[English] 日本語
 Yorodumi
Yorodumi- EMDB-25173: Cryo-EM structure of ACKR3 in complex with CXCL12 and an intracel... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of ACKR3 in complex with CXCL12 and an intracellular Fab | ||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Atypical Chemokine Receptor / MEMBRANE PROTEIN / SIGNALING PROTEIN / SIGNALING PROTEIN-IMMUNE SYSTEM complex | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationoculomotor nerve development / positive regulation of mesenchymal stem cell migration / telencephalon cell migration / chemokine (C-X-C motif) ligand 12 signaling pathway / response to ultrasound / negative regulation of leukocyte tethering or rolling / regulation of actin polymerization or depolymerization / C-X-C chemokine binding / CXCL12-activated CXCR4 signaling pathway / chemokine receptor binding ...oculomotor nerve development / positive regulation of mesenchymal stem cell migration / telencephalon cell migration / chemokine (C-X-C motif) ligand 12 signaling pathway / response to ultrasound / negative regulation of leukocyte tethering or rolling / regulation of actin polymerization or depolymerization / C-X-C chemokine binding / CXCL12-activated CXCR4 signaling pathway / chemokine receptor binding / positive regulation of axon extension involved in axon guidance / CXCR chemokine receptor binding / C-X-C chemokine receptor activity / positive regulation of vasculature development / Signaling by ROBO receptors / induction of positive chemotaxis / negative regulation of dendritic cell apoptotic process / integrin activation / positive regulation of dopamine secretion / negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage / cellular response to chemokine / C-C chemokine receptor activity / C-C chemokine binding / positive regulation of monocyte chemotaxis / chemokine activity / scavenger receptor activity / blood circulation / Chemokine receptors bind chemokines / positive regulation of calcium ion import / detection of temperature stimulus involved in sensory perception of pain / animal organ regeneration / detection of mechanical stimulus involved in sensory perception of pain / vasculogenesis / positive regulation of T cell migration / Nuclear signaling by ERBB4 / coreceptor activity / positive regulation of endothelial cell proliferation / clathrin-coated pit / positive regulation of neuron differentiation / positive regulation of cell adhesion / axon guidance / chemokine-mediated signaling pathway / growth factor activity / cell chemotaxis / adult locomotory behavior / calcium-mediated signaling / defense response / receptor internalization / recycling endosome / response to peptide hormone / integrin binding / neuron migration / response to virus / chemotaxis / intracellular calcium ion homeostasis / extracellular matrix / positive regulation of cytosolic calcium ion concentration / angiogenesis / Estrogen-dependent gene expression / G alpha (i) signalling events / response to hypoxia / early endosome / positive regulation of ERK1 and ERK2 cascade / cell adhesion / endosome / immune response / positive regulation of cell migration / G protein-coupled receptor signaling pathway / negative regulation of cell population proliferation / signaling receptor binding / external side of plasma membrane / cell surface / signal transduction / : / extracellular exosome / extracellular region / membrane / plasma membrane Similarity search - Function | ||||||||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||||||||||||||
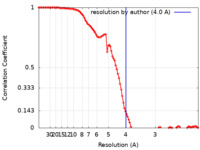
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | ||||||||||||||||||||||||
 Authors Authors | Yen YC / Schafer CT / Gustavsson M / Handel TM / Tesmer JJG | ||||||||||||||||||||||||
| Funding support |  United States, 7 items United States, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Structures of atypical chemokine receptor 3 reveal the basis for its promiscuity and signaling bias. Authors: Yu-Chen Yen / Christopher T Schafer / Martin Gustavsson / Stefanie A Eberle / Pawel K Dominik / Dawid Deneka / Penglie Zhang / Thomas J Schall / Anthony A Kossiakoff / John J G Tesmer / Tracy M Handel /    Abstract: Both CXC chemokine receptor 4 (CXCR4) and atypical chemokine receptor 3 (ACKR3) are activated by the chemokine CXCL12 yet evoke distinct cellular responses. CXCR4 is a canonical G protein-coupled ...Both CXC chemokine receptor 4 (CXCR4) and atypical chemokine receptor 3 (ACKR3) are activated by the chemokine CXCL12 yet evoke distinct cellular responses. CXCR4 is a canonical G protein-coupled receptor (GPCR), whereas ACKR3 is intrinsically biased for arrestin. The molecular basis for this difference is not understood. Here, we describe cryo-EM structures of ACKR3 in complex with CXCL12, a more potent CXCL12 variant, and a small-molecule agonist. The bound chemokines adopt an unexpected pose relative to those established for CXCR4 and observed in other receptor-chemokine complexes. Along with functional studies, these structures provide insight into the ligand-binding promiscuity of ACKR3, why it fails to couple to G proteins, and its bias toward β-arrestin. The results lay the groundwork for understanding the physiological interplay of ACKR3 with other GPCRs. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25173.map.gz emd_25173.map.gz | 97.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25173-v30.xml emd-25173-v30.xml emd-25173.xml emd-25173.xml | 21.2 KB 21.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25173_fsc.xml emd_25173_fsc.xml | 10.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_25173.png emd_25173.png | 85.2 KB | ||
| Filedesc metadata |  emd-25173.cif.gz emd-25173.cif.gz | 7.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25173 http://ftp.pdbj.org/pub/emdb/structures/EMD-25173 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25173 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25173 | HTTPS FTP |
-Related structure data
| Related structure data |  7sk5MC  7sk3C  7sk4C  7sk6C  7sk7C  7sk8C  7sk9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25173.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25173.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Complex structure of ACKR3-CXCL12-CID24-NB
| Entire | Name: Complex structure of ACKR3-CXCL12-CID24-NB |
|---|---|
| Components |
|
-Supramolecule #1: Complex structure of ACKR3-CXCL12-CID24-NB
| Supramolecule | Name: Complex structure of ACKR3-CXCL12-CID24-NB / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|
-Macromolecule #1: Atypical chemokine receptor 3
| Macromolecule | Name: Atypical chemokine receptor 3 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.196406 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GAPDLHLFDY SEPGNFSDIS WPCNSSDCIV VDTVMCPNMP NKSVLLYTLS FIYIFIFVIG MIANSVVVWV NIQAKTTGYD THCYILNLA IADLWVVLTI PVWVVSLVQH NQWPMGELTC KVTHLIFSIN LFGSIFFLTC MSVDRYLSIT YFTNTPSSRK K MVRRVVCI ...String: GAPDLHLFDY SEPGNFSDIS WPCNSSDCIV VDTVMCPNMP NKSVLLYTLS FIYIFIFVIG MIANSVVVWV NIQAKTTGYD THCYILNLA IADLWVVLTI PVWVVSLVQH NQWPMGELTC KVTHLIFSIN LFGSIFFLTC MSVDRYLSIT YFTNTPSSRK K MVRRVVCI LVWLLAFCVS LPDTYYLKTV TSASNNETYC RSFYPEHSIK EWLIGMELVS VVLGFAVPFS IIAVFYFLLA RA ISASSDQ EKHSSRKIIF SYVVVFLVCW LPYHVAVLLD IFSILHYIPF TCRLEHALFT ALHVTQCLSL VHCCVNPVLY SFI NRNYRY ELMKAFIFKY SAKTGLTKLI DASRVSETEY SALEQSTKGR PLEVLFQGPH HHHHHHHHHD YKDDDDK UniProtKB: Atypical chemokine receptor 3 |
-Macromolecule #2: CID24 Fab heavy chain
| Macromolecule | Name: CID24 Fab heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.380223 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NISSSSIHWV RQAPGKGLEW VASISPSYGY TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARVSYWDWTW GWSKYEGMDY WGQGTLVTVS SASTKGPSVF PLAPSSKSTS GGTAALGCLV K DYFPEPVT ...String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NISSSSIHWV RQAPGKGLEW VASISPSYGY TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARVSYWDWTW GWSKYEGMDY WGQGTLVTVS SASTKGPSVF PLAPSSKSTS GGTAALGCLV K DYFPEPVT VSWNSGALTS GVHTFPAVLQ SSGLYSLSSV VTVPSSSLGT QTYICNVNHK PSNTKVDKKV EPKSCDKTHT |
-Macromolecule #3: Stromal cell-derived factor 1
| Macromolecule | Name: Stromal cell-derived factor 1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.97846 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KPVSLSYRCP CRFFESHVAR ANVKHLKILN TPNCALQIVA RLKNNNRQVC IDPKLKWIQE YLEKALNK UniProtKB: Stromal cell-derived factor 1 |
-Macromolecule #4: CID24 Fab light chain
| Macromolecule | Name: CID24 Fab light chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.471031 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQSYYYPITF GQGTKVEIKR TVAAPSVFIF PPSDSQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQSYYYPITF GQGTKVEIKR TVAAPSVFIF PPSDSQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC |
-Macromolecule #5: Anti-Fab nanobody
| Macromolecule | Name: Anti-Fab nanobody / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.390644 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSQVQLQESG GGLVQPGGSL RLSCAASGRT ISRYAMSWFR QAPGKEREFV AVARRSGDGA FYADSVQGRF TVSRDDAKNT VYLQMNSLK PEDTAVYYCA IDSDTFYSGS YDYWGQGTQV TVSS |
-Macromolecule #6: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 6 / Number of copies: 7 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 53.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)