[English] 日本語
 Yorodumi
Yorodumi- EMDB-25914: Cardiac thin filament decorated with regulatory M-domain of cardi... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
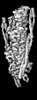
| Title | Cardiac thin filament decorated with regulatory M-domain of cardiac myosin binding protein C | |||||||||
 Map data Map data | cardiac thin filament decorated with THM of regulatory M-domain of cMyBP-C | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cardiac contraction regulator / muscle protein / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationC zone / regulation of muscle filament sliding / striated muscle myosin thick filament / regulation of striated muscle contraction / cardiac myofibril / A band / Striated Muscle Contraction / regulation of cardiac muscle cell contraction / M band / structural constituent of muscle ...C zone / regulation of muscle filament sliding / striated muscle myosin thick filament / regulation of striated muscle contraction / cardiac myofibril / A band / Striated Muscle Contraction / regulation of cardiac muscle cell contraction / M band / structural constituent of muscle / sarcomere organization / ventricular cardiac muscle tissue morphogenesis / myosin heavy chain binding / myosin binding / ATPase activator activity / heart morphogenesis / cardiac muscle contraction / titin binding / sarcomere / actin binding / cytoskeleton / cell adhesion / ATP binding / metal ion binding / identical protein binding / cytosol Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 8.0 Å | |||||||||
 Authors Authors | Risi CM / Galkin VE | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2022 Journal: J Mol Biol / Year: 2022Title: Cryo-Electron Microscopy Reveals Cardiac Myosin Binding Protein-C M-Domain Interactions with the Thin Filament. Authors: Cristina M Risi / Edwin Villanueva / Betty Belknap / Rachel L Sadler / Samantha P Harris / Howard D White / Vitold E Galkin /  Abstract: Cardiac myosin binding protein C (cMyBP-C) modulates cardiac contraction via direct interactions with cardiac thick (myosin) and thin (actin) filaments (cTFs). While its C-terminal domains (e.g. C8- ...Cardiac myosin binding protein C (cMyBP-C) modulates cardiac contraction via direct interactions with cardiac thick (myosin) and thin (actin) filaments (cTFs). While its C-terminal domains (e.g. C8-C10) anchor cMyBP-C to the backbone of the thick filament, its N-terminal domains (NTDs) (e.g. C0, C1, M, and C2) bind to both myosin and actin to accomplish its dual roles of inhibiting thick filaments and activating cTFs. While the positions of C0, C1 and C2 on cTF have been reported, the binding site of the M-domain on the surface of the cTF is unknown. Here, we used cryo-EM to reveal that the M-domain interacts with actin via helix 3 of its ordered tri-helix bundle region, while the unstructured part of the M-domain does not maintain extensive interactions with actin. We combined the recently obtained structure of the cTF with the positions of all the four NTDs on its surface to propose a complete model of the NTD binding to the cTF. The model predicts that the interactions of the NTDs with the cTF depend on the activation state of the cTF. At the peak of systole, when bound to the extensively activated cTF, NTDs would inhibit actomyosin interactions. In contrast, at falling Ca levels, NTDs would not compete with the myosin heads for binding to the cTF, but would rather promote formation of active cross-bridges at the adjacent regulatory units located at the opposite cTF strand. Our structural data provides a testable model of the cTF regulation by the cMyBP-C. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25914.map.gz emd_25914.map.gz | 3.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25914-v30.xml emd-25914-v30.xml emd-25914.xml emd-25914.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25914.png emd_25914.png | 117.1 KB | ||
| Filedesc metadata |  emd-25914.cif.gz emd-25914.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25914 http://ftp.pdbj.org/pub/emdb/structures/EMD-25914 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25914 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25914 | HTTPS FTP |
-Related structure data
| Related structure data |  7titMC  7tijC  7tj7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25914.map.gz / Format: CCP4 / Size: 4.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25914.map.gz / Format: CCP4 / Size: 4.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cardiac thin filament decorated with THM of regulatory M-domain of cMyBP-C | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.345 Å | ||||||||||||||||||||||||||||||||||||
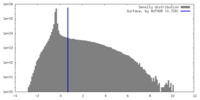
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Complex of cardiac thin filament with C1-M fragment of cardiac my...
| Entire | Name: Complex of cardiac thin filament with C1-M fragment of cardiac myosin binding protein C (cMyBP-C) |
|---|---|
| Components |
|
-Supramolecule #1: Complex of cardiac thin filament with C1-M fragment of cardiac my...
| Supramolecule | Name: Complex of cardiac thin filament with C1-M fragment of cardiac myosin binding protein C (cMyBP-C) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Only triple helix motif of the M-domain is visible in the map. C1 domain is not bound and hence not visible in the map. |
|---|
-Macromolecule #1: cardiac actin
| Macromolecule | Name: cardiac actin / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.830551 KDa |
| Sequence | String: DDEETTALVC DNGSGLVKAG FAGDDAPRAV FPSIVGRPRH QGVMVGMGQK DSYVGDEAQS KRGILTLKYP IEHGIITNWD DMEKIWHHT FYNELRVAPE EHPTLLTEAP LNPKANREKM TQIMFETFNV PAMYVAIQAV LSLYASGRTT GIVLDSGDGV T HNVPIYEG ...String: DDEETTALVC DNGSGLVKAG FAGDDAPRAV FPSIVGRPRH QGVMVGMGQK DSYVGDEAQS KRGILTLKYP IEHGIITNWD DMEKIWHHT FYNELRVAPE EHPTLLTEAP LNPKANREKM TQIMFETFNV PAMYVAIQAV LSLYASGRTT GIVLDSGDGV T HNVPIYEG YALPHAIMRL DLAGRDLTDY LMKILTERGY SFVTTAEREI VRDIKEKLCY VALDFENEMA TAASSSSLEK SY ELPDGQV ITIGNERFRC PETLFQPSFI GMESAGIHET TYNSIMKCDI DIRKDLYANN VLSGGTTMYP GIADRMQKEI TAL APSTMK IKIIAPPERK YSVWIGGSIL ASLSTFQQMW ISKQEYDEAG PSIVHRKCF UniProtKB: Actin alpha cardiac muscle 1 |
-Macromolecule #2: Myosin-binding protein C, cardiac-type
| Macromolecule | Name: Myosin-binding protein C, cardiac-type / type: protein_or_peptide / ID: 2 Details: only triple helix motif from C1-M construct bound to thin filament is visible in the map and contained in the model Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.803123 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DDPIGLFVMR PQDGEVTVGG SITFSARVAG ASLLKPPVVK WFKGKWVDLS SKVGQHLQLH DSYDRASKVY LFELHITDAQ PAFTGSYRC EVSTKDKFDC SNFNLTVHEA MGTGDLDLLS AFRRTSLAGG GRRISDSHED TGILDFSSLL KKRDSFRTPR D SKLEAPAE ...String: DDPIGLFVMR PQDGEVTVGG SITFSARVAG ASLLKPPVVK WFKGKWVDLS SKVGQHLQLH DSYDRASKVY LFELHITDAQ PAFTGSYRC EVSTKDKFDC SNFNLTVHEA MGTGDLDLLS AFRRTSLAGG GRRISDSHED TGILDFSSLL KKRDSFRTPR D SKLEAPAE EDVWEILRQA PPSEYERIAF QYGVTDLRGM LKRLKGMRRD EKKSTAFQKK LE UniProtKB: Myosin-binding protein C, cardiac-type |
-Macromolecule #3: tropomyosin model
| Macromolecule | Name: tropomyosin model / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.507176 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: PELCO Ultrathin Carbon with Lacey Carbon / Material: COPPER / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 20 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 275 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 34.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 27.4 Å Applied symmetry - Helical parameters - Δ&Phi: -166.7 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 8.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: IHRSR / Number images used: 14282 |
|---|---|
| Startup model | Type of model: OTHER / Details: F-actin model filtered to 100A resolution |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: IHRSR |
-Atomic model buiding 1
| Initial model |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT | ||||||||
| Output model |  PDB-7tit: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)