[English] 日本語
 Yorodumi
Yorodumi- EMDB-26756: C1 symmetric cryoEM structure of Azotobacter vinelandii MoFeP und... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | C1 symmetric cryoEM structure of Azotobacter vinelandii MoFeP under non-turnover conditions | |||||||||||||||
 Map data Map data | sharpened map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | nitrogenase / MoFeP / nitrogen fixation / OXIDOREDUCTASE | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationmolybdenum-iron nitrogenase complex / nitrogenase / nitrogenase activity / iron-sulfur cluster binding / ATP binding / metal ion binding Similarity search - Function | |||||||||||||||
| Biological species |  Azotobacter vinelandii DJ (bacteria) Azotobacter vinelandii DJ (bacteria) | |||||||||||||||
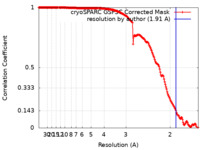
| Method | single particle reconstruction / cryo EM / Resolution: 1.91 Å | |||||||||||||||
 Authors Authors | Rutledge HL / Cook B / Tezcan FA / Herzik MA | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Structures of the nitrogenase complex prepared under catalytic turnover conditions. Authors: Hannah L Rutledge / Brian D Cook / Hoang P M Nguyen / Mark A Herzik / F Akif Tezcan /  Abstract: The enzyme nitrogenase couples adenosine triphosphate (ATP) hydrolysis to the multielectron reduction of atmospheric dinitrogen into ammonia. Despite extensive research, the mechanistic details of ...The enzyme nitrogenase couples adenosine triphosphate (ATP) hydrolysis to the multielectron reduction of atmospheric dinitrogen into ammonia. Despite extensive research, the mechanistic details of ATP-dependent energy transduction and dinitrogen reduction by nitrogenase are not well understood, requiring new strategies to monitor its structural dynamics during catalytic action. Here, we report cryo-electron microscopy structures of the nitrogenase complex prepared under enzymatic turnover conditions. We observe that asymmetry governs all aspects of the nitrogenase mechanism, including ATP hydrolysis, protein-protein interactions, and catalysis. Conformational changes near the catalytic iron-molybdenum cofactor are correlated with the nucleotide-hydrolysis state of the enzyme. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26756.map.gz emd_26756.map.gz | 484.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26756-v30.xml emd-26756-v30.xml emd-26756.xml emd-26756.xml | 24.7 KB 24.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26756_fsc.xml emd_26756_fsc.xml | 16.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_26756.png emd_26756.png | 121.9 KB | ||
| Masks |  emd_26756_msk_1.map emd_26756_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-26756.cif.gz emd-26756.cif.gz | 6.9 KB | ||
| Others |  emd_26756_additional_1.map.gz emd_26756_additional_1.map.gz emd_26756_half_map_1.map.gz emd_26756_half_map_1.map.gz emd_26756_half_map_2.map.gz emd_26756_half_map_2.map.gz | 258.4 MB 474.7 MB 474.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26756 http://ftp.pdbj.org/pub/emdb/structures/EMD-26756 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26756 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26756 | HTTPS FTP |
-Related structure data
| Related structure data |  7ut6MC  7ut7C  7ut8C  7ut9C  7utaC  8dpnC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26756.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26756.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.815 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_26756_msk_1.map emd_26756_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
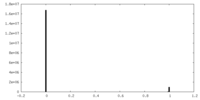
| Density Histograms |
-Additional map: unsharpened map
| File | emd_26756_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
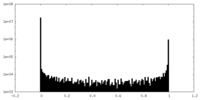
| Density Histograms |
-Half map: gold-standard half map B
| File | emd_26756_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | gold-standard half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
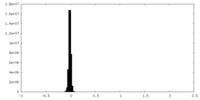
| Density Histograms |
-Half map: gold-standard half map A
| File | emd_26756_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | gold-standard half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
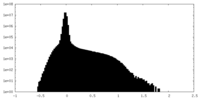
| Density Histograms |
- Sample components
Sample components
-Entire : Heterotetrameric MoFeP from Azotobacter vinelandii
| Entire | Name: Heterotetrameric MoFeP from Azotobacter vinelandii |
|---|---|
| Components |
|
-Supramolecule #1: Heterotetrameric MoFeP from Azotobacter vinelandii
| Supramolecule | Name: Heterotetrameric MoFeP from Azotobacter vinelandii / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: Wild-type MoFeP was purified from the native organism, Azotobacter vinelandii |
|---|---|
| Source (natural) | Organism:  Azotobacter vinelandii DJ (bacteria) Azotobacter vinelandii DJ (bacteria) |
| Molecular weight | Theoretical: 233.21 KDa |
-Macromolecule #1: Nitrogenase molybdenum-iron protein alpha chain
| Macromolecule | Name: Nitrogenase molybdenum-iron protein alpha chain / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: nitrogenase |
|---|---|
| Source (natural) | Organism:  Azotobacter vinelandii DJ (bacteria) Azotobacter vinelandii DJ (bacteria) |
| Molecular weight | Theoretical: 55.363043 KDa |
| Sequence | String: MTGMSREEVE SLIQEVLEVY PEKARKDRNK HLAVNDPAVT QSKKCIISNK KSQPGLMTIR GCAYAGSKGV VWGPIKDMIH ISHGPVGCG QYSRAGRRNY YIGTTGVNAF VTMNFTSDFQ EKDIVFGGDK KLAKLIDEVE TLFPLNKGIS VQSECPIGLI G DDIESVSK ...String: MTGMSREEVE SLIQEVLEVY PEKARKDRNK HLAVNDPAVT QSKKCIISNK KSQPGLMTIR GCAYAGSKGV VWGPIKDMIH ISHGPVGCG QYSRAGRRNY YIGTTGVNAF VTMNFTSDFQ EKDIVFGGDK KLAKLIDEVE TLFPLNKGIS VQSECPIGLI G DDIESVSK VKGAELSKTI VPVRCEGFRG VSQSLGHHIA NDAVRDWVLG KRDEDTTFAS TPYDVAIIGD YNIGGDAWSS RI LLEEMGL RCVAQWSGDG SISEIELTPK VKLNLVHCYR SMNYISRHME EKYGIPWMEY NFFGPTKTIE SLRAIAAKFD ESI QKKCEE VIAKYKPEWE AVVAKYRPRL EGKRVMLYIG GLRPRHVIGA YEDLGMEVVG TGYEFAHNDD YDRTMKEMGD STLL YDDVT GYEFEEFVKR IKPDLIGSGI KEKFIFQKMG IPFREMHSWD YSGPYHGFDG FAIFARDMDM TLNNPCWKKL QAPWE ASEG AEKVAASA UniProtKB: Nitrogenase molybdenum-iron protein alpha chain |
-Macromolecule #2: Nitrogenase molybdenum-iron protein beta chain
| Macromolecule | Name: Nitrogenase molybdenum-iron protein beta chain / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: nitrogenase |
|---|---|
| Source (natural) | Organism:  Azotobacter vinelandii DJ (bacteria) Azotobacter vinelandii DJ (bacteria) |
| Molecular weight | Theoretical: 59.535879 KDa |
| Sequence | String: MSQQVDKIKA SYPLFLDQDY KDMLAKKRDG FEEKYPQDKI DEVFQWTTTK EYQELNFQRE ALTVNPAKAC QPLGAVLCAL GFEKTMPYV HGSQGCVAYF RSYFNRHFRE PVSCVSDSMT EDAAVFGGQQ NMKDGLQNCK ATYKPDMIAV STTCMAEVIG D DLNAFINN ...String: MSQQVDKIKA SYPLFLDQDY KDMLAKKRDG FEEKYPQDKI DEVFQWTTTK EYQELNFQRE ALTVNPAKAC QPLGAVLCAL GFEKTMPYV HGSQGCVAYF RSYFNRHFRE PVSCVSDSMT EDAAVFGGQQ NMKDGLQNCK ATYKPDMIAV STTCMAEVIG D DLNAFINN SKKEGFIPDE FPVPFAHTPS FVGSHVTGWD NMFEGIARYF TLKSMDDKVV GSNKKINIVP GFETYLGNFR VI KRMLSEM GVGYSLLSDP EEVLDTPADG QFRMYAGGTT QEEMKDAPNA LNTVLLQPWH LEKTKKFVEG TWKHEVPKLN IPM GLDWTD EFLMKVSEIS GQPIPASLTK ERGRLVDMMT DSHTWLHGKR FALWGDPDFV MGLVKFLLEL GCEPVHILCH NGNK RWKKA VDAILAASPY GKNATVYIGK DLWHLRSLVF TDKPDFMIGN SYGKFIQRDT LHKGKEFEVP LIRIGFPIFD RHHLH RSTT LGYEGAMQIL TTLVNSILER LDEETRGMQA TDYNHDLVR UniProtKB: Nitrogenase molybdenum-iron protein beta chain |
-Macromolecule #3: 3-HYDROXY-3-CARBOXY-ADIPIC ACID
| Macromolecule | Name: 3-HYDROXY-3-CARBOXY-ADIPIC ACID / type: ligand / ID: 3 / Number of copies: 2 / Formula: HCA |
|---|---|
| Molecular weight | Theoretical: 206.15 Da |
| Chemical component information | 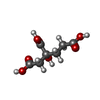 ChemComp-HCA: |
-Macromolecule #4: iron-sulfur-molybdenum cluster with interstitial carbon
| Macromolecule | Name: iron-sulfur-molybdenum cluster with interstitial carbon type: ligand / ID: 4 / Number of copies: 2 / Formula: ICS |
|---|---|
| Molecular weight | Theoretical: 787.451 Da |
| Chemical component information | 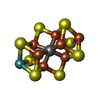 ChemComp-ICE: |
-Macromolecule #5: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #6: FE(8)-S(7) CLUSTER
| Macromolecule | Name: FE(8)-S(7) CLUSTER / type: ligand / ID: 6 / Number of copies: 2 / Formula: CLF |
|---|---|
| Molecular weight | Theoretical: 671.215 Da |
| Chemical component information |  ChemComp-CLF: |
-Macromolecule #7: water
| Macromolecule | Name: water / type: ligand / ID: 7 / Number of copies: 415 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.44 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Solutions were prepared and filtered immediately prior to the experiment. | |||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING | |||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER Details: Custom manual plunger. Greater than 95% humidity.. | |||||||||
| Details | 1.44 mg/mL MoFeP |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Temperature | Min: 93.0 K / Max: 123.0 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Number real images: 3773 / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)