+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22621 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cardiac Sodium channel with toxin bound | |||||||||
 Map data Map data | cryo-EM map of sodium channel with toxin | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ion channel / Toxin bound / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationvoltage-gated sodium channel activity involved in AV node cell action potential / voltage-gated sodium channel activity involved in bundle of His cell action potential / voltage-gated sodium channel activity involved in Purkinje myocyte action potential / voltage-gated sodium channel activity involved in SA node cell action potential / bundle of His cell action potential / regulation of ventricular cardiac muscle cell membrane depolarization / AV node cell action potential / SA node cell action potential / AV node cell to bundle of His cell communication / membrane depolarization during SA node cell action potential ...voltage-gated sodium channel activity involved in AV node cell action potential / voltage-gated sodium channel activity involved in bundle of His cell action potential / voltage-gated sodium channel activity involved in Purkinje myocyte action potential / voltage-gated sodium channel activity involved in SA node cell action potential / bundle of His cell action potential / regulation of ventricular cardiac muscle cell membrane depolarization / AV node cell action potential / SA node cell action potential / AV node cell to bundle of His cell communication / membrane depolarization during SA node cell action potential / response to denervation involved in regulation of muscle adaptation / membrane depolarization during atrial cardiac muscle cell action potential / sodium channel complex / regulation of atrial cardiac muscle cell membrane repolarization / voltage-gated sodium channel activity involved in cardiac muscle cell action potential / cardiac ventricle development / brainstem development / membrane depolarization during AV node cell action potential / regulation of atrial cardiac muscle cell membrane depolarization / positive regulation of action potential / membrane depolarization during bundle of His cell action potential / atrial cardiac muscle cell action potential / membrane depolarization during Purkinje myocyte cell action potential / telencephalon development / membrane depolarization during cardiac muscle cell action potential / membrane depolarization during action potential / positive regulation of sodium ion transport / regulation of sodium ion transmembrane transport / ventricular cardiac muscle cell action potential / regulation of ventricular cardiac muscle cell membrane repolarization / cardiac muscle cell action potential involved in contraction / voltage-gated sodium channel complex / sodium channel inhibitor activity / sodium ion import across plasma membrane / regulation of cardiac muscle cell contraction / voltage-gated sodium channel activity / ankyrin binding / sodium ion transport / nitric-oxide synthase binding / odontogenesis of dentin-containing tooth / regulation of heart rate by cardiac conduction / fibroblast growth factor binding / intercalated disc / lateral plasma membrane / membrane depolarization / positive regulation of heart rate / neuronal action potential / cardiac muscle contraction / T-tubule / regulation of heart rate / cellular response to calcium ion / cerebellum development / bioluminescence / sodium ion transmembrane transport / positive regulation of epithelial cell proliferation / generation of precursor metabolites and energy / defense response / sarcolemma / caveola / Z disc / toxin activity / scaffold protein binding / transmembrane transporter binding / calmodulin binding / protein domain specific binding / axon / ubiquitin protein ligase binding / protein kinase binding / perinuclear region of cytoplasm / enzyme binding / cell surface / endoplasmic reticulum / extracellular region / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Leiurus hebraeus (scorpion) / Leiurus hebraeus (scorpion) /  | |||||||||
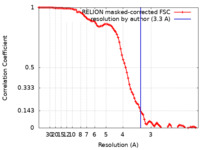
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Jiang D / Catterall WA | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Structural basis for voltage-sensor trapping of the cardiac sodium channel by a deathstalker scorpion toxin. Authors: Daohua Jiang / Lige Tonggu / Tamer M Gamal El-Din / Richard Banh / Régis Pomès / Ning Zheng / William A Catterall /   Abstract: Voltage-gated sodium (Na) channels initiate action potentials in excitable cells, and their function is altered by potent gating-modifier toxins. The α-toxin LqhIII from the deathstalker scorpion ...Voltage-gated sodium (Na) channels initiate action potentials in excitable cells, and their function is altered by potent gating-modifier toxins. The α-toxin LqhIII from the deathstalker scorpion inhibits fast inactivation of cardiac Na1.5 channels with IC = 11.4 nM. Here we reveal the structure of LqhIII bound to Na1.5 at 3.3 Å resolution by cryo-EM. LqhIII anchors on top of voltage-sensing domain IV, wedged between the S1-S2 and S3-S4 linkers, which traps the gating charges of the S4 segment in a unique intermediate-activated state stabilized by four ion-pairs. This conformational change is propagated inward to weaken binding of the fast inactivation gate and favor opening the activation gate. However, these changes do not permit Na permeation, revealing why LqhIII slows inactivation of Na channels but does not open them. Our results provide important insights into the structural basis for gating-modifier toxin binding, voltage-sensor trapping, and fast inactivation of Na channels. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22621.map.gz emd_22621.map.gz | 59.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22621-v30.xml emd-22621-v30.xml emd-22621.xml emd-22621.xml | 16.7 KB 16.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22621_fsc.xml emd_22621_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_22621.png emd_22621.png | 141.4 KB | ||
| Filedesc metadata |  emd-22621.cif.gz emd-22621.cif.gz | 7.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22621 http://ftp.pdbj.org/pub/emdb/structures/EMD-22621 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22621 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22621 | HTTPS FTP |
-Related structure data
| Related structure data |  7k18MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22621.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22621.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM map of sodium channel with toxin | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.056 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Cryo-EM map for sodium channel and scorpion toxin complex
| Entire | Name: Cryo-EM map for sodium channel and scorpion toxin complex |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM map for sodium channel and scorpion toxin complex
| Supramolecule | Name: Cryo-EM map for sodium channel and scorpion toxin complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Molecular weight | Theoretical: 250 KDa |
-Supramolecule #2: Sodium channel protein type 5 subunit alpha
| Supramolecule | Name: Sodium channel protein type 5 subunit alpha / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Alpha-like toxin Lqh3
| Supramolecule | Name: Alpha-like toxin Lqh3 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Leiurus hebraeus (scorpion) Leiurus hebraeus (scorpion) |
-Macromolecule #1: Sodium channel protein type 5 subunit alpha, Enhanced Green fluor...
| Macromolecule | Name: Sodium channel protein type 5 subunit alpha, Enhanced Green fluorescent protein type: protein_or_peptide / ID: 1 Details: Loops truncated sodium channel with EGFP and FLAG tag fused at the C-terminus Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 209.037781 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MANLLLPRGT SSFRRFTRES LAAIEKRMAE KQARGGSATS QESREGLQEE EAPRPQLDLQ ASKKLPDLYG NPPRELIGEP LEDLDPFYS TQKTFIVLNK GKTIFRFSAT NALYVLSPFH PVRRAAVKIL VHSLFSMLIM CTILTNCVFM AQHDPPPWTK Y VEYTFTAI ...String: MANLLLPRGT SSFRRFTRES LAAIEKRMAE KQARGGSATS QESREGLQEE EAPRPQLDLQ ASKKLPDLYG NPPRELIGEP LEDLDPFYS TQKTFIVLNK GKTIFRFSAT NALYVLSPFH PVRRAAVKIL VHSLFSMLIM CTILTNCVFM AQHDPPPWTK Y VEYTFTAI YTFESLVKIL ARGFCLHAFT FLRDPWNWLD FSVIVMAYTT EFVDLGNVSA LRTFRVLRAL KTISVISGLK TI VGALIQS VKKLADVMVL TVFCLSVFAL IGLQLFMGNL RHKCVRNFTE LNGTNGSVEA DGLVWNSLDV YLNDPANYLL KNG TTDVLL CGNSSDAGTC PEGYRCLKAG ENPDHGYTSF DSFAWAFLAL FRLMTQDCWE RLYQQTLRSA GKIYMIFFML VIFL GSFYL VNLILAVVAM AYEEQNQATI AETEEKEKRF QEAMEMLKKE HEALTIRGVD TVSRSSARQR ALSAVSVLTS ALEEL EESH RKCPPCWNRF AQHYLIWECC PLWMSIKQKV KFVVMDPFAD LTITMCIVLN TLFMALEHYN MTAEFEEMLQ VGNLVF TGI FTAEMTFKII ALDPYYYFQQ GWNIFDSIIV ILSLMELGLS RMGNLSVLRS FRLLRVFKLA KSWPTLNTLI KIIGNSV GA LGNLTLVLAI IVFIFAVVGM QLFGKNYSEL RHRISDSGLL PRWHMMDFFH AFLIIFRILC GEWIETMWDC MEVSGQSL C LLVFLLVMVI GNLVVLNLFL ALLLSSFSAD NLTAPDEDGE MNNLQLALAR IQRGLRFVKR TTWDFCCGIL RRRPKKPAA LATHSQLPSC ITAPRSPPPP EVEKVPPARK ETRFEEDKRP GQGTPGDSEP VCVPIAVAES DTEDQEEDEE NGKVWWRLRK TCYRIVEHS WFETFIIFMI LLSSGALAFE DIYLEERKTI KVLLEYADKM FTYVFVLEML LKWVAYGFKK YFTNAWCWLD F LIVDVSLV SLVANTLGFA EMGPIKSLRT LRALRPLRAL SRFEGMRVVV NALVGAIPSI MNVLLVCLIF WLIFSIMGVN LF AGKFGRC INQTEGDLPL NYTIVNNKSE CESFNVTGEL YWTKVKVNFD NVGAGYLALL QVATFKGWMD IMYAAVDSRG YEE QPQWED NLYMYIYFVV FIIFGSFFTL NLFIGVIIDN FNQQKKKLGG QDIFMTEEQK KYYNAMKKLG SKKPQKPIPR PLNK YQGFI FDIVTKQAFD VTIMFLICLN MVTMMVETDD QSPEKVNILA KINLLFVAIF TGECIVKMAA LRHYYFTNSW NIFDF VVVI LSIVGTVLSD IIQKYFFSPT LFRVIRLARI GRILRLIRGA KGIRTLLFAL MMSLPALFNI GLLLFLVMFI YSIFGM ANF AYVKWEAGID DMFNFQTFAN SMLCLFQITT SAGWDGLLSP ILNTGPPYCD PNLPNSNGSR GNCGSPAVGI LFFTTYI II SFLIVVNMYI AIILENFSVA TEESTEPLSE DDFDMFYEIW EKFDPEATQF IEYLALSDFA DALSEPLRIA KPNQISLI N MDLPMVSGDR IHCMDILFAF TKRVLGESGE MDALKIQMEE KFMAANPSKI SYEPITTTLE VLFQGPGSMV SKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIF FKDDGNYKTR AEVKFEGDTL VNRIELKGID FKEDGNILGH KLEYNYNSHN VYIMADKQKN GIKVNFKIRH N IEDGSVQL ADHYQQNTPI GDGPVLLPDN HYLSTQSALS KDPNEKRDHM VLLEFVTAAG ITLGMDELYK GSDYKDDDDK UniProtKB: Sodium channel protein type 5 subunit alpha, Green fluorescent protein |
-Macromolecule #2: Alpha-like toxin Lqh3
| Macromolecule | Name: Alpha-like toxin Lqh3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Leiurus hebraeus (scorpion) Leiurus hebraeus (scorpion) |
| Molecular weight | Theoretical: 7.065984 KDa |
| Sequence | String: VRDGYIAQPE NCVYHCFPGS SGCDTLCKEK GGTSGHCGFK VGHGLACWCN ALPDNVGIIV EGEKCHS UniProtKB: Alpha-like toxin Lqh3 |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #6: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyl...
| Macromolecule | Name: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyloxy-propan-2-yl] (~{Z})-octadec-9-enoate type: ligand / ID: 6 / Number of copies: 11 / Formula: 6OU |
|---|---|
| Molecular weight | Theoretical: 717.996 Da |
| Chemical component information |  ChemComp-6OU: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 6 Component:
| ||||||||||||
| Grid | Model: UltrAuFoil / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 59 |
|---|---|
| Output model |  PDB-7k18: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)