+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | EcMscK G924S mutant in a closed conformation | |||||||||
 Map data Map data | EcMscK G924S mutant in a closed conformation | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRANSPORT PROTEIN / MEMBRANE PROTEIN / MECHANOSENSATION / ION CHANNEL | |||||||||
| Function / homology |  Function and homology information Function and homology informationintracellular water homeostasis / response to potassium ion / mechanosensitive monoatomic ion channel activity / potassium ion transport / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.84 Å | |||||||||
 Authors Authors | Mount JW / Yuan P | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural basis for mechanotransduction in a potassium-dependent mechanosensitive ion channel. Authors: Jonathan Mount / Grigory Maksaev / Brock T Summers / James A J Fitzpatrick / Peng Yuan /  Abstract: Mechanosensitive channels of small conductance, found in many living organisms, open under elevated membrane tension and thus play crucial roles in biological response to mechanical stress. Amongst ...Mechanosensitive channels of small conductance, found in many living organisms, open under elevated membrane tension and thus play crucial roles in biological response to mechanical stress. Amongst these channels, MscK is unique in that its activation also requires external potassium ions. To better understand this dual gating mechanism by force and ligand, we elucidate distinct structures of MscK along the gating cycle using cryo-electron microscopy. The heptameric channel comprises three layers: a cytoplasmic domain, a periplasmic gating ring, and a markedly curved transmembrane domain that flattens and expands upon channel opening, which is accompanied by dilation of the periplasmic ring. Furthermore, our results support a potentially unifying mechanotransduction mechanism in ion channels depicted as flattening and expansion of the transmembrane domain. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26823.map.gz emd_26823.map.gz | 211.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26823-v30.xml emd-26823-v30.xml emd-26823.xml emd-26823.xml | 17.8 KB 17.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26823_fsc.xml emd_26823_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_26823.png emd_26823.png | 162 KB | ||
| Filedesc metadata |  emd-26823.cif.gz emd-26823.cif.gz | 6.7 KB | ||
| Others |  emd_26823_half_map_1.map.gz emd_26823_half_map_1.map.gz emd_26823_half_map_2.map.gz emd_26823_half_map_2.map.gz | 226.1 MB 226.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26823 http://ftp.pdbj.org/pub/emdb/structures/EMD-26823 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26823 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26823 | HTTPS FTP |
-Related structure data
| Related structure data |  7uw5MC  7ux1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26823.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26823.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EcMscK G924S mutant in a closed conformation | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.94 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: EcMscK G924S mutant in a closed conformation, half map A
| File | emd_26823_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EcMscK G924S mutant in a closed conformation, half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
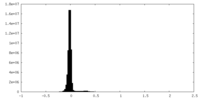
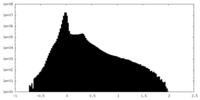
| Density Histograms |
-Half map: EcMscK G924S mutant in a closed conformation, half map B
| File | emd_26823_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EcMscK G924S mutant in a closed conformation, half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
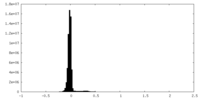
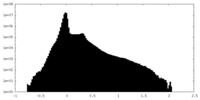
| Density Histograms |
- Sample components
Sample components
-Entire : E.coli MscK
| Entire | Name: E.coli MscK |
|---|---|
| Components |
|
-Supramolecule #1: E.coli MscK
| Supramolecule | Name: E.coli MscK / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Homomeric Heptamer |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Mechanosensitive channel MscK
| Macromolecule | Name: Mechanosensitive channel MscK / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 127.378789 KDa |
| Recombinant expression | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
| Sequence | String: MTMFQYYKRS RHFVFSAFIA FVFVLLCQNT AFARASSNGD LPTKADLQAQ LDSLNKQKDL SAQDKLVQQD LTDTLATLDK IDRIKEETV QLRQKVAEAP EKMRQATAAL TALSDVDNDE ETRKILSTLS LRQLETRVAQ ALDDLQNAQN DLASYNSQLV S LQTQPERV ...String: MTMFQYYKRS RHFVFSAFIA FVFVLLCQNT AFARASSNGD LPTKADLQAQ LDSLNKQKDL SAQDKLVQQD LTDTLATLDK IDRIKEETV QLRQKVAEAP EKMRQATAAL TALSDVDNDE ETRKILSTLS LRQLETRVAQ ALDDLQNAQN DLASYNSQLV S LQTQPERV QNAMYNASQQ LQQIRSRLDG TDVGETALRP SQKVLMQAQQ ALLNAEIDQQ RKSLEGNTVL QDTLQKQRDY VT ANSARLE HQLQLLQEAV NSKRLTLTEK TAQEAVSPDE AARIQANPLV KQELEINQQL SQRLITATEN GNQLMQQNIK VKN WLERAL QSERNIKEQI AVLKGSLLLS RILYQQQQTL PSADELENMT NRIADLRLEQ FEVNQQRDAL FQSDAFVNKL EEGH TNEVN SEVHDALLQV VDMRRELLDQ LNKQLGNQLM MAINLQINQQ QLMSVSKNLK SILTQQIFWV NSNRPMDWDW IKAFP QSLK DEFKSMKITV NWQKAWPAVF IAFLAGLPLL LIAGLIHWRL GWLKAYQQKL ASAVGSLRND SQLNTPKAIL IDLIRA LPV CLIILAVGLI LLTMQLNISE LLWSFSKKLA IFWLVFGLCW KVLEKNGVAV RHFGMPEQQT SHWRRQIVRI SLALLPI HF WSVVAELSPL HLMDDVLGQA MIFFNLLLIA FLVWPMCRES WRDKESHTMR LVTITVLSII PIALMVLTAT GYFYTTLR L AGRWIETVYL VIIWNLLYQT VLRGLSVAAR RIAWRRALAR RQNLVKEGAE GAEPPEEPTI ALEQVNQQTL RITMLLMFA LFGVMFWAIW SDLITVFSYL DSITLWHYNG TEAGAAVVKN VTMGSLLFAI IASMVAWALI RNLPGLLEVL VLSRLNMRQG ASYAITTIL NYIIIAVGAM TVFGSLGVSW DKLQWLAAAL SVGLSFGLQE IFGNFVSGLI ILFERPVRIG DTVTIGSFSG T VSKIRIRA TTITDFDRKE VIIPNKAFVT ERLINWSLTD TTTRLVIRLG VAYGSDLEKV RKVLLKAATE HPRVMHEPMP EV FFTAFGA STLDHELRLY VRELRDRSRT VDELNRTIDQ LCRENDINIA FNQLEVHLHN EKGDEVTEVK RDYKGDDPTP AVG UniProtKB: Mechanosensitive channel MscK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Micrographs were pooled from protein purified in the presence of either 150mM NaCl or 150mM KCl, respectively. Buffers were prepared fresh, degassed, and filtered through a 0.45 um Durapore PVDF membrane | |||||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | The specimen was homogeneous and monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Detector mode: COUNTING / Average electron dose: 46.16 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 150000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)