+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | native AMPA receptor | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Yu J / Rao P / Gouaux E | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2019 Journal: Science / Year: 2019Title: Architecture and subunit arrangement of native AMPA receptors elucidated by cryo-EM. Authors: Yan Zhao / Shanshuang Chen / Adam C Swensen / Wei-Jun Qian / Eric Gouaux /  Abstract: Glutamate-gated AMPA receptors mediate the fast component of excitatory signal transduction at chemical synapses throughout all regions of the mammalian brain. AMPA receptors are tetrameric ...Glutamate-gated AMPA receptors mediate the fast component of excitatory signal transduction at chemical synapses throughout all regions of the mammalian brain. AMPA receptors are tetrameric assemblies composed of four subunits, GluA1-GluA4. Despite decades of study, the subunit composition, subunit arrangement, and molecular structure of native AMPA receptors remain unknown. Here we elucidate the structures of 10 distinct native AMPA receptor complexes by single-particle cryo-electron microscopy (cryo-EM). We find that receptor subunits are arranged nonstochastically, with the GluA2 subunit preferentially occupying the B and D positions of the tetramer and with triheteromeric assemblies comprising a major population of native AMPA receptors. Cryo-EM maps define the structure for S2-M4 linkers between the ligand-binding and transmembrane domains, suggesting how neurotransmitter binding is coupled to ion channel gating. | |||||||||
| History |
|
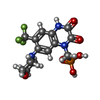
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23286.map.gz emd_23286.map.gz | 483.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23286-v30.xml emd-23286-v30.xml emd-23286.xml emd-23286.xml | 31.3 KB 31.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23286.png emd_23286.png | 101.1 KB | ||
| Others |  emd_23286_additional_1.map.gz emd_23286_additional_1.map.gz emd_23286_additional_2.map.gz emd_23286_additional_2.map.gz emd_23286_additional_3.map.gz emd_23286_additional_3.map.gz | 199.2 MB 214.9 MB 483.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23286 http://ftp.pdbj.org/pub/emdb/structures/EMD-23286 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23286 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23286 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_23286.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23286.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.00687 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_23286_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #3
| File | emd_23286_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #2
| File | emd_23286_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : GluA1/A2/A3-asymmetric-conformation2 bound to Ab fragments
+Supramolecule #1: GluA1/A2/A3-asymmetric-conformation2 bound to Ab fragments
+Supramolecule #2: GluA1/A2/A3-asymmetric-conformation2
+Supramolecule #3: 11B8 scFv
+Supramolecule #4: 15F1 Fab
+Supramolecule #5: 5B2 Fab
+Macromolecule #1: Glutamate receptor 1
+Macromolecule #2: Glutamate receptor
+Macromolecule #3: Glutamate receptor 3
+Macromolecule #4: Protein cornichon homolog 2
+Macromolecule #5: Voltage-dependent calcium channel gamma-8 subunit
+Macromolecule #6: 11B8 scFv
+Macromolecule #7: 15F1 Fab light chain
+Macromolecule #8: 15F1 Fab heavy chain
+Macromolecule #9: 5B2 Fab
+Macromolecule #12: {[7-morpholin-4-yl-2,3-dioxo-6-(trifluoromethyl)-3,4-dihydroquino...
+Macromolecule #13: 2-acetamido-2-deoxy-beta-D-glucopyranose
+Macromolecule #14: 6-[2-chloro-6-(trifluoromethoxy)phenyl]-1H-benzimidazol-2-ol
+Macromolecule #15: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 142831 |
|---|---|
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)