+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of D-site Rac1-bound WAVE Regulatory Complex | |||||||||
 Map data Map data | WRC230VCA-Rac1 complex sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | actin regulator / GTPase binding protein / cytoskeletal regulator / CELL INVASION | |||||||||
| Function / homology |  Function and homology information Function and homology informationzygotic determination of anterior/posterior axis, embryo / dendritic transport of mitochondrion / SCAR complex / notochord morphogenesis / mesodermal cell migration / positive regulation of neurotrophin TRK receptor signaling pathway / lamellipodium morphogenesis / basal protein localization / paraxial mesoderm morphogenesis / positive regulation of Arp2/3 complex-mediated actin nucleation ...zygotic determination of anterior/posterior axis, embryo / dendritic transport of mitochondrion / SCAR complex / notochord morphogenesis / mesodermal cell migration / positive regulation of neurotrophin TRK receptor signaling pathway / lamellipodium morphogenesis / basal protein localization / paraxial mesoderm morphogenesis / positive regulation of Arp2/3 complex-mediated actin nucleation / embryonic body morphogenesis / establishment or maintenance of actin cytoskeleton polarity / cell migration involved in gastrulation / regulation of actin polymerization or depolymerization / Arp2/3 complex binding / dendrite extension / embryonic olfactory bulb interneuron precursor migration / anatomical structure arrangement / regulation of ERK5 cascade / angiotensin-activated signaling pathway involved in heart process / cerebral cortex GABAergic interneuron development / regulation of respiratory burst / positive regulation of ovarian follicle development / regulation of translation at postsynapse, modulating synaptic transmission / auditory receptor cell morphogenesis / cerebral cortex radially oriented cell migration / erythrocyte enucleation / regulation of neutrophil migration / negative regulation of interleukin-23 production / Activated NTRK2 signals through CDK5 / localization within membrane / modification of postsynaptic actin cytoskeleton / kinocilium / regulation of cell adhesion involved in heart morphogenesis / interneuron migration / ruffle assembly / embryonic foregut morphogenesis / NTRK2 activates RAC1 / filopodium tip / NADPH oxidase complex / cochlea morphogenesis / Inactivation of CDC42 and RAC1 / engulfment of apoptotic cell / regulation of hydrogen peroxide metabolic process / regulation of neuron maturation / respiratory burst / WNT5:FZD7-mediated leishmania damping / regulation of actin filament polymerization / SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion / cortical cytoskeleton organization / regulation of modification of postsynaptic actin cytoskeleton / positive regulation of skeletal muscle acetylcholine-gated channel clustering / endoderm development / ruffle organization / epithelial cell morphogenesis / midbrain dopaminergic neuron differentiation / positive regulation of bicellular tight junction assembly / GTP-dependent protein binding / RNA 7-methylguanosine cap binding / cell projection assembly / thioesterase binding / regulation of lamellipodium assembly / negative regulation of fibroblast migration / regulation of neuron migration / regulation of stress fiber assembly / axon extension / RHO GTPases activate CIT / embryonic heart tube development / cell-cell junction organization / motor neuron axon guidance / Nef and signal transduction / PCP/CE pathway / Activation of RAC1 / hepatocyte growth factor receptor signaling pathway / sphingosine-1-phosphate receptor signaling pathway / RHO GTPases activate KTN1 / regulation of nitric oxide biosynthetic process / quinolinate biosynthetic process / DCC mediated attractive signaling / MET activates RAP1 and RAC1 / cortical actin cytoskeleton organization / Azathioprine ADME / Sema4D mediated inhibition of cell attachment and migration / hyperosmotic response / CD28 dependent Vav1 pathway / Ephrin signaling / positive regulation of neutrophil chemotaxis / apical protein localization / Wnt signaling pathway, planar cell polarity pathway / superoxide anion generation / regulation of receptor signaling pathway via JAK-STAT / lamellipodium assembly / NRAGE signals death through JNK / positive regulation of ruffle assembly / dendrite morphogenesis / small GTPase-mediated signal transduction / Rho GDP-dissociation inhibitor binding / Activation of RAC1 downstream of NMDARs / protein kinase A regulatory subunit binding / positive regulation of Rho protein signal transduction Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
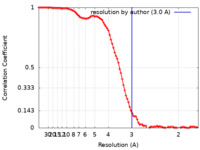
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Ding B / Yang S / Chen B / Chowdhury S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structures reveal a key mechanism of WAVE regulatory complex activation by Rac1 GTPase. Authors: Bojian Ding / Sheng Yang / Matthias Schaks / Yijun Liu / Abbigale J Brown / Klemens Rottner / Saikat Chowdhury / Baoyu Chen /    Abstract: The Rho-family GTPase Rac1 activates the WAVE regulatory complex (WRC) to drive Arp2/3 complex-mediated actin polymerization in many essential processes. Rac1 binds to WRC at two distinct sites-the ...The Rho-family GTPase Rac1 activates the WAVE regulatory complex (WRC) to drive Arp2/3 complex-mediated actin polymerization in many essential processes. Rac1 binds to WRC at two distinct sites-the A and D sites. Precisely how Rac1 binds and how the binding triggers WRC activation remain unknown. Here we report WRC structures by itself, and when bound to single or double Rac1 molecules, at ~3 Å resolutions by cryogenic-electron microscopy. The structures reveal that Rac1 binds to the two sites by distinct mechanisms, and binding to the A site, but not the D site, drives WRC activation. Activation involves a series of unique conformational changes leading to the release of sequestered WCA (WH2-central-acidic) polypeptide, which stimulates the Arp2/3 complex to polymerize actin. Together with biochemical and cellular analyses, the structures provide a novel mechanistic understanding of how the Rac1-WRC-Arp2/3-actin signaling axis is regulated in diverse biological processes and diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26733.map.gz emd_26733.map.gz | 12.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26733-v30.xml emd-26733-v30.xml emd-26733.xml emd-26733.xml | 30.9 KB 30.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26733_fsc.xml emd_26733_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_26733.png emd_26733.png | 115.7 KB | ||
| Masks |  emd_26733_msk_1.map emd_26733_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-26733.cif.gz emd-26733.cif.gz | 9.1 KB | ||
| Others |  emd_26733_additional_1.map.gz emd_26733_additional_1.map.gz emd_26733_half_map_1.map.gz emd_26733_half_map_1.map.gz emd_26733_half_map_2.map.gz emd_26733_half_map_2.map.gz | 9.2 MB 59 MB 59 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26733 http://ftp.pdbj.org/pub/emdb/structures/EMD-26733 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26733 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26733 | HTTPS FTP |
-Related structure data
| Related structure data |  7usdMC  7uscC  7useC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26733.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26733.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-Rac1 complex sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8757 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_26733_msk_1.map emd_26733_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: WRC230VCA-Rac1 complex masked unsharpened map
| File | emd_26733_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-Rac1 complex masked unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: WRC230VCA-Rac1 complex unmasked unfiltered unsharpened first half map
| File | emd_26733_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-Rac1 complex unmasked unfiltered unsharpened first half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: WRC230VCA-Rac1 complex unmasked unfiltered unsharpened second half map...
| File | emd_26733_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-Rac1 complex unmasked unfiltered unsharpened second half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : WAVE regulatory complex with Rac1 bound to D-site
| Entire | Name: WAVE regulatory complex with Rac1 bound to D-site |
|---|---|
| Components |
|
-Supramolecule #1: WAVE regulatory complex with Rac1 bound to D-site
| Supramolecule | Name: WAVE regulatory complex with Rac1 bound to D-site / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 360 KDa |
-Macromolecule #1: Cytoplasmic FMR1-interacting protein 1
| Macromolecule | Name: Cytoplasmic FMR1-interacting protein 1 / type: protein_or_peptide / ID: 1 Details: This construct contains two additional uncleaved residues "GA" in the N terminus from the construct design and purification procedure. Densities for these residues are not observed in the ...Details: This construct contains two additional uncleaved residues "GA" in the N terminus from the construct design and purification procedure. Densities for these residues are not observed in the map and were not included in the sample sequence to avoid numbering shifts. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 145.36375 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MAAQVTLEDA LSNVDLLEEL PLPDQQPCIE PPPSSLLYQP NFNTNFEDRN AFVTGIARYI EQATVHSSMN EMLEEGQEYA VMLYTWRSC SRAIPQVKCN EQPNRVEIYE KTVEVLEPEV TKLMNFMYFQ RNAIERFCGE VRRLCHAERR KDFVSEAYLI T LGKFINMF ...String: MAAQVTLEDA LSNVDLLEEL PLPDQQPCIE PPPSSLLYQP NFNTNFEDRN AFVTGIARYI EQATVHSSMN EMLEEGQEYA VMLYTWRSC SRAIPQVKCN EQPNRVEIYE KTVEVLEPEV TKLMNFMYFQ RNAIERFCGE VRRLCHAERR KDFVSEAYLI T LGKFINMF AVLDELKNMK CSVKNDHSAY KRAAQFLRKM ADPQSIQESQ NLSMFLANHN KITQSLQQQL EVISGYEELL AD IVNLCVD YYENRMYLTP SEKHMLLKVM GFGLYLMDGS VSNIYKLDAK KRINLSKIDK YFKQLQVVPL FGDMQIELAR YIK TSAHYE ENKSRWTCTS SGSSPQYNIC EQMIQIREDH MRFISELARY SNSEVVTGSG RQEAQKTDAE YRKLFDLALQ GLQL LSQWS AHVMEVYSWK LVHPTDKYSN KDCPDSAEEY ERATRYNYTS EEKFALVEVI AMIKGLQVLM GRMESVFNHA IRHTV YAAL QDFSQVTLRE PLRQAIKKKK NVIQSVLQAI RKTVCDWETG HEPFNDPALR GEKDPKSGFD IKVPRRAVGP SSTQLY MVR TMLESLIADK SGSKKTLRSS LEGPTILDIE KFHRESFFYT HLINFSETLQ QCCDLSQLWF REFFLELTMG RRIQFPI EM SMPWILTDHI LETKEASMME YVLYSLDLYN DSAHYALTRF NKQFLYDEIE AEVNLCFDQF VYKLADQIFA YYKVMAGS L LLDKRLRSEC KNQGATIHLP PSNRYETLLK QRHVQLLGRS IDLNRLITQR VSAAMYKSLE LAIGRFESED LTSIVELDG LLEINRMTHK LLSRYLTLDG FDAMFREANH NVSAPYGRIT LHVFWELNYD FLPNYCYNGS TNRFVRTVLP FSQEFQRDKQ PNAQPQYLH GSKALNLAYS SIYGSYRNFV GPPHFQVICR LLGYQGIAVV MEELLKVVKS LLQGTILQYV KTLMEVMPKI C RLPRHEYG SPGILEFFHH QLKDIVEYAE LKTVCFQNLR EVGNAILFCL LIEQSLSLEE VCDLLHAAPF QNILPRVHVK EG ERLDAKM KRLESKYAPL HLVPLIERLG TPQQIAIARE GDLLTKERLC CGLSMFEVIL TRIRSFLDDP IWRGPLPSNG VMH VDECVE FHRLWSAMQF VYCIPVGTHE FTVEQCFGDG LHWAGCMIIV LLGQQRRFAV LDFCYHLLKV QKHDGKDEII KNVP LKKMV ERIRKFQILN DEIITILDKY LKSGDGEGTP VEHVRCFQPP IHQSLASS UniProtKB: Cytoplasmic FMR1-interacting protein 1 |
-Macromolecule #2: Nck-associated protein 1
| Macromolecule | Name: Nck-associated protein 1 / type: protein_or_peptide / ID: 2 Details: This construct contains two additional uncleaved residues "GA" in the N terminus from the construct design and purification procedure. Densities for these residues are not observed in the ...Details: This construct contains two additional uncleaved residues "GA" in the N terminus from the construct design and purification procedure. Densities for these residues are not observed in the map and were not included in the sample sequence to avoid numbering shifts. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 128.940727 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSRSVLQPSQ QKLAEKLTIL NDRGVGMLTR LYNIKKACGD PKAKPSYLID KNLESAVKFI VRKFPAVETR NNNQQLAQLQ KEKSEILKN LALYYFTFVD VMEFKDHVCE LLNTIDVCQV FFDITVNFDL TKNYLDLIIT YTTLMILLSR IEERKAIIGL Y NYAHEMTH ...String: MSRSVLQPSQ QKLAEKLTIL NDRGVGMLTR LYNIKKACGD PKAKPSYLID KNLESAVKFI VRKFPAVETR NNNQQLAQLQ KEKSEILKN LALYYFTFVD VMEFKDHVCE LLNTIDVCQV FFDITVNFDL TKNYLDLIIT YTTLMILLSR IEERKAIIGL Y NYAHEMTH GASDREYPRL GQMIVDYENP LKKMMEEFVP HSKSLSDALI SLQMVYPRRN LSADQWRNAQ LLSLISAPST ML NPAQSDT MPCEYLSLDA MEKWIIFGFI LCHGILNTDA TALNLWKLAL QSSSCLSLFR DEVFHIHKAA EDLFVNIRGY NKR INDIRE CKEAAVSHAG SMHRERRKFL RSALKELATV LSDQPGLLGP KALFVFMALS FARDEIIWLL RHADNMPKKS ADDF IDKHI AELIFYMEEL RAHVRKYGPV MQRYYVQYLS GFDAVVLNEL VQNLSVCPED ESIIMSSFVN TMTSLSVKQV EDGEV FDFR GMRLDWFRLQ AYTSVSKASL GLADHRELGK MMNTIIFHTK MVDSLVEMLV ETSDLSIFCF YSRAFEKMFQ QCLELP SQS RYSIAFPLLC THFMSCTHEL CPEERHHIGD RSLSLCNMFL DEMAKQARNL ITDICTEQCT LSDQLLPKHC AKTISQA VN KKSKKQTGKK GEPEREKPGV ESMRKNRLVV TNLDKLHTAL SELCFSINYV PNMVVWEHTF TPREYLTSHL EIRFTKSI V GMTMYNQATQ EIAKPSELLT SVRAYMTVLQ SIENYVQIDI TRVFNNVLLQ QTQHLDSHGE PTITSLYTNW YLETLLRQV SNGHIAYFPA MKAFVNLPTE NELTFNAEEY SDISEMRSLS ELLGPYGMKF LSESLMWHIS SQVAELKKLV VENVDVLTQM RTSFDKPDQ MAALFKRLSS VDSVLKRMTI IGVILSFRSL AQEALRDVLS YHIPFLVSSI EDFKDHIPRE TDMKVAMNVY E LSSAAGLP CEIDPALVVA LSSQKSENIS PEEEYKIACL LMVFVAVSLP TLASNVMSQY SPAIEGHCNN IHCLAKAINQ IA AALFTIH KGSIEDRLKE FLALASSSLL KIGQETDKTT TRNRESVYLL LDMIVQESPF LTMDLLESCF PYVLLRNAYH AVY KQSVTS SA UniProtKB: Nck-associated protein 1 |
-Macromolecule #3: Wiskott-Aldrich syndrome protein family member 1
| Macromolecule | Name: Wiskott-Aldrich syndrome protein family member 1 / type: protein_or_peptide / ID: 3 Details: Residues 231-248 are inserted as a flexible linker sequence. This construct contains two additional uncleaved residues "GA" in the N terminus from the construct design and purification ...Details: Residues 231-248 are inserted as a flexible linker sequence. This construct contains two additional uncleaved residues "GA" in the N terminus from the construct design and purification procedure. Densities for these residues are not observed in the map and were not included in the sample sequence to avoid numbering shifts. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.009406 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPLVKRNIDP RHLCHTALPR GIKNELECVT NISLANIIRQ LSSLSKYAED IFGELFNEAH SFSFRVNSLQ ERVDRLSVSV TQLDPKEEE LSLQDITMRK AFRSSTIQDQ QLFDRKTLPI PLQETYDVCE QPPPLNILTP YRDDGKEGLK FYTNPSYFFD L WKEKMLQD ...String: MPLVKRNIDP RHLCHTALPR GIKNELECVT NISLANIIRQ LSSLSKYAED IFGELFNEAH SFSFRVNSLQ ERVDRLSVSV TQLDPKEEE LSLQDITMRK AFRSSTIQDQ QLFDRKTLPI PLQETYDVCE QPPPLNILTP YRDDGKEGLK FYTNPSYFFD L WKEKMLQD TEDKRKEKRK QKQKNLDRPH EPEKVPRAPH DRRREWQKLA QGPELAEDDA NLLHKHIEVA NGGGSGGSGG SG GSGGSGG SKRHPSTLPV ISDARSVLLE AIRKGIQLRK VEEQREQEAK HERIENDVAT ILSRRIAVEY SDSEDDSEFD EVD WLE UniProtKB: Actin-binding protein WASF1, Actin-binding protein WASF1 |
-Macromolecule #4: Protein BRICK1
| Macromolecule | Name: Protein BRICK1 / type: protein_or_peptide / ID: 4 Details: This construct contains uncleaved residues "GHMGAA" in the N terminus from the construct design and purification procedure. Densities for the residues are not observed in the map and were ...Details: This construct contains uncleaved residues "GHMGAA" in the N terminus from the construct design and purification procedure. Densities for the residues are not observed in the map and were not included in the sample sequence to avoid numbering shifts. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.756915 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGQEDPVQR EIHQDWANRE YIEIITSSIK KIADFLNSFD MSCRSRLATL NEKLTALERR IEYIEARVTK GETLT UniProtKB: Protein BRICK1 |
-Macromolecule #5: Abl interactor 2
| Macromolecule | Name: Abl interactor 2 / type: protein_or_peptide / ID: 5 Details: The sequence only contains residues 1-158. Also, there are two additional uncleaved residues "GH" in the N terminus from the construct design and purification procedure. Densities for these ...Details: The sequence only contains residues 1-158. Also, there are two additional uncleaved residues "GH" in the N terminus from the construct design and purification procedure. Densities for these residues are not observed in the map and were not included in the sample sequence to avoid numbering shifts. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.041482 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAELQMLLEE EIPGGRRALF DSYTNLERVA DYCENNYIQS ADKQRALEET KAYTTQSLAS VAYLINTLAN NVLQMLDIQA SQLRRMESS INHISQTVDI HKEKVARREI GILTTNKNTS RTHKIIAPAN LERPVRYIRK PIDYTILDDI GHGVKVSTQ UniProtKB: Abl interactor 2 |
-Macromolecule #6: Ras-related C3 botulinum toxin substrate 1
| Macromolecule | Name: Ras-related C3 botulinum toxin substrate 1 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO / EC number: small monomeric GTPase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 21.010486 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MQAIKCVVVG DGAVGKTCLL ISYTTNAFSG EYIPTVFDNY SANVMVDGKP VNLGLWDTAG LEDYDRLRPL SYPQTDVFLI CFSLVSPAS FENVRAKWYP EVRHHCPNTP IILVGTKLDL RDDKDTIEKL KEKKLTPITY PQGLAMAKEI GAVKYLECSA L TQRGLKTV FDEAIRAVLC PPPVKKRKRK UniProtKB: Ras-related C3 botulinum toxin substrate 1 |
-Macromolecule #7: GUANOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 7 / Number of copies: 1 / Formula: GTP |
|---|---|
| Molecular weight | Theoretical: 523.18 Da |
| Chemical component information |  ChemComp-GTP: |
-Macromolecule #8: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 8 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.41 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.019 kPa / Details: 30 mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Details | Data were collected by shifting the stage to target exposure positions. |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 2512 / Average exposure time: 40.0 sec. / Average electron dose: 45.27 e/Å2 Details: Each micrograph was acquired as dose-fractionated movies consisting of 62 frames per movie. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 120000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)