[English] 日本語
 Yorodumi
Yorodumi- EMDB-2716: Dynactin 3D structure: Implications for assembly and dynein binding -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2716 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dynactin 3D structure: Implications for assembly and dynein binding | |||||||||
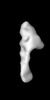
 Map data Map data | Random conical tilt 3D reconstruction of negatively stained dynactin complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | dynein / microtubule / actin / single particle analysis | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 34.0 Å | |||||||||
 Authors Authors | Imai H / Narita A / Maeda Y / Schroer TA | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2014 Journal: J Mol Biol / Year: 2014Title: Dynactin 3D structure: implications for assembly and dynein binding. Authors: Hiroshi Imai / Akihiro Narita / Yuichiro Maéda / Trina A Schroer /   Abstract: The multisubunit protein complex, dynactin, is an essential component of the cytoplasmic dynein motor. High-resolution structural work on dynactin and the dynein/dynactin supercomplex has been ...The multisubunit protein complex, dynactin, is an essential component of the cytoplasmic dynein motor. High-resolution structural work on dynactin and the dynein/dynactin supercomplex has been limited to small subunits and recombinant fragments that do not report fully on either ≈1MDa assembly. In the present study, we used negative-stain electron microscopy and image analysis based on random conical tilt reconstruction to obtain a three-dimensional (3D) structure of native vertebrate dynactin. The 35-nm-long dynactin molecule has a V-shaped shoulder at one end and a flattened tip at the other end, both offset relative to the long axis of the actin-related protein (Arp) backbone. The shoulder projects dramatically away from the Arp filament core in a way that cannot be appreciated in two-dimensional images, which has implications for the mechanism of dynein binding. The 3D structure allows the helical parameters of the entire Arp filament core, which includes the actin capping protein, CP, to be determined for the first time. This structure exhibits near identity to F-actin and can be well fitted into the dynactin envelope. Molecular fitting of modeled CP-Arp polymers into the envelope shows that the filament contains between 7 and 9 Arp protomers and is capped at both ends. In the 7 Arp model, which agrees best with measured Arp stoichiometry and other structural information, actin capping protein (CP) is not present at the distal tip of the structure, unlike what is seen in the other models. The 3D structure suggests a mechanism for dynactin assembly and length specification. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2716.map.gz emd_2716.map.gz | 3.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2716-v30.xml emd-2716-v30.xml emd-2716.xml emd-2716.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2716-EMDBdynactin500x500pixelA.tif EMD-2716-EMDBdynactin500x500pixelA.tif | 43.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2716 http://ftp.pdbj.org/pub/emdb/structures/EMD-2716 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2716 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2716 | HTTPS FTP |
-Validation report
| Summary document |  emd_2716_validation.pdf.gz emd_2716_validation.pdf.gz | 164.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2716_full_validation.pdf.gz emd_2716_full_validation.pdf.gz | 164 KB | Display | |
| Data in XML |  emd_2716_validation.xml.gz emd_2716_validation.xml.gz | 3.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2716 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2716 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2716 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2716 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2716.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2716.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Random conical tilt 3D reconstruction of negatively stained dynactin complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : The dynactin complex
+Supramolecule #1000: The dynactin complex
+Macromolecule #1: p150Glued
+Macromolecule #2: p50/dynamitin
+Macromolecule #3: p24
+Macromolecule #4: Arp1
+Macromolecule #5: Arp11
+Macromolecule #6: actin
+Macromolecule #7: p25
+Macromolecule #8: p27
+Macromolecule #9: p62
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.015 mg/mL |
|---|---|
| Buffer | pH: 7.1 Details: 35 mM Pipes-KOH, 5 mM MgSO4, 1 mM EGTA, and 0.5 mM EDTA, ~30 mM KCl |
| Staining | Type: NEGATIVE Details: Dynactin complex was negatively stained with 1% uranyl acetate on glow-discharged carbon grids. |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010HC |
|---|---|
| Date | Jan 19, 2008 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: OTHER / Average electron dose: 18 e/Å2 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 100 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 20000 |
| Sample stage | Specimen holder: room temperature, high tilt holder / Specimen holder model: JEOL / Tilt angle max: 60 |
- Image processing
Image processing
| Details | Random conical tilt reconstruction |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 34.0 Å / Resolution method: OTHER / Software - Name: Eos, SPIDER Details: Images of the dynactin structure were divided into two groups and reconstructed into two 3D maps that were compared by Fourier Shell Correlation (Van Heel, 1987). Number images used: 3637 |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)