[English] 日本語
 Yorodumi
Yorodumi- EMDB-27701: Focused map (monomer A) for Arabidopsis SPY in complex with GDP-fucose -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Focused map (monomer A) for Arabidopsis SPY in complex with GDP-fucose | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | O-Fucosyltransferase / PLANT PROTEIN | |||||||||
| Biological species |  | |||||||||
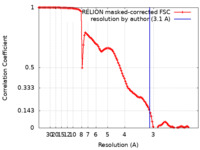
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Kumar S / Zhou Y / Dillard L / Borgnia MJ / Bartesaghi A / Zhou P | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structure and dynamics of the Arabidopsis O-fucosyltransferase SPINDLY. Authors: Shivesh Kumar / Yan Wang / Ye Zhou / Lucas Dillard / Fay-Wei Li / Carly A Sciandra / Ning Sui / Rodolfo Zentella / Emily Zahn / Jeffrey Shabanowitz / Donald F Hunt / Mario J Borgnia / ...Authors: Shivesh Kumar / Yan Wang / Ye Zhou / Lucas Dillard / Fay-Wei Li / Carly A Sciandra / Ning Sui / Rodolfo Zentella / Emily Zahn / Jeffrey Shabanowitz / Donald F Hunt / Mario J Borgnia / Alberto Bartesaghi / Tai-Ping Sun / Pei Zhou /  Abstract: SPINDLY (SPY) in Arabidopsis thaliana is a novel nucleocytoplasmic protein O-fucosyltransferase (POFUT), which regulates diverse developmental processes. Sequence analysis indicates that SPY is ...SPINDLY (SPY) in Arabidopsis thaliana is a novel nucleocytoplasmic protein O-fucosyltransferase (POFUT), which regulates diverse developmental processes. Sequence analysis indicates that SPY is distinct from ER-localized POFUTs and contains N-terminal tetratricopeptide repeats (TPRs) and a C-terminal catalytic domain resembling the O-linked-N-acetylglucosamine (GlcNAc) transferases (OGTs). However, the structural feature that determines the distinct enzymatic selectivity of SPY remains unknown. Here we report the cryo-electron microscopy (cryo-EM) structure of SPY and its complex with GDP-fucose, revealing distinct active-site features enabling GDP-fucose instead of UDP-GlcNAc binding. SPY forms an antiparallel dimer instead of the X-shaped dimer in human OGT, and its catalytic domain interconverts among multiple conformations. Analysis of mass spectrometry, co-IP, fucosylation activity, and cryo-EM data further demonstrates that the N-terminal disordered peptide in SPY contains trans auto-fucosylation sites and inhibits the POFUT activity, whereas TPRs 1-5 dynamically regulate SPY activity by interfering with protein substrate binding. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27701.map.gz emd_27701.map.gz | 118 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27701-v30.xml emd-27701-v30.xml emd-27701.xml emd-27701.xml | 17.3 KB 17.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27701_fsc.xml emd_27701_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_27701.png emd_27701.png | 49.3 KB | ||
| Masks |  emd_27701_msk_1.map emd_27701_msk_1.map | 125 MB |  Mask map Mask map | |
| Others |  emd_27701_half_map_1.map.gz emd_27701_half_map_1.map.gz emd_27701_half_map_2.map.gz emd_27701_half_map_2.map.gz | 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27701 http://ftp.pdbj.org/pub/emdb/structures/EMD-27701 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27701 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27701 | HTTPS FTP |
-Validation report
| Summary document |  emd_27701_validation.pdf.gz emd_27701_validation.pdf.gz | 759.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27701_full_validation.pdf.gz emd_27701_full_validation.pdf.gz | 759.1 KB | Display | |
| Data in XML |  emd_27701_validation.xml.gz emd_27701_validation.xml.gz | 18.1 KB | Display | |
| Data in CIF |  emd_27701_validation.cif.gz emd_27701_validation.cif.gz | 23.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27701 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27701 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27701 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27701 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27701.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27701.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27701_msk_1.map emd_27701_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27701_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_27701_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SPY GDP-fucose complex
| Entire | Name: SPY GDP-fucose complex |
|---|---|
| Components |
|
-Supramolecule #1: SPY GDP-fucose complex
| Supramolecule | Name: SPY GDP-fucose complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: SPY GDP-fucose complex
| Macromolecule | Name: SPY GDP-fucose complex / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MVGLEDDTER ERSPVVENGF SNGSRSSSSS AGVLSPSRKV TQGNDTLSYA NILRARNKFA DALALYEAML EKDSKNVEAH IGKGICLQTQ NKGNLAFDCF SEAIRLDPHN ACALTHCGIL HKEEGRLVEA AESYQKALMA DASYKPAAEC LAIVLTDLGT SLKLAGNTQE ...String: MVGLEDDTER ERSPVVENGF SNGSRSSSSS AGVLSPSRKV TQGNDTLSYA NILRARNKFA DALALYEAML EKDSKNVEAH IGKGICLQTQ NKGNLAFDCF SEAIRLDPHN ACALTHCGIL HKEEGRLVEA AESYQKALMA DASYKPAAEC LAIVLTDLGT SLKLAGNTQE GIQKYYEALK IDPHYAPAYY NLGVVYSEMM QYDNALSCYE KAALERPMYA EAYCNMGVIY KNRGDLEMAI TCYERCLAVS PNFEIAKNNM AIALTDLGTK VKLEGDVTQG VAYYKKALYY NWHYADAMYN LGVAYGEMLK FDMAIVFYEL AFHFNPHCAE ACNNLGVLYK DRDNLDKAVE CYQMALSIKP NFAQSLNNLG VVYTVQGKMD AAASMIEKAI LANPTYAEAF NNLGVLYRDA GNITMAIDAY EECLKIDPDS RNAGQNRLLA MNYINEGLDD KLFEAHRDWG WRFTRLHPQY TSWDNLKDPE RPITIGYISP DFFTHSVSYF IEAPLTHHDY TKYKVVVYSA VVKADAKTYR FRDKVLKKGG VWKDIYGIDE KKIASMVRED KIDILVELTG HTANNKLGTM ACRPAPVQVT WIGYPNTTGL PTVDYRITDS LADPPDTKQK QVEELVRLPD CFLCYTPSPE AGPVCPTPAL SNGFVTFGSF NNLAKITPKV LQVWARILCA VPNSRLVVKC KPFCCDSIRQ RFLTTLEQLG LESKRVDLLP LILFNHDHMQ AYSLMDISLD TFPYAGTTTT CESLYMGVPC VTMAGSVHAH NVGVSLLTKV GLGHLVAKNE DEYVQLSVDL ASDVTALSKL RMSLRDLMAG SPVCNGPSFA VGLESAYRNM WKKYCKGEVP SLRRMEMLQK EVHDDPLISK DLGPSRVSVT GEATPSLKAN GSAPVPSSLP TQSPQLSKRM DSTSGGSENL YFQGGSHHHH HHHHHHGGWS HPQFEK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Support film - Film thickness: 40 / Pretreatment - Type: PLASMA CLEANING | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 289.15 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Average electron dose: 52.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 81000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X