+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | apo NEXT overall reconstruction | |||||||||
 Map data Map data | apo NEXT overall map | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
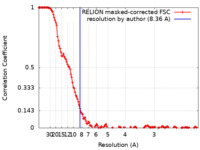
| Method | single particle reconstruction / cryo EM / Resolution: 8.36 Å | |||||||||
 Authors Authors | Puno MR / Lima CD | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2022 Journal: Cell / Year: 2022Title: Structural basis for RNA surveillance by the human nuclear exosome targeting (NEXT) complex. Authors: M Rhyan Puno / Christopher D Lima /  Abstract: RNA quality control relies on co-factors and adaptors to identify and prepare substrates for degradation by ribonucleases such as the 3' to 5' ribonucleolytic RNA exosome. Here, we determined ...RNA quality control relies on co-factors and adaptors to identify and prepare substrates for degradation by ribonucleases such as the 3' to 5' ribonucleolytic RNA exosome. Here, we determined cryogenic electron microscopy structures of human nuclear exosome targeting (NEXT) complexes bound to RNA that reveal mechanistic insights to substrate recognition and early steps that precede RNA handover to the exosome. The structures illuminate ZCCHC8 as a scaffold, mediating homodimerization while embracing the MTR4 helicase and flexibly anchoring RBM7 to the helicase core. All three subunits collaborate to bind the RNA, with RBM7 and ZCCHC8 surveying sequences upstream of the 3' end to facilitate RNA capture by MTR4. ZCCHC8 obscures MTR4 surfaces important for RNA binding and extrusion as well as MPP6-dependent recruitment and docking onto the RNA exosome core, interactions that contribute to RNA surveillance by coordinating RNA capture, translocation, and extrusion from the helicase to the exosome for decay. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24884.map.gz emd_24884.map.gz | 15.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24884-v30.xml emd-24884-v30.xml emd-24884.xml emd-24884.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24884_fsc.xml emd_24884_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_24884.png emd_24884.png | 37 KB | ||
| Others |  emd_24884_half_map_1.map.gz emd_24884_half_map_1.map.gz emd_24884_half_map_2.map.gz emd_24884_half_map_2.map.gz | 171.1 MB 171 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24884 http://ftp.pdbj.org/pub/emdb/structures/EMD-24884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24884 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24884.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24884.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | apo NEXT overall map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.088 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map 1
| File | emd_24884_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_24884_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human Nuclear Exosome Targeting (NEXT) complex
| Entire | Name: human Nuclear Exosome Targeting (NEXT) complex |
|---|---|
| Components |
|
-Supramolecule #1: human Nuclear Exosome Targeting (NEXT) complex
| Supramolecule | Name: human Nuclear Exosome Targeting (NEXT) complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Macromolecule #1: MTR4
| Macromolecule | Name: MTR4 / type: protein_or_peptide / ID: 1 Details: The first three amino acid residues (SGD) in the sample sequence are cloning artefact. Enantiomer: LEVO / EC number: RNA helicase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: SGDMADAFGD ELFSVFEGDS TTAAGTKKDK EKDKGKWKGP PGSADKAGKR FDGKLQSEST NNGKNKRDVD FEGTDEPIFG KKPRIEESI TEDLSLADLM PRVKVQSVET VEGCTHEVAL PAEEDYLPLK PRVGKAAKEY PFILDAFQRE AIQCVDNNQS V LVSAHTSA ...String: SGDMADAFGD ELFSVFEGDS TTAAGTKKDK EKDKGKWKGP PGSADKAGKR FDGKLQSEST NNGKNKRDVD FEGTDEPIFG KKPRIEESI TEDLSLADLM PRVKVQSVET VEGCTHEVAL PAEEDYLPLK PRVGKAAKEY PFILDAFQRE AIQCVDNNQS V LVSAHTSA GKTVCAEYAI ALALREKQRV IFTSPIKALS NQKYREMYEE FQDVGLMTGD VTINPTASCL VMTTEILRSM LY RGSEVMR EVAWVIFDEI HYMRDSERGV VWEETIILLP DNVHYVFLSA TIPNARQFAE WICHLHKQPC HVIYTDYRPT PLQ HYIFPA GGDGLHLVVD ENGDFREDNF NTAMQVLRDA GDLAKGDQKG RKGGTKGPSN VFKIVKMIME RNFQPVIIFS FSKK DCEAY ALQMTKLDFN TDEEKKMVEE VFSNAIDCLS DEDKKLPQVE HVLPLLKRGI GIHHGGLLPI LKETIEILFS EGLIK ALFA TETFAMGINM PARTVLFTNA RKFDGKDFRW ISSGEYIQMS GRAGRRGMDD RGIVILMVDE KMSPTIGKQL LKGSAD PLN SAFHLTYNMV LNLLRVEEIN PEYMLEKSFY QFQHYRAIPG VVEKVKNSEE QYNKIVIPNE ESVVIYYKIR QQLAKLG KE IEEYIHKPKY CLPFLQPGRL VKVKNEGDDF GWGVVVNFSK KSNVKPNSGE LDPLYVVEVL LRCSKESLKN SATEAAKP A KPDEKGEMQV VPVLVHLLSA ISSVRLYIPK DLRPVDNRQS VLKSIQEVQK RFPDGIPLLD PIDDMGIQDQ GLKKVIQKV EAFEHRMYSH PLHNDPNLET VYTLCEKKAQ IAIDIKSAKR ELKKARTVLQ MDELKCRKRV LRRLGFATSS DVIEMKGRVA CEISSADEL LLTEMMFNGL FNDLSAEQAT ALLSCFVFQE NSSEMPKLTE QLAGPLRQMQ ECAKRIAKVS AEAKLEIDEE T YLSSFKPH LMDVVYTWAT GATFAHICKM TDVFEGSIIR CMRRLEELLR QMCQAAKAIG NTELENKFAE GITKIKRDIV FA ASLYL |
-Macromolecule #2: ZCCHC8
| Macromolecule | Name: ZCCHC8 / type: protein_or_peptide / ID: 2 Details: The first three amino acid residues (SGD) in the sample sequence are cloning artefact. Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: SGDMAAEVYF GDLELFEPFD HPEESIPKPV HTRFKDDDGD EEDENGVGDA ELRERLRQCE ETIEQLRAEN QELKRKLNIL TRPSGILVN DTKLDGPILQ ILFMNNAISK QYHQEIEEFV SNLVKRFEEQ QKNDVEKTSF NLLPQPSSIV LEEDHKVEES C AIKNNKEA ...String: SGDMAAEVYF GDLELFEPFD HPEESIPKPV HTRFKDDDGD EEDENGVGDA ELRERLRQCE ETIEQLRAEN QELKRKLNIL TRPSGILVN DTKLDGPILQ ILFMNNAISK QYHQEIEEFV SNLVKRFEEQ QKNDVEKTSF NLLPQPSSIV LEEDHKVEES C AIKNNKEA FSVVGSVLYF TNFCLDKLGQ PLLNENPQLS EGWEIPKYHQ VFSHIVSLEG QEIQVKAKRP KPHCFNCGSE EH QMKDCPM PRNAARISEK RKEYMDACGE ANNQNFQQRY HAEEVEERFG RFKPGVISEE LQDALGVTDK SLPPFIYRMR QLG YPPGWL KEAELENSGL ALYDGKDGTD GETEVGEIQQ NKSVTYDLSK LVNYPGFNIS TPRGIPDEWR IFGSIPMQAC QQKD VFANY LTSNFQAPGV KSGGAVDEDA LTLEELEEQQ RRIWAALEQA ESVNSDSDVP VDTPLTGNSV ASSPCPNELD LPVPE GKTS EKQTLDEPEV PEIFTKKSEA GHASSPDSEV TSLCQKEKAE LAPVNTEGAL LDNGSVVPNC DISNGGSQKL FPADTS PST ATKIHSPIPD MSKFATGITP FEFENMAEST GMYLRIRSLL KNSPRNQQKN KKASE |
-Macromolecule #3: RBM7
| Macromolecule | Name: RBM7 / type: protein_or_peptide / ID: 3 Details: The first three amino acid residues (SGD) in the sample sequence are cloning artefact. Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: SGDEADRTLF VGNLETKVTE ELLFELFHQA GPVIKVKIPK DKDGKPKQFA FVNFKHEVSV PYAMNLLNGI KLYGRPIKIQ FRS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV / Details: 30 s wait time, blot for 2.5 s before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 67.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)