[English] 日本語
 Yorodumi
Yorodumi- EMDB-25648: CryoEM structure of the adenosine 2A receptor-BRIL/Anti BRIL Fab ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the adenosine 2A receptor-BRIL/Anti BRIL Fab complex with ZM241385 | |||||||||
 Map data Map data | Structure of the adenosine 2A receptor-BRIL/Anti BRIL Fab with ZM241385 | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of acetylcholine secretion, neurotransmission / positive regulation of circadian sleep/wake cycle, sleep / regulation of norepinephrine secretion / negative regulation of alpha-beta T cell activation / Adenosine P1 receptors / G protein-coupled adenosine receptor activity / G protein-coupled adenosine receptor signaling pathway / response to purine-containing compound / sensory perception / positive regulation of urine volume ...positive regulation of acetylcholine secretion, neurotransmission / positive regulation of circadian sleep/wake cycle, sleep / regulation of norepinephrine secretion / negative regulation of alpha-beta T cell activation / Adenosine P1 receptors / G protein-coupled adenosine receptor activity / G protein-coupled adenosine receptor signaling pathway / response to purine-containing compound / sensory perception / positive regulation of urine volume / NGF-independant TRKA activation / Surfactant metabolism / synaptic transmission, dopaminergic / : / inhibitory postsynaptic potential / negative regulation of vascular permeability / synaptic transmission, cholinergic / type 5 metabotropic glutamate receptor binding / positive regulation of glutamate secretion / blood circulation / response to caffeine / intermediate filament / eating behavior / presynaptic active zone / alpha-actinin binding / membrane depolarization / regulation of calcium ion transport / asymmetric synapse / axolemma / : / cellular defense response / prepulse inhibition / phagocytosis / response to amphetamine / presynaptic modulation of chemical synaptic transmission / excitatory postsynaptic potential / positive regulation of synaptic transmission, glutamatergic / neuron projection morphogenesis / regulation of mitochondrial membrane potential / synaptic transmission, glutamatergic / positive regulation of long-term synaptic potentiation / locomotory behavior / central nervous system development / astrocyte activation / positive regulation of synaptic transmission, GABAergic / positive regulation of protein secretion / apoptotic signaling pathway / electron transport chain / positive regulation of apoptotic signaling pathway / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / adenylate cyclase-activating G protein-coupled receptor signaling pathway / negative regulation of inflammatory response / vasodilation / blood coagulation / cell-cell signaling / presynaptic membrane / G alpha (s) signalling events / postsynaptic membrane / negative regulation of neuron apoptotic process / periplasmic space / electron transfer activity / calmodulin binding / response to xenobiotic stimulus / inflammatory response / iron ion binding / negative regulation of cell population proliferation / neuronal cell body / glutamatergic synapse / lipid binding / apoptotic process / dendrite / heme binding / protein-containing complex binding / regulation of DNA-templated transcription / enzyme binding / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Zhang KH / Wu H / Hoppe N / Manglik A / Cheng YF | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Fusion protein strategies for cryo-EM study of G protein-coupled receptors. Authors: Kaihua Zhang / Hao Wu / Nicholas Hoppe / Aashish Manglik / Yifan Cheng /  Abstract: Single particle cryogenic-electron microscopy (cryo-EM) is used extensively to determine structures of activated G protein-coupled receptors (GPCRs) in complex with G proteins or arrestins. However, ...Single particle cryogenic-electron microscopy (cryo-EM) is used extensively to determine structures of activated G protein-coupled receptors (GPCRs) in complex with G proteins or arrestins. However, applying it to GPCRs without signaling proteins remains challenging because most receptors lack structural features in their soluble domains to facilitate image alignment. In GPCR crystallography, inserting a fusion protein between transmembrane helices 5 and 6 is a highly successful strategy for crystallization. Although a similar strategy has the potential to broadly facilitate cryo-EM structure determination of GPCRs alone without signaling protein, the critical determinants that make this approach successful are not yet clear. Here, we address this shortcoming by exploring different fusion protein designs, which lead to structures of antagonist bound A adenosine receptor at 3.4 Å resolution and unliganded Smoothened at 3.7 Å resolution. The fusion strategies explored here are likely applicable to cryo-EM interrogation of other GPCRs and small integral membrane proteins. #1:  Journal: Science / Year: 2012 Journal: Science / Year: 2012Title: Structural Basis for Allosteric Regulation of GPCRs by Sodium Ions Authors: Liu W | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25648.map.gz emd_25648.map.gz | 97.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25648-v30.xml emd-25648-v30.xml emd-25648.xml emd-25648.xml | 11 KB 11 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25648.png emd_25648.png | 33.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25648 http://ftp.pdbj.org/pub/emdb/structures/EMD-25648 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25648 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25648 | HTTPS FTP |
-Validation report
| Summary document |  emd_25648_validation.pdf.gz emd_25648_validation.pdf.gz | 395 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25648_full_validation.pdf.gz emd_25648_full_validation.pdf.gz | 394.5 KB | Display | |
| Data in XML |  emd_25648_validation.xml.gz emd_25648_validation.xml.gz | 6.3 KB | Display | |
| Data in CIF |  emd_25648_validation.cif.gz emd_25648_validation.cif.gz | 7.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25648 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25648 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25648 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25648 | HTTPS FTP |
-Related structure data
| Related structure data |  7t32MC  8cxoC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25648.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25648.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the adenosine 2A receptor-BRIL/Anti BRIL Fab with ZM241385 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.835 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : A2A adenosine receptor-BRIL/Anti BRIL Fab complex
| Entire | Name: A2A adenosine receptor-BRIL/Anti BRIL Fab complex |
|---|---|
| Components |
|
-Supramolecule #1: A2A adenosine receptor-BRIL/Anti BRIL Fab complex
| Supramolecule | Name: A2A adenosine receptor-BRIL/Anti BRIL Fab complex / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Adenosine receptor A2a/Soluble cytochrome b562 Fusion Protein
| Macromolecule | Name: Adenosine receptor A2a/Soluble cytochrome b562 Fusion Protein type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 43.415855 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VYITVELAIA VLAILGNVLV CWAVWLNSNL QNVTNYFVVS LAAADIAVGV LAIPFAITIS TGFCAACHGC LFIACFVLVL TQSSIFSLL AIAIDRYIAI RIPLRYNGLV TGTRAKGIIA ICWVLSFAIG LTPMLGWNNC GQPKEGKNHS QGCGEGQVAC L FEDVVPMN ...String: VYITVELAIA VLAILGNVLV CWAVWLNSNL QNVTNYFVVS LAAADIAVGV LAIPFAITIS TGFCAACHGC LFIACFVLVL TQSSIFSLL AIAIDRYIAI RIPLRYNGLV TGTRAKGIIA ICWVLSFAIG LTPMLGWNNC GQPKEGKNHS QGCGEGQVAC L FEDVVPMN YMVYFNFFAC VLVPLLLMLG VYLRIFLAAR RQLADLEDNW ETLNDNLKVI EKADNAAQVK DALTKMRAAA LD AQKATPP KLEDKSPDSP EMKDFRHGFD ILVGQIDDAL KLANEGKVKE AQAAAEQLKT TRNAYIQKYL ERARSTLQKE VHA AKSLAI IVGLFALCWL PLHIINCFTF FCPDCSHAPL WLMYLAIVLS HTNSVVNPFI YAYRIREFRQ TFRKI |
-Macromolecule #2: 4-{2-[(7-amino-2-furan-2-yl[1,2,4]triazolo[1,5-a][1,3,5]triazin-5...
| Macromolecule | Name: 4-{2-[(7-amino-2-furan-2-yl[1,2,4]triazolo[1,5-a][1,3,5]triazin-5-yl)amino]ethyl}phenol type: ligand / ID: 2 / Number of copies: 1 / Formula: ZMA |
|---|---|
| Molecular weight | Theoretical: 337.336 Da |
| Chemical component information | 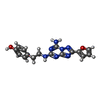 ChemComp-ZMA: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 67.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.4 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 215946 |
|---|---|
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller


















