[English] 日本語
 Yorodumi
Yorodumi- EMDB-26426: Meprin alpha helix in complex with fetuin-B [subparticle localise... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Meprin alpha helix in complex with fetuin-B [subparticle localised reconstruction, masked focused refinement] | |||||||||
 Map data Map data | B-factor amplitude corrected sharpened map, filtered by local resolution | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Metalloprotease / complex / helical / extracellular / ONCOPROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmeprin A / meprin A complex / epidermal growth factor receptor ligand maturation / negative regulation of endopeptidase activity / metalloendopeptidase inhibitor activity / metallodipeptidase activity / binding of sperm to zona pellucida / signaling receptor ligand precursor processing / single fertilization / cysteine-type endopeptidase inhibitor activity ...meprin A / meprin A complex / epidermal growth factor receptor ligand maturation / negative regulation of endopeptidase activity / metalloendopeptidase inhibitor activity / metallodipeptidase activity / binding of sperm to zona pellucida / signaling receptor ligand precursor processing / single fertilization / cysteine-type endopeptidase inhibitor activity / metalloendopeptidase activity / metallopeptidase activity / : / extracellular exosome / extracellular region / zinc ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
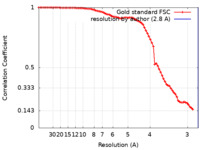
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Bayly-Jones C / Lupton CJ / Fritz C / Schlenzig D / Whisstock JC | |||||||||
| Funding support |  Germany, Germany,  Australia, 2 items Australia, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Helical ultrastructure of the metalloprotease meprin α in complex with a small molecule inhibitor. Authors: Charles Bayly-Jones / Christopher J Lupton / Claudia Fritz / Hariprasad Venugopal / Daniel Ramsbeck / Michael Wermann / Christian Jäger / Alex de Marco / Stephan Schilling / Dagmar ...Authors: Charles Bayly-Jones / Christopher J Lupton / Claudia Fritz / Hariprasad Venugopal / Daniel Ramsbeck / Michael Wermann / Christian Jäger / Alex de Marco / Stephan Schilling / Dagmar Schlenzig / James C Whisstock /   Abstract: The zinc-dependent metalloprotease meprin α is predominantly expressed in the brush border membrane of proximal tubules in the kidney and enterocytes in the small intestine and colon. In normal ...The zinc-dependent metalloprotease meprin α is predominantly expressed in the brush border membrane of proximal tubules in the kidney and enterocytes in the small intestine and colon. In normal tissue homeostasis meprin α performs key roles in inflammation, immunity, and extracellular matrix remodelling. Dysregulated meprin α is associated with acute kidney injury, sepsis, urinary tract infection, metastatic colorectal carcinoma, and inflammatory bowel disease. Accordingly, meprin α is the target of drug discovery programs. In contrast to meprin β, meprin α is secreted into the extracellular space, whereupon it oligomerises to form giant assemblies and is the largest extracellular protease identified to date (~6 MDa). Here, using cryo-electron microscopy, we determine the high-resolution structure of the zymogen and mature form of meprin α, as well as the structure of the active form in complex with a prototype small molecule inhibitor and human fetuin-B. Our data reveal that meprin α forms a giant, flexible, left-handed helical assembly of roughly 22 nm in diameter. We find that oligomerisation improves proteolytic and thermal stability but does not impact substrate specificity or enzymatic activity. Furthermore, structural comparison with meprin β reveal unique features of the active site of meprin α, and helical assembly more broadly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26426.map.gz emd_26426.map.gz | 1.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26426-v30.xml emd-26426-v30.xml emd-26426.xml emd-26426.xml | 32.1 KB 32.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26426_fsc.xml emd_26426_fsc.xml | 7 KB | Display |  FSC data file FSC data file |
| Images |  emd_26426.png emd_26426.png | 104 KB | ||
| Masks |  emd_26426_msk_1.map emd_26426_msk_1.map | 38.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-26426.cif.gz emd-26426.cif.gz | 8.6 KB | ||
| Others |  emd_26426_additional_1.map.gz emd_26426_additional_1.map.gz emd_26426_additional_2.map.gz emd_26426_additional_2.map.gz emd_26426_half_map_1.map.gz emd_26426_half_map_1.map.gz emd_26426_half_map_2.map.gz emd_26426_half_map_2.map.gz | 34.2 MB 33.1 MB 35.7 MB 35.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26426 http://ftp.pdbj.org/pub/emdb/structures/EMD-26426 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26426 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26426 | HTTPS FTP |
-Related structure data
| Related structure data |  7uaiMC  7uabC  7uacC  7uaeC  7uafC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26426.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26426.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | B-factor amplitude corrected sharpened map, filtered by local resolution | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.41333 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_26426_msk_1.map emd_26426_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: DeepEMhancer (mask) sharpened final map. Low pass filtered...
| File | emd_26426_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DeepEMhancer (mask) sharpened final map. Low pass filtered to global FSC. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: DeepEMhancer (highres) sharpened final map. Low pass filtered...
| File | emd_26426_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DeepEMhancer (highres) sharpened final map. Low pass filtered to global FSC. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Unfiltered half-map (1 of 2).
| File | emd_26426_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered half-map (1 of 2). | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Unfiltered half-map (2 of 2).
| File | emd_26426_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered half-map (2 of 2). | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Tetrameric portion of meprin alpha helix in complex with fetuin-B...
| Entire | Name: Tetrameric portion of meprin alpha helix in complex with fetuin-B [subparticle localised reconstruction] |
|---|---|
| Components |
|
-Supramolecule #1: Tetrameric portion of meprin alpha helix in complex with fetuin-B...
| Supramolecule | Name: Tetrameric portion of meprin alpha helix in complex with fetuin-B [subparticle localised reconstruction] type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: Subparticle localised reconstruction of tetrameric region of recombinant, secreted helical meprin alpha in complex with fetuin-B |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 85 kDa/nm |
-Macromolecule #1: Meprin A subunit alpha
| Macromolecule | Name: Meprin A subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: meprin A |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 67.28582 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: WSHPQFEKVP IKYLPEENVH DADFGEQKDI SEINLAAGLD LFQGDILLQK SRNGLRDPNT RWTFPIPYIL ADNLGLNAKG AILYAFEMF RLKSCVDFKP YEGESSYIIF QQFDGCWSEV GDQHVGQNIS IGQGCAYKAI IEHEILHALG FYHEQSRTDR D DYVNIWWD ...String: WSHPQFEKVP IKYLPEENVH DADFGEQKDI SEINLAAGLD LFQGDILLQK SRNGLRDPNT RWTFPIPYIL ADNLGLNAKG AILYAFEMF RLKSCVDFKP YEGESSYIIF QQFDGCWSEV GDQHVGQNIS IGQGCAYKAI IEHEILHALG FYHEQSRTDR D DYVNIWWD QILSGYQHNF DTYDDSLITD LNTPYDYESL MHYQPFSFNK NASVPTITAK IPEFNSIIGQ RLDFSAIDLE RL NRMYNCT TTHTLLDHCT FEKANICGMI QGTRDDTDWA HQDSAQAGEV DHTLLGQCTG AGYFMQFSTS SGSAEEAALL ESR ILYPKR KQQCLQFFYK MTGSPSDRLV VWVRRDDSTG NVRKLVKVQT FQGDDDHNWK IAHVVLKEEQ KFRYLFQGTK GDPQ NSTGG IYLDDITLTE TPCPTGVWTV RNFSQVLENT SKGDKLQSPR FYNSEGYGFG VTLYPNSRES SGYLRLAFHV CSGEN DAIL EWPVENRQVI ITILDQEPDV RNRMSSSMVF TTSKSHTSPA INDTVIWDRP SRVGTYHTDC NCFRSIDLGW SGFISH QML KRRSFLKNDD LIIFVDFEDI THLS UniProtKB: Meprin A subunit alpha |
-Macromolecule #2: Fetuin-B
| Macromolecule | Name: Fetuin-B / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.926691 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGLLLPLALC ILVLCCGAMS PPQLALNPSA LLSRGCNDSD VLAVAGFALR DINKDRKDGY VLRLNRVNDA QEYRRGGLGS LFYLTLDVL ETDCHVLRKK AWQDCGMRIF FESVYGQCKA IFYMNNPSRV LYLAAYNCTL RPVSKKKIYM TCPDCPSSIP T DSSNHQVL ...String: MGLLLPLALC ILVLCCGAMS PPQLALNPSA LLSRGCNDSD VLAVAGFALR DINKDRKDGY VLRLNRVNDA QEYRRGGLGS LFYLTLDVL ETDCHVLRKK AWQDCGMRIF FESVYGQCKA IFYMNNPSRV LYLAAYNCTL RPVSKKKIYM TCPDCPSSIP T DSSNHQVL EAATESLAKY NNENTSKQYS LFKVTRASSQ WVVGPSYFVE YLIKESPCTK SQASSCSLQS SDSVPVGLCK GS LTRTHWE KFVSVTCDFF ESQAPATGSE NSAVNQKPTN LPKVEESQQK NTPPTDSPSK AGPRGSVQYL PDLDDKNSQE KGP QEAFPV HLDLTTNPQG ETLDISFLFL EPMEEKLVVL PFPKEKARTA ECPGPAQNAS PLVLPPHHHH HH UniProtKB: Fetuin-B |
-Macromolecule #6: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 6 / Number of copies: 8 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #7: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 7 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #8: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 8 / Number of copies: 4 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: Pelco EasyGlow | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV / Details: 3 s blot, -3 force. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-25 / Number grids imaged: 1 / Number real images: 4068 / Average electron dose: 44.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)