[English] 日本語
 Yorodumi
Yorodumi- EMDB-23474: Cryo-EM structure of full-length TRPV1 with capsaicin at 4 degree... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23474 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of full-length TRPV1 with capsaicin at 4 degrees Celsius | |||||||||
 Map data Map data | map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Heat sensing ion channel / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of iodide transmembrane transport / positive regulation of membrane depolarization / negative regulation of establishment of blood-brain barrier / response to capsazepine / sensory perception of mechanical stimulus / peptide secretion / cellular response to temperature stimulus / positive regulation of sensory perception of pain / temperature-gated ion channel activity / detection of chemical stimulus involved in sensory perception of pain ...negative regulation of iodide transmembrane transport / positive regulation of membrane depolarization / negative regulation of establishment of blood-brain barrier / response to capsazepine / sensory perception of mechanical stimulus / peptide secretion / cellular response to temperature stimulus / positive regulation of sensory perception of pain / temperature-gated ion channel activity / detection of chemical stimulus involved in sensory perception of pain / positive regulation of renal sodium excretion / negative regulation of axon regeneration / TRP channels / positive regulation of cardiac muscle cell differentiation / smooth muscle contraction involved in micturition / fever generation / detection of temperature stimulus involved in thermoception / thermoception / urinary bladder smooth muscle contraction / response to pH / glutamate secretion / monoatomic cation transmembrane transporter activity / negative regulation of systemic arterial blood pressure / positive regulation of urine volume / excitatory extracellular ligand-gated monoatomic ion channel activity / dendritic spine membrane / negative regulation of heart rate / response to acidic pH / response to pain / diet induced thermogenesis / cellular response to alkaloid / temperature homeostasis / cellular response to cytokine stimulus / cellular response to ATP / intracellularly gated calcium channel activity / detection of temperature stimulus involved in sensory perception of pain / negative regulation of mitochondrial membrane potential / calcium ion import across plasma membrane / behavioral response to pain / positive regulation of vasoconstriction / monoatomic ion channel activity / ligand-gated monoatomic ion channel activity / monoatomic cation channel activity / cellular response to acidic pH / extracellular ligand-gated monoatomic ion channel activity / phosphatidylinositol binding / sensory perception of pain / axon terminus / positive regulation of excitatory postsynaptic potential / sarcoplasmic reticulum / lipid metabolic process / phosphoprotein binding / microglial cell activation / cellular response to nerve growth factor stimulus / response to peptide hormone / cellular response to growth factor stimulus / GABA-ergic synapse / cellular response to tumor necrosis factor / calcium ion transmembrane transport / calcium channel activity / positive regulation of nitric oxide biosynthetic process / calcium ion transport / transmembrane signaling receptor activity / cellular response to heat / sensory perception of taste / response to heat / positive regulation of cytosolic calcium ion concentration / monoatomic ion transmembrane transport / protein homotetramerization / calmodulin binding / postsynaptic membrane / neuron projection / positive regulation of apoptotic process / external side of plasma membrane / neuronal cell body / dendrite / negative regulation of transcription by RNA polymerase II / ATP binding / membrane / metal ion binding / identical protein binding / nucleus / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.37 Å | |||||||||
 Authors Authors | Kwon DH / Zhang F | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2021 Journal: Nat Struct Mol Biol / Year: 2021Title: Heat-dependent opening of TRPV1 in the presence of capsaicin. Authors: Do Hoon Kwon / Feng Zhang / Yang Suo / Jonathan Bouvette / Mario J Borgnia / Seok-Yong Lee /  Abstract: Transient receptor potential vanilloid member 1 (TRPV1) is a Ca-permeable cation channel that serves as the primary heat and capsaicin sensor in humans. Using cryo-EM, we have determined the ...Transient receptor potential vanilloid member 1 (TRPV1) is a Ca-permeable cation channel that serves as the primary heat and capsaicin sensor in humans. Using cryo-EM, we have determined the structures of apo and capsaicin-bound full-length rat TRPV1 reconstituted into lipid nanodiscs over a range of temperatures. This has allowed us to visualize the noxious heat-induced opening of TRPV1 in the presence of capsaicin. Notably, noxious heat-dependent TRPV1 opening comprises stepwise conformational transitions. Global conformational changes across multiple subdomains of TRPV1 are followed by the rearrangement of the outer pore, leading to gate opening. Solvent-accessible surface area analyses and functional studies suggest that a subset of residues form an interaction network that is directly involved in heat sensing. Our study provides a glimpse of the molecular principles underlying noxious physical and chemical stimuli sensing by TRPV1, which can be extended to other thermal sensing ion channels. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23474.map.gz emd_23474.map.gz | 60 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23474-v30.xml emd-23474-v30.xml emd-23474.xml emd-23474.xml | 20.5 KB 20.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23474.png emd_23474.png | 26.6 KB | ||
| Filedesc metadata |  emd-23474.cif.gz emd-23474.cif.gz | 6.9 KB | ||
| Others |  emd_23474_half_map_1.map.gz emd_23474_half_map_1.map.gz emd_23474_half_map_2.map.gz emd_23474_half_map_2.map.gz | 45.9 MB 45.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23474 http://ftp.pdbj.org/pub/emdb/structures/EMD-23474 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23474 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23474 | HTTPS FTP |
-Related structure data
| Related structure data |  7lpaMC  7lp9C  7lpbC  7lpcC  7lpdC  7lpeC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23474.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23474.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
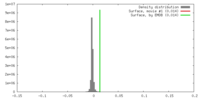
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: half map 1
| File | emd_23474_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
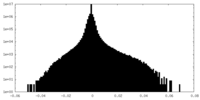
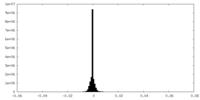
| Density Histograms |
-Half map: half map 2
| File | emd_23474_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
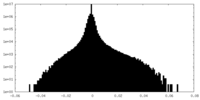
| Density Histograms |
- Sample components
Sample components
-Entire : Transient receptor potential cation channel subfamily V member 1
| Entire | Name: Transient receptor potential cation channel subfamily V member 1 |
|---|---|
| Components |
|
-Supramolecule #1: Transient receptor potential cation channel subfamily V member 1
| Supramolecule | Name: Transient receptor potential cation channel subfamily V member 1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 400 KDa |
-Macromolecule #1: Transient receptor potential cation channel subfamily V member 1
| Macromolecule | Name: Transient receptor potential cation channel subfamily V member 1 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 98.719758 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEQRASLDSE ESESPPQENS CLDPPDRDPN CKPPPVKPHI FTTRSRTRLF GKGDSEEASP LDCPYEEGGL ASCPIITVSS VLTIQRPGD GPASVRPSSQ DSVSAGEKPP RLYDRRSIFD AVAQSNCQEL ESLLPFLQRS KKRLTDSEFK DPETGKTCLL K AMLNLHNG ...String: MEQRASLDSE ESESPPQENS CLDPPDRDPN CKPPPVKPHI FTTRSRTRLF GKGDSEEASP LDCPYEEGGL ASCPIITVSS VLTIQRPGD GPASVRPSSQ DSVSAGEKPP RLYDRRSIFD AVAQSNCQEL ESLLPFLQRS KKRLTDSEFK DPETGKTCLL K AMLNLHNG QNDTIALLLD VARKTDSLKQ FVNASYTDSY YKGQTALHIA IERRNMTLVT LLVENGADVQ AAANGDFFKK TK GRPGFYF GELPLSLAAC TNQLAIVKFL LQNSWQPADI SARDSVGNTV LHALVEVADN TVDNTKFVTS MYNEILILGA KLH PTLKLE EITNRKGLTP LALAASSGKI GVLAYILQRE IHEPECRHLS RKFTEWAYGP VHSSLYDLSC IDTCEKNSVL EVIA YSSSE TPNRHDMLLV EPLNRLLQDK WDRFVKRIFY FNFFVYCLYM IIFTAAAYYR PVEGLPPYKL KNTVGDYFRV TGEIL SVSG GVYFFFRGIQ YFLQRRPSLK SLFVDSYSEI LFFVQSLFML VSVVLYFSQR KEYVASMVFS LAMGWTNMLY YTRGFQ QMG IYAVMIEKMI LRDLCRFMFV YLVFLFGFST AVVTLIEDGK NNSLPMESTP HKCRGSACKP GNSYNSLYST CLELFKF TI GMGDLEFTEN YDFKAVFIIL LLAYVILTYI LLLNMLIALM GETVNKIAQE SKNIWKLQRA ITILDTEKSF LKCMRKAF R SGKLLQVGFT PDGKDDYRWC FRVDEVNWTT WNTNVGIINE DPGNCEGVKR TLSFSLRSGR VSGRNWKNFA LVPLLRDAS TRDRHATQQE EVQLKHYTGS LKPEDAEVFK DSMVPGEKEN SLEVLFQGPD YKDDDDKAHH HHHHHHHH UniProtKB: Transient receptor potential cation channel subfamily V member 1 |
-Macromolecule #2: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine
| Macromolecule | Name: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine / type: ligand / ID: 2 / Number of copies: 8 / Formula: LBN |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-LBN: |
-Macromolecule #3: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyl...
| Macromolecule | Name: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyloxy-propan-2-yl] (~{Z})-octadec-9-enoate type: ligand / ID: 3 / Number of copies: 12 / Formula: 6OU |
|---|---|
| Molecular weight | Theoretical: 717.996 Da |
| Chemical component information |  ChemComp-6OU: |
-Macromolecule #4: (6E)-N-(4-hydroxy-3-methoxybenzyl)-8-methylnon-6-enamide
| Macromolecule | Name: (6E)-N-(4-hydroxy-3-methoxybenzyl)-8-methylnon-6-enamide type: ligand / ID: 4 / Number of copies: 4 / Formula: 4DY |
|---|---|
| Molecular weight | Theoretical: 305.412 Da |
| Chemical component information |  ChemComp-4DY: |
-Macromolecule #5: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoglycerol
| Macromolecule | Name: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoglycerol / type: ligand / ID: 5 / Number of copies: 4 / Formula: YFP |
|---|---|
| Molecular weight | Theoretical: 749.007 Da |
| Chemical component information |  ChemComp-YFP: |
-Macromolecule #6: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 6 / Number of copies: 2 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL | ||||||
|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||
| Grid | Model: Quantifoil / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 150 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa | ||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 3275 / Average exposure time: 4.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.75 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)