic signaling pathway / transcription corepressor binding / TP53 Regulates Transcription of DNA Repair Genes / transcription initiation at RNA polymerase II promoter / transcription elongation by RNA polymerase II / P-body / G1/S transition of mitotic cell cycle / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Regulation of RUNX3 expression and activity / RING-type E3 ubiquitin transferase / Vif-mediated degradation of APOBEC3G / Inactivation of CSF3 (G-CSF) signaling / Evasion by RSV of host interferon responses / virion component / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / calcium channel activity / Downregulation of ERBB2 signaling / Transcriptional regulation of granulopoiesis / Regulation of expression of SLITs and ROBOs / protein polyubiquitination / ubiquitin-protein transferase activity / osteoblast differentiation / Regulation of RUNX2 expression and activity / ubiquitin protein ligase activity / cell migration / Antigen processing: Ubiquitination & Proteasome degradation / positive regulation of proteasomal ubiquitin-dependent protein catabolic process / RUNX1 regulates transcription of genes involved in differentiation of HSCs / Neddylation Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
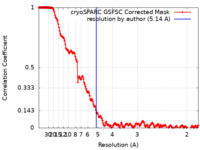
| Method | single particle reconstruction / cryo EM / Resolution: 5.14 Å | |||||||||
 Authors Authors | Ito F / Alvarez-Cabrera AL / Zhou ZH / Chen XS | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural basis of HIV-1 Vif-mediated E3 ligase targeting of host APOBEC3H. Authors: Fumiaki Ito / Ana L Alvarez-Cabrera / Kyumin Kim / Z Hong Zhou / Xiaojiang S Chen /  Abstract: Human APOBEC3 (A3) cytidine deaminases are antiviral factors that are particularly potent against retroviruses. As a countermeasure, HIV-1 uses a viral infectivity factor (Vif) to target specific ...Human APOBEC3 (A3) cytidine deaminases are antiviral factors that are particularly potent against retroviruses. As a countermeasure, HIV-1 uses a viral infectivity factor (Vif) to target specific human A3s for proteasomal degradation. Vif recruits cellular transcription cofactor CBF-β and Cullin-5 (CUL5) RING E3 ubiquitin ligase to bind different A3s distinctively, but how this is accomplished remains unclear in the absence of the atomic structure of the complex. Here, we present the cryo-EM structures of HIV-1 Vif in complex with human A3H, CBF-β and components of CUL5 ubiquitin ligase (CUL5, ELOB, and ELOC). Vif nucleates the entire complex by directly binding four human proteins, A3H, CBF-β, CUL5, and ELOC. The structures reveal a large interface area between A3H and Vif, primarily mediated by an α-helical side of A3H and a five-stranded β-sheet of Vif. This A3H-Vif interface unveils the basis for sensitivity-modulating polymorphism of both proteins, including a previously reported gain-of-function mutation in Vif isolated from HIV/AIDS patients. Our structural and functional results provide insights into the remarkable interplay between HIV and humans and would inform development efforts for anti-HIV therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29490.map.gz emd_29490.map.gz | 483.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29490-v30.xml emd-29490-v30.xml emd-29490.xml emd-29490.xml | 21.7 KB 21.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29490_fsc.xml emd_29490_fsc.xml | 17 KB | Display |  FSC data file FSC data file |
| Images |  emd_29490.png emd_29490.png | 51.2 KB | ||
| Filedesc metadata |  emd-29490.cif.gz emd-29490.cif.gz | 6.2 KB | ||
| Others |  emd_29490_half_map_1.map.gz emd_29490_half_map_1.map.gz emd_29490_half_map_2.map.gz emd_29490_half_map_2.map.gz | 475.8 MB 475.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29490 http://ftp.pdbj.org/pub/emdb/structures/EMD-29490 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29490 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29490 | HTTPS FTP |
-Related structure data
| Related structure data |  8fviC  8fvjC C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29490.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29490.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.92 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_29490_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_29490_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Hetero-heptameric complex of HIV-1 Vif and human APOBEC3H, CBF-be...
| Entire | Name: Hetero-heptameric complex of HIV-1 Vif and human APOBEC3H, CBF-beta, ELOB, ELOC, CUL5, and RBX2 |
|---|---|
| Components |
|
-Supramolecule #1: Hetero-heptameric complex of HIV-1 Vif and human APOBEC3H, CBF-be...
| Supramolecule | Name: Hetero-heptameric complex of HIV-1 Vif and human APOBEC3H, CBF-beta, ELOB, ELOC, CUL5, and RBX2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Core-binding factor subunit beta
| Macromolecule | Name: Core-binding factor subunit beta / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MPRVVPDQRS KFENEEFFR K LSRECEIK YT GFRDRPH EER QARFQN ACRD GRSEI AFVAT GTNL SLQFFP ASW QGEQRQT PS REYVDLER E AGKVYLKAP MILNGVCVIW KGWIDLQRL D GMGCLEFD EE RAQQEDA LAQ QAFEEA RRRT REFED RD UniProtKB: Core-binding factor subunit beta |
-Macromolecule #2: Virion infectivity factor
| Macromolecule | Name: Virion infectivity factor / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 / Strain: pNL4-3 Human immunodeficiency virus 1 / Strain: pNL4-3 |
| Recombinant expression | Organism:  |
| Sequence | String: GPMENRWQVM IV WQVDRMR INT WKRLVK HHMY ISRKA KDWFY RHHY ESTHPK ISS EVHIPLG DA KLVITTYW G LHTGERDWH LGQGVSIEWR KKRYSTQVD P DLADQLIH LH YFDCFSE SAI RNTILG RIVS PRCEY QAGHN KVGS LQYLAL AAL ...String: GPMENRWQVM IV WQVDRMR INT WKRLVK HHMY ISRKA KDWFY RHHY ESTHPK ISS EVHIPLG DA KLVITTYW G LHTGERDWH LGQGVSIEWR KKRYSTQVD P DLADQLIH LH YFDCFSE SAI RNTILG RIVS PRCEY QAGHN KVGS LQYLAL AAL IKPKQIK PP LPSVRKLT E DRWNK UniProtKB: Virion infectivity factor |
-Macromolecule #3: Elongin-B
| Macromolecule | Name: Elongin-B / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MDVFLMIRRH KTTIFTDAK E SSTVFELK RI VEGILKR PPD EQRLYK DDQL LDDGK TLGEC GFTS QTARPQ APA TVGLAFR AD DTFEALCI E PFSSPPELP DV UniProtKB: Elongin-B |
-Macromolecule #4: Elongin-C
| Macromolecule | Name: Elongin-C / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MYVKLISSDG HEFIVKREH A LTSGTIKA ML SGPGQFA ENE TNEVNF REIP SHVLS KVCMY FTYK VRYTNS STE IPEFPIA PE IALELLMA A NFLDC UniProtKB: Elongin-C |
-Macromolecule #5: Cullin-5
| Macromolecule | Name: Cullin-5 / type: protein_or_peptide / ID: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: SLQFEDKWD FMRPIVLKLL RQESVTKQQ W FDLFSDVH AV CLWDDKG PAK IHQALK EDIL EFIKQ AQARV LSHQ DDTALL KAY IVEWRKF FT QCDILPKP F CQLEITLMG KQGSNKKSNV EDSIVRKLM L DTWNESIF SN IKNRLQD SAM KLVHAE RLGE ...String: SLQFEDKWD FMRPIVLKLL RQESVTKQQ W FDLFSDVH AV CLWDDKG PAK IHQALK EDIL EFIKQ AQARV LSHQ DDTALL KAY IVEWRKF FT QCDILPKP F CQLEITLMG KQGSNKKSNV EDSIVRKLM L DTWNESIF SN IKNRLQD SAM KLVHAE RLGE AFDSQ LVIGV RESY VNLCSN PED KLQIYRD NF EKAYLDST E RFYRTQAPS YLQQNGVQNY MKYADAKLK E EEKRALRY LE TRRECNS VEA LMECCV NALV TSFKE TILAE CQGM IKRNET EKL HLMFSLM DK VPNGIEPM L KDLEEHIIS AGLADMVAAA ETITTDSEK Y VEQLLTLF NR FSKLVKE AFQ DDPRFL TARD KAYKA VVNDA TIFK LELPLK QKG VGLKTQP ES KCPELLAN Y CDMLLRKTP LSKKLTSEEI EAKLKEVLL V LKYVQNKD VF MRYHKAH LTR RLILDI SADS EIEEN MVEWL REVG MPADYV NKL ARMFQDI KV SEDLNQAF K EMHKNNKLA LPADSVNIKI LNAGAWSRS S EKVFVSLP TE LEDLIPE VEE FYKKNH SGRK LHWHH LMSNG IITF KNEVGQ YDL EVTTFQL AV LFAWNQRP R EKISFENLK LATELPDAEL RRTLWSLVA F PKLKRQVL LY EPQVNSP KDF TEGTLF SVNQ EFSLI KNAKV QKRG KINLIG RLQ LTTERMR EE ENEGIVQL R ILRTQEAII QIMKMRKKIS NAQLQTELV E ILKNMFLP QK KMIKEQI EWL IEHKYI RRDE SDINT FIYMA UniProtKB: Cullin-5 |
-Macromolecule #6: RING-box protein 2
| Macromolecule | Name: RING-box protein 2 / type: protein_or_peptide / ID: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MADVEDGEET CALASHSGS S GSKSGGDK MF SLKKWNA VAM WSWDVE CDTC AICRV QVMDA CLRC QAENKQ EDC VVVWGEC NH SFHNCCMS L WVKQNNRCP LCQQDWVVQR IGK UniProtKB: RING-box protein 2 |
-Macromolecule #7: DNA dC->dU-editing enzyme APOBEC-3H
| Macromolecule | Name: DNA dC->dU-editing enzyme APOBEC-3H / type: protein_or_peptide / ID: 7 / Enantiomer: LEVO / EC number: single-stranded DNA cytosine deaminase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: GPGGSGGMAL LTAETFR LQ FNNKRRLR R PYYPRKALL CYQLTPQNGS TPTRGYFEN K KKCHAEIC FI NEIKSMG LDE TQCYQV TCYL TWSPC SSCAW ELVD FIQAHD HLN LRIFASR LY YHWCKPQQ D GLRLLCGSQ VPVEVMGFPE FADCWENFV D ...String: GPGGSGGMAL LTAETFR LQ FNNKRRLR R PYYPRKALL CYQLTPQNGS TPTRGYFEN K KKCHAEIC FI NEIKSMG LDE TQCYQV TCYL TWSPC SSCAW ELVD FIQAHD HLN LRIFASR LY YHWCKPQQ D GLRLLCGSQ VPVEVMGFPE FADCWENFV D HEKPLSFN PY KMLEELD KNS RAIKRR LDRI KS UniProtKB: DNA dC->dU-editing enzyme APOBEC-3H |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.15 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 12546 / Average exposure time: 8.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 150000 |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)