[English] 日本語
 Yorodumi
Yorodumi- EMDB-26852: Cryo-EM structure of the mTORC1-TFEB-Rag-Ragulator complex with C... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the mTORC1-TFEB-Rag-Ragulator complex with C2 symmetry | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Cui Z / Hurley J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Structure of the lysosomal mTORC1-TFEB-Rag-Ragulator megacomplex. Authors: Zhicheng Cui / Gennaro Napolitano / Mariana E G de Araujo / Alessandra Esposito / Jlenia Monfregola / Lukas A Huber / Andrea Ballabio / James H Hurley /    Abstract: The transcription factor TFEB is a master regulator of lysosomal biogenesis and autophagy. The phosphorylation of TFEB by the mechanistic target of rapamycin complex 1 (mTORC1) is unique in its ...The transcription factor TFEB is a master regulator of lysosomal biogenesis and autophagy. The phosphorylation of TFEB by the mechanistic target of rapamycin complex 1 (mTORC1) is unique in its mTORC1 substrate recruitment mechanism, which is strictly dependent on the amino acid-mediated activation of the RagC GTPase activating protein FLCN. TFEB lacks the TOR signalling motif responsible for the recruitment of other mTORC1 substrates. We used cryogenic-electron microscopy to determine the structure of TFEB as presented to mTORC1 for phosphorylation, which we refer to as the 'megacomplex'. Two full Rag-Ragulator complexes present each molecule of TFEB to the mTOR active site. One Rag-Ragulator complex is bound to Raptor in the canonical mode seen previously in the absence of TFEB. A second Rag-Ragulator complex (non-canonical) docks onto the first through a RagC GDP-dependent contact with the second Ragulator complex. The non-canonical Rag dimer binds the first helix of TFEB with a RagC-dependent aspartate clamp in the cleft between the Rag G domains. In cellulo mutation of the clamp drives TFEB constitutively into the nucleus while having no effect on mTORC1 localization. The remainder of the 108-amino acid TFEB docking domain winds around Raptor and then back to RagA. The double use of RagC GDP contacts in both Rag dimers explains the strong dependence of TFEB phosphorylation on FLCN and the RagC GDP state. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26852.map.gz emd_26852.map.gz | 732.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26852-v30.xml emd-26852-v30.xml emd-26852.xml emd-26852.xml | 12.6 KB 12.6 KB | Display Display |  EMDB header EMDB header |
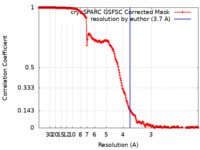
| FSC (resolution estimation) |  emd_26852_fsc.xml emd_26852_fsc.xml | 19.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_26852.png emd_26852.png | 89.8 KB | ||
| Others |  emd_26852_half_map_1.map.gz emd_26852_half_map_1.map.gz emd_26852_half_map_2.map.gz emd_26852_half_map_2.map.gz | 720.1 MB 720.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26852 http://ftp.pdbj.org/pub/emdb/structures/EMD-26852 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26852 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26852 | HTTPS FTP |
-Validation report
| Summary document |  emd_26852_validation.pdf.gz emd_26852_validation.pdf.gz | 949.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_26852_full_validation.pdf.gz emd_26852_full_validation.pdf.gz | 948.8 KB | Display | |
| Data in XML |  emd_26852_validation.xml.gz emd_26852_validation.xml.gz | 29.3 KB | Display | |
| Data in CIF |  emd_26852_validation.cif.gz emd_26852_validation.cif.gz | 38.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26852 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26852 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26852 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26852 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26852.map.gz / Format: CCP4 / Size: 775.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26852.map.gz / Format: CCP4 / Size: 775.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_26852_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_26852_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The mTORC1-TFEB-Rag-Ragulator complex
| Entire | Name: The mTORC1-TFEB-Rag-Ragulator complex |
|---|---|
| Components |
|
-Supramolecule #1: The mTORC1-TFEB-Rag-Ragulator complex
| Supramolecule | Name: The mTORC1-TFEB-Rag-Ragulator complex / type: complex / ID: 1 / Chimera: Yes / Parent: 0 |
|---|
-Supramolecule #2: mTORC1-TFEB-Rag
| Supramolecule | Name: mTORC1-TFEB-Rag / type: complex / ID: 2 / Chimera: Yes / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Ragulator complex
| Supramolecule | Name: Ragulator complex / type: complex / ID: 3 / Chimera: Yes / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)