+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Meprin alpha helix (activated) - full C1 consensus reconstruction | |||||||||
 マップデータ マップデータ | Unsharpened final map. Low pass filtered to the global FSC. | |||||||||
 試料 試料 |
| |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.4 Å | |||||||||
 データ登録者 データ登録者 | Bayly-Jones C / Lupton CJ / Fritz C / Schlenzig D / Whisstock JC | |||||||||
| 資金援助 |  ドイツ, ドイツ,  オーストラリア, 2件 オーストラリア, 2件
| |||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2022 ジャーナル: Nat Commun / 年: 2022タイトル: Helical ultrastructure of the metalloprotease meprin α in complex with a small molecule inhibitor. 著者: Charles Bayly-Jones / Christopher J Lupton / Claudia Fritz / Hariprasad Venugopal / Daniel Ramsbeck / Michael Wermann / Christian Jäger / Alex de Marco / Stephan Schilling / Dagmar Schlenzig ...著者: Charles Bayly-Jones / Christopher J Lupton / Claudia Fritz / Hariprasad Venugopal / Daniel Ramsbeck / Michael Wermann / Christian Jäger / Alex de Marco / Stephan Schilling / Dagmar Schlenzig / James C Whisstock /   要旨: The zinc-dependent metalloprotease meprin α is predominantly expressed in the brush border membrane of proximal tubules in the kidney and enterocytes in the small intestine and colon. In normal ...The zinc-dependent metalloprotease meprin α is predominantly expressed in the brush border membrane of proximal tubules in the kidney and enterocytes in the small intestine and colon. In normal tissue homeostasis meprin α performs key roles in inflammation, immunity, and extracellular matrix remodelling. Dysregulated meprin α is associated with acute kidney injury, sepsis, urinary tract infection, metastatic colorectal carcinoma, and inflammatory bowel disease. Accordingly, meprin α is the target of drug discovery programs. In contrast to meprin β, meprin α is secreted into the extracellular space, whereupon it oligomerises to form giant assemblies and is the largest extracellular protease identified to date (~6 MDa). Here, using cryo-electron microscopy, we determine the high-resolution structure of the zymogen and mature form of meprin α, as well as the structure of the active form in complex with a prototype small molecule inhibitor and human fetuin-B. Our data reveal that meprin α forms a giant, flexible, left-handed helical assembly of roughly 22 nm in diameter. We find that oligomerisation improves proteolytic and thermal stability but does not impact substrate specificity or enzymatic activity. Furthermore, structural comparison with meprin β reveal unique features of the active site of meprin α, and helical assembly more broadly. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_26421.map.gz emd_26421.map.gz | 122 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-26421-v30.xml emd-26421-v30.xml emd-26421.xml emd-26421.xml | 20.5 KB 20.5 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_26421_fsc.xml emd_26421_fsc.xml | 14.4 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_26421.png emd_26421.png | 70.8 KB | ||
| マスクデータ |  emd_26421_msk_1.map emd_26421_msk_1.map | 244.1 MB |  マスクマップ マスクマップ | |
| その他 |  emd_26421_half_map_1.map.gz emd_26421_half_map_1.map.gz emd_26421_half_map_2.map.gz emd_26421_half_map_2.map.gz | 226.9 MB 226.9 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26421 http://ftp.pdbj.org/pub/emdb/structures/EMD-26421 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26421 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26421 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_26421.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_26421.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Unsharpened final map. Low pass filtered to the global FSC. | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
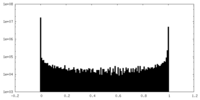
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-マスク #1
| ファイル |  emd_26421_msk_1.map emd_26421_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
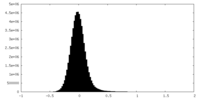
| 密度ヒストグラム |
-ハーフマップ: Unfiltered half-map (1 of 2).
| ファイル | emd_26421_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Unfiltered half-map (1 of 2). | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
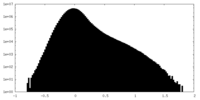
| 密度ヒストグラム |
-ハーフマップ: Unfiltered half-map (2 of 2).
| ファイル | emd_26421_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Unfiltered half-map (2 of 2). | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Full helical meprin alpha in the activated state
| 全体 | 名称: Full helical meprin alpha in the activated state |
|---|---|
| 要素 |
|
-超分子 #1: Full helical meprin alpha in the activated state
| 超分子 | 名称: Full helical meprin alpha in the activated state / タイプ: complex / キメラ: Yes / ID: 1 / 親要素: 0 / 含まれる分子: all 詳細: Full C1 reconstruction of the filamentous helix of recombinant, secreted helical meprin alpha in the activated state |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 組換発現 | 生物種:  組換細胞: Schneider-2 / 組換プラスミド: pMT/BiP/V5 |
| 分子量 | 理論値: 85 kDa/nm |
-分子 #1: Human meprin alpha
| 分子 | 名称: Human meprin alpha / タイプ: protein_or_peptide / ID: 1 / 光学異性体: LEVO / EC番号: meprin A |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 組換発現 | 生物種:  |
| 配列 | 文字列: WSHPQFEKVP IKYLPEENVH DADFGEQKDI SEINLAAGLD LFQGDILLQK SR NGLRDPN TRWTFPIPYI LADNLGLNAK GAILYAFEMF RLKSCVDFKP YEGESSYIIF QQF DGCWSE VGDQHVGQNI SIGQGCAYKA IIEHEILHAL GFYHEQSRTD RDDYVNIWWD ...文字列: WSHPQFEKVP IKYLPEENVH DADFGEQKDI SEINLAAGLD LFQGDILLQK SR NGLRDPN TRWTFPIPYI LADNLGLNAK GAILYAFEMF RLKSCVDFKP YEGESSYIIF QQF DGCWSE VGDQHVGQNI SIGQGCAYKA IIEHEILHAL GFYHEQSRTD RDDYVNIWWD QILS GYQHN FDTYDDSLIT DLNTPYDYES LMHYQPFSFN KNASVPTITA KIPEFNSIIG QRLDF SAID LERLNRMYNC TTTHTLLDHC TFEKANICGM IQGTRDDTDW AHQDSAQAGE VDHTLL GQC TGAGYFMQFS TSSGSAEEAA LLESRILYPK RKQQCLQFFY KMTGSPSDRL VVWVRRD DS TGNVRKLVKV QTFQGDDDHN WKIAHVVLKE EQKFRYLFQG TKGDPQNSTG GIYLDDIT L TETPCPTGVW TVRNFSQVLE NTSKGDKLQS PRFYNSEGYG FGVTLYPNSR ESSGYLRLA FHVCSGENDA ILEWPVENRQ VIITILDQEP DVRNRMSSSM VFTTSKSHTS PAINDTVIWD RPSRVGTYH TDCNCFRSID LGWSGFISHQ MLKRRSFLKN DDLIIFVDFE DITHLS |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | filament |
- 試料調製
試料調製
| 濃度 | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7.4 構成要素:
| |||||||||
| グリッド | モデル: Quantifoil R2/2 / 材質: COPPER / メッシュ: 200 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 雰囲気: AIR | |||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277.15 K / 装置: FEI VITROBOT MARK IV / 詳細: 3 s blot, -3 force. | |||||||||
| 詳細 | Polydisperse |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / デジタル化 - 画像ごとのフレーム数: 1-50 / 撮影したグリッド数: 1 / 実像数: 2648 / 平均電子線量: 44.5 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): 1.5 µm / 最小 デフォーカス(公称値): 0.5 µm / 倍率(公称値): 130000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル | PDB ID: |
|---|---|
| 詳細 | Rigid body fit only |
| 精密化 | 空間: REAL / プロトコル: OTHER |
 ムービー
ムービー コントローラー
コントローラー
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)