[English] 日本語
 Yorodumi
Yorodumi- EMDB-28630: CX3CR1 nucleosome and PU.1 complex containing disulfide bond mutations -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
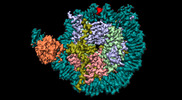
| Title | CX3CR1 nucleosome and PU.1 complex containing disulfide bond mutations | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | nucleosome / transcription factor / transcription / chromatin binding protein-DNA complex / TRANSCRIPTION-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of antifungal innate immune response / regulation of myeloid progenitor cell differentiation / positive regulation of myeloid dendritic cell chemotaxis / anatomical structure regression / pro-T cell differentiation / negative regulation of neutrophil degranulation / follicular B cell differentiation / myeloid leukocyte differentiation / positive regulation of microglial cell mediated cytotoxicity / germinal center B cell differentiation ...positive regulation of antifungal innate immune response / regulation of myeloid progenitor cell differentiation / positive regulation of myeloid dendritic cell chemotaxis / anatomical structure regression / pro-T cell differentiation / negative regulation of neutrophil degranulation / follicular B cell differentiation / myeloid leukocyte differentiation / positive regulation of microglial cell mediated cytotoxicity / germinal center B cell differentiation / lymphocyte differentiation / TRAIL-activated apoptotic signaling pathway / granulocyte differentiation / negative regulation of MHC class II biosynthetic process / endothelial to hematopoietic transition / negative regulation of adipose tissue development / apoptotic process involved in blood vessel morphogenesis / pericyte cell differentiation / lymphoid progenitor cell differentiation / immature B cell differentiation / myeloid dendritic cell differentiation / defense response to tumor cell / vasculature development / positive regulation of p38MAPK cascade / oncogene-induced cell senescence / negative regulation of non-canonical NF-kappaB signal transduction / negative regulation of protein localization to chromatin / positive regulation of B cell differentiation / STAT family protein binding / interleukin-6-mediated signaling pathway / NFAT protein binding / cellular response to ethanol / macrophage differentiation / somatic stem cell population maintenance / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / cis-regulatory region sequence-specific DNA binding / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / negative regulation of canonical NF-kappaB signal transduction / Deposition of new CENPA-containing nucleosomes at the centromere / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / telomere organization / Interleukin-7 signaling / Inhibition of DNA recombination at telomere / RNA Polymerase I Promoter Opening / transforming growth factor beta receptor signaling pathway / Meiotic synapsis / transcription initiation-coupled chromatin remodeling / protein sequestering activity / Assembly of the ORC complex at the origin of replication / osteoclast differentiation / lipopolysaccharide-mediated signaling pathway / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / SUMOylation of chromatin organization proteins / DNA methylation / epigenetic regulation of gene expression / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / innate immune response in mucosa / erythrocyte differentiation / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / HDACs deacetylate histones / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / Transcriptional regulation by small RNAs / regulation of erythrocyte differentiation / Formation of the beta-catenin:TCF transactivating complex / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / HDMs demethylate histones / G2/M DNA damage checkpoint / NoRC negatively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / B-WICH complex positively regulates rRNA expression / PKMTs methylate histone lysines / Meiotic recombination / Pre-NOTCH Transcription and Translation / positive regulation of miRNA transcription / DNA-binding transcription repressor activity, RNA polymerase II-specific / Metalloprotease DUBs / histone deacetylase binding / RMTs methylate histone arginines / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Transcriptional regulation of granulopoiesis / HCMV Early Events / antimicrobial humoral immune response mediated by antimicrobial peptide / structural constituent of chromatin / UCH proteinases Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.64 Å | |||||||||
 Authors Authors | Lian T / Guan R / Bai Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Structural mechanism of synergistic targeting of the CX3CR1 nucleosome by PU.1 and C/EBPα. Authors: Tengfei Lian / Ruifang Guan / Bing-Rui Zhou / Yawen Bai /  Abstract: Pioneer transcription factors are vital for cell fate changes. PU.1 and C/EBPα work together to regulate hematopoietic stem cell differentiation. However, how they recognize in vivo nucleosomal DNA ...Pioneer transcription factors are vital for cell fate changes. PU.1 and C/EBPα work together to regulate hematopoietic stem cell differentiation. However, how they recognize in vivo nucleosomal DNA targets remains elusive. Here we report the structures of the nucleosome containing the mouse genomic CX3CR1 enhancer DNA and its complexes with PU.1 alone and with both PU.1 and the C/EBPα DNA binding domain. Our structures reveal that PU.1 binds the DNA motif at the exit linker, shifting 17 bp of DNA into the core region through interactions with H2A, unwrapping ~20 bp of nucleosomal DNA. C/EBPα binding, aided by PU.1's repositioning, unwraps ~25 bp of entry DNA. The PU.1 Q218H mutation, linked to acute myeloid leukemia, disrupts PU.1-H2A interactions. PU.1 and C/EBPα jointly displace linker histone H1 and open the H1-condensed nucleosome array. Our study unveils how two pioneer factors can work cooperatively to open closed chromatin by altering DNA positioning in the nucleosome. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28630.map.gz emd_28630.map.gz | 49.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28630-v30.xml emd-28630-v30.xml emd-28630.xml emd-28630.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28630.png emd_28630.png | 84.6 KB | ||
| Filedesc metadata |  emd-28630.cif.gz emd-28630.cif.gz | 6.7 KB | ||
| Others |  emd_28630_half_map_1.map.gz emd_28630_half_map_1.map.gz emd_28630_half_map_2.map.gz emd_28630_half_map_2.map.gz | 49 MB 49 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28630 http://ftp.pdbj.org/pub/emdb/structures/EMD-28630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28630 | HTTPS FTP |
-Validation report
| Summary document |  emd_28630_validation.pdf.gz emd_28630_validation.pdf.gz | 977.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28630_full_validation.pdf.gz emd_28630_full_validation.pdf.gz | 976.9 KB | Display | |
| Data in XML |  emd_28630_validation.xml.gz emd_28630_validation.xml.gz | 12 KB | Display | |
| Data in CIF |  emd_28630_validation.cif.gz emd_28630_validation.cif.gz | 14 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28630 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28630 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28630 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28630 | HTTPS FTP |
-Related structure data
| Related structure data |  8eviMC  8evhC  8evjC  8sypC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28630.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28630.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.056 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_28630_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_28630_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : nucleosome PU.1 mutant complex
+Supramolecule #1: nucleosome PU.1 mutant complex
+Macromolecule #1: DNA (167-MER)
+Macromolecule #2: DNA (167-MER)
+Macromolecule #3: Histone H3.1
+Macromolecule #4: Histone H4
+Macromolecule #5: Histone H2A type 2-C
+Macromolecule #6: Histone H2B type 2-E
+Macromolecule #7: Histone H2A type 2-C
+Macromolecule #8: Single-chain variable fragment
+Macromolecule #9: Transcription factor PU.1
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.3 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 53.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.64 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 127327 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)