[English] 日本語
 Yorodumi
Yorodumi- EMDB-27278: Saccharomyces cerevisiae Ufd1/Npl4/Cdc48 complex bound to three u... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Saccharomyces cerevisiae Ufd1/Npl4/Cdc48 complex bound to three ubiquitin moieties and one unfolded ubiquitin in presence of SUMO-ubiquitin(K48polyUb)-mEOS and ATP, state 2 (uD) | |||||||||
 Map data Map data | composite map of the ubiquitin unfolded state 'uD' | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ATPASE / ATPASE COMPLEX / UBIQUITIN / SUMO / SMT3 / QUALITY CONTROL / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information: / RAS processing / Regulation of PTEN localization / : / UCH proteinases / : / Aggrephagy / Pexophagy / PINK1-PRKN Mediated Mitophagy / : ...: / RAS processing / Regulation of PTEN localization / : / UCH proteinases / : / Aggrephagy / Pexophagy / PINK1-PRKN Mediated Mitophagy / : / EGAD pathway / SCF complex disassembly in response to cadmium stress / mitotic DNA replication termination / Ovarian tumor domain proteases / Cdc48p-Npl4p-Vms1p AAA ATPase complex / Doa10p ubiquitin ligase complex / stress-induced homeostatically regulated protein degradation pathway / Peroxisomal protein import / : / protein localization to vacuole / endoplasmic reticulum membrane fusion / sister chromatid biorientation / Hrd1p ubiquitin ligase ERAD-L complex / DNA replication termination / RQC complex / : / Endosomal Sorting Complex Required For Transport (ESCRT) / mitochondria-associated ubiquitin-dependent protein catabolic process / positive regulation of mitochondrial fusion / cytoplasm protein quality control by the ubiquitin-proteasome system / HSF1 activation / protein-containing complex disassembly / : / protein transport to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway / nuclear protein quality control by the ubiquitin-proteasome system / endosome to plasma membrane protein transport / ribophagy / Translesion synthesis by REV1 / : / : / : / protein phosphatase regulator activity / piecemeal microautophagy of the nucleus / : / mating projection tip / : / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / Protein methylation / replisome / mitotic spindle disassembly / VCP-NPL4-UFD1 AAA ATPase complex / ribosome-associated ubiquitin-dependent protein catabolic process / vesicle-fusing ATPase / K48-linked polyubiquitin modification-dependent protein binding / nuclear outer membrane-endoplasmic reticulum membrane network / nonfunctional rRNA decay / Regulation of PTEN stability and activity / retrograde protein transport, ER to cytosol / CDK-mediated phosphorylation and removal of Cdc6 / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / KEAP1-NFE2L2 pathway / Neddylation / Formation of TC-NER Pre-Incision Complex / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / Orc1 removal from chromatin / MAPK6/MAPK4 signaling / SRP-dependent cotranslational protein targeting to membrane / Gap-filling DNA repair synthesis and ligation in TC-NER / GTP hydrolysis and joining of the 60S ribosomal subunit / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Antigen processing: Ubiquitination & Proteasome degradation / protein quality control for misfolded or incompletely synthesized proteins / L13a-mediated translational silencing of Ceruloplasmin expression / Dual incision in TC-NER / autophagosome maturation / Ub-specific processing proteases / polyubiquitin modification-dependent protein binding / mRNA transport / ATP metabolic process / ERAD pathway / Neutrophil degranulation / rescue of stalled cytosolic ribosome / macroautophagy / ubiquitin binding / positive regulation of protein localization to nucleus / modification-dependent protein catabolic process / protein tag activity / peroxisome / nuclear membrane / ubiquitin-dependent protein catabolic process / proteasome-mediated ubiquitin-dependent protein catabolic process / protein ubiquitination / ubiquitin protein ligase binding / endoplasmic reticulum membrane / perinuclear region of cytoplasm / ATP hydrolysis activity / mitochondrion / ATP binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
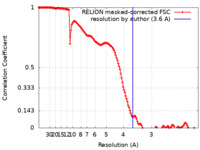
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Lee HG / Lima CD | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: SUMO enhances unfolding of SUMO-polyubiquitin-modified substrates by the Ufd1/Npl4/Cdc48 complex. Authors: Hyein G Lee / Abigail A Lemmon / Christopher D Lima /  Abstract: The Ufd1/Npl4/Cdc48 complex is a universal protein segregase that plays key roles in eukaryotic cellular processes. Its functions orchestrating the clearance or removal of polyubiquitylated targets ...The Ufd1/Npl4/Cdc48 complex is a universal protein segregase that plays key roles in eukaryotic cellular processes. Its functions orchestrating the clearance or removal of polyubiquitylated targets are established; however, prior studies suggest that the complex also targets substrates modified by the ubiquitin-like protein SUMO. Here, we show that interactions between Ufd1 and SUMO enhance unfolding of substrates modified by SUMO-polyubiquitin hybrid chains by the budding yeast Ufd1/Npl4/Cdc48 complex compared to substrates modified by polyubiquitin chains, a difference that is accentuated when the complex has a choice between these substrates. Incubating Ufd1/Npl4/Cdc48 with a substrate modified by a SUMO-polyubiquitin hybrid chain produced a series of single-particle cryo-EM structures that reveal features of interactions between Ufd1/Npl4/Cdc48 and ubiquitin prior to and during unfolding of ubiquitin. These results are consistent with cellular functions for SUMO and ubiquitin modifications and support a physical model wherein Ufd1/Npl4/Cdc48, SUMO, and ubiquitin conjugation pathways converge to promote clearance of proteins modified with SUMO and polyubiquitin. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27278.map.gz emd_27278.map.gz | 25.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27278-v30.xml emd-27278-v30.xml emd-27278.xml emd-27278.xml | 54 KB 54 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27278_fsc.xml emd_27278_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_27278.png emd_27278.png | 96 KB | ||
| Filedesc metadata |  emd-27278.cif.gz emd-27278.cif.gz | 8.3 KB | ||
| Others |  emd_27278_additional_1.map.gz emd_27278_additional_1.map.gz emd_27278_additional_10.map.gz emd_27278_additional_10.map.gz emd_27278_additional_11.map.gz emd_27278_additional_11.map.gz emd_27278_additional_12.map.gz emd_27278_additional_12.map.gz emd_27278_additional_13.map.gz emd_27278_additional_13.map.gz emd_27278_additional_2.map.gz emd_27278_additional_2.map.gz emd_27278_additional_3.map.gz emd_27278_additional_3.map.gz emd_27278_additional_4.map.gz emd_27278_additional_4.map.gz emd_27278_additional_5.map.gz emd_27278_additional_5.map.gz emd_27278_additional_6.map.gz emd_27278_additional_6.map.gz emd_27278_additional_7.map.gz emd_27278_additional_7.map.gz emd_27278_additional_8.map.gz emd_27278_additional_8.map.gz emd_27278_additional_9.map.gz emd_27278_additional_9.map.gz emd_27278_half_map_1.map.gz emd_27278_half_map_1.map.gz emd_27278_half_map_2.map.gz emd_27278_half_map_2.map.gz | 17.5 MB 9.9 MB 171.2 MB 171.2 MB 8 MB 13.5 MB 170.7 MB 170.7 MB 171.4 MB 171.2 MB 12.2 MB 171.2 MB 171.4 MB 171.5 MB 171.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27278 http://ftp.pdbj.org/pub/emdb/structures/EMD-27278 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27278 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27278 | HTTPS FTP |
-Related structure data
| Related structure data |  8dawMC  8darC  8dasC  8datC  8dauC  8davC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27278.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27278.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | composite map of the ubiquitin unfolded state 'uD' | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.064 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
+Additional map: post-processed overall refinement map of the ubiquitin unfolded...
+Additional map: post-processed focused refinement map of the Ufd1/Npl4 tower...
+Additional map: focused refinement half map 1 of Cdc48 of...
+Additional map: focused refinement half map 1 of the Ufd1/Npl4...
+Additional map: post-processed focused refinement map of upper Ufd1/Npl4 tower...
+Additional map: post-processed focused refinement map of Cdc48 of the...
+Additional map: focused refinement half map 1 of upper Ufd1/Npl4...
+Additional map: focused refinement half map 2 of upper Ufd1/Npl4...
+Additional map: focused refinement half map 2 of the D1/D2...
+Additional map: focused refinement half map 2 of the Ufd1/Npl4...
+Additional map: post-processed focused refinement map of the D1/D2 ATPase...
+Additional map: focused refinement half map 2 of Cdc48 of...
+Additional map: focused refinement half map 1 of the D1/D2...
+Half map: overall refinement half map 2 of the ubiquitin unfolded state 'uD'
+Half map: overall refinement half map 1 of the ubiquitin unfolded state 'uD'
- Sample components
Sample components
+Entire : Saccharomyces cerevisiae Ufd1/Npl4/Cdc48 complex bound to three u...
+Supramolecule #1: Saccharomyces cerevisiae Ufd1/Npl4/Cdc48 complex bound to three u...
+Supramolecule #2: Saccharomyces cerevisiae Ufd1/Npl4/Cdc48 complex bound to three u...
+Supramolecule #3: Cdc48 hexamer
+Supramolecule #4: Three folded ubiqutin moieties and one unfolded ubiquitin bound t...
+Supramolecule #5: D1/D2 ATPase domains of the Cdc48 hexamer
+Supramolecule #6: Three folded Ubiqutin moieties bound to Npl4
+Macromolecule #1: Cell division control protein 48
+Macromolecule #2: Nuclear protein localization protein 4
+Macromolecule #3: Ubiquitin fusion degradation protein 1
+Macromolecule #4: Ubiquitin
+Macromolecule #5: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #6: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #7: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM HEPES pH 8.0, 150 mM NaCl, 0.1 mM TCEP, 1 mM MgCl2, 5 mM ATP. Added 0.05% CHAPSO before vitrification. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV / Details: 8 s wait, 4 s blot before plunging. |
| Details | Ufd1/Npl4/Cdc48 was pre-incubated with SUMO-ubiquitin(K48polyUb)-mEOS and ATP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 70.577 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



























 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)