+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22363 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Drosophila ORC in the active conformation | |||||||||
 Map data Map data | unsharpened, unmasked cryo-EM map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | origin recognition complex / DNA replication initiation / REPLICATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationalpha-heterochromatin / CDC6 association with the ORC:origin complex / septin cytoskeleton / larval feeding behavior / septin cytoskeleton organization / Activation of ATR in response to replication stress / Activation of the pre-replicative complex / eggshell chorion gene amplification / Orc1 removal from chromatin / DNA amplification ...alpha-heterochromatin / CDC6 association with the ORC:origin complex / septin cytoskeleton / larval feeding behavior / septin cytoskeleton organization / Activation of ATR in response to replication stress / Activation of the pre-replicative complex / eggshell chorion gene amplification / Orc1 removal from chromatin / DNA amplification / origin recognition complex / positive regulation of border follicle cell migration / Assembly of the ORC complex at the origin of replication / nuclear origin of replication recognition complex / nuclear pre-replicative complex / olfactory learning / DNA replication preinitiation complex / mitotic chromosome condensation / mitotic DNA replication checkpoint signaling / chromosome condensation / DNA replication origin binding / DNA replication initiation / ribonucleoprotein complex binding / nuclear pore / heterochromatin / mitotic spindle organization / GTPase activator activity / learning / DNA-templated DNA replication / mitotic cell cycle / learning or memory / DNA replication / chromatin binding / protein homodimerization activity / ATP hydrolysis activity / DNA binding / ATP binding / metal ion binding / nucleus / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
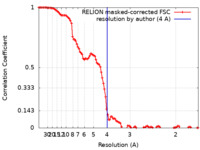
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Schmidt JM / Bleichert F | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6. Authors: Jan Marten Schmidt / Franziska Bleichert /   Abstract: Eukaryotic DNA replication initiation relies on the origin recognition complex (ORC), a DNA-binding ATPase that loads the Mcm2-7 replicative helicase onto replication origins. Here, we report cryo- ...Eukaryotic DNA replication initiation relies on the origin recognition complex (ORC), a DNA-binding ATPase that loads the Mcm2-7 replicative helicase onto replication origins. Here, we report cryo-electron microscopy (cryo-EM) structures of DNA-bound Drosophila ORC with and without the co-loader Cdc6. These structures reveal that Orc1 and Orc4 constitute the primary DNA binding site in the ORC ring and cooperate with the winged-helix domains to stabilize DNA bending. A loop region near the catalytic Walker B motif of Orc1 directly contacts DNA, allosterically coupling DNA binding to ORC's ATPase site. Correlating structural and biochemical data show that DNA sequence modulates DNA binding and remodeling by ORC, and that DNA bending promotes Mcm2-7 loading in vitro. Together, these findings explain the distinct DNA sequence-dependencies of metazoan and S. cerevisiae initiators in origin recognition and support a model in which DNA geometry and bendability contribute to Mcm2-7 loading site selection in metazoans. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22363.map.gz emd_22363.map.gz | 80.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22363-v30.xml emd-22363-v30.xml emd-22363.xml emd-22363.xml | 27.3 KB 27.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22363_fsc.xml emd_22363_fsc.xml | 10.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_22363.png emd_22363.png | 109.3 KB | ||
| Filedesc metadata |  emd-22363.cif.gz emd-22363.cif.gz | 8 KB | ||
| Others |  emd_22363_additional.map.gz emd_22363_additional.map.gz emd_22363_half_map_1.map.gz emd_22363_half_map_1.map.gz emd_22363_half_map_2.map.gz emd_22363_half_map_2.map.gz | 9.5 MB 81 MB 81 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22363 http://ftp.pdbj.org/pub/emdb/structures/EMD-22363 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22363 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22363 | HTTPS FTP |
-Related structure data
| Related structure data |  7jk6MC  7jgrC  7jgsC  7jk2C  7jk3C  7jk4C  7jk5C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22363.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22363.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened, unmasked cryo-EM map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: masked, sharpened cryo-EM map
| File | emd_22363_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | masked, sharpened cryo-EM map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: cryo-EM half map
| File | emd_22363_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: cryo-EM half map
| File | emd_22363_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Drosophila ORC (active state)
| Entire | Name: Drosophila ORC (active state) |
|---|---|
| Components |
|
-Supramolecule #1: Drosophila ORC (active state)
| Supramolecule | Name: Drosophila ORC (active state) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Origin recognition complex subunit 4
| Macromolecule | Name: Origin recognition complex subunit 4 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 52.277109 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: SNAMPEADRE LVSIRRFLKE RLQRDYTTLR GYAKERSNVR LLLQRTAEMG ESNSLLLLGP RGSGKTTLIN SVLADLLPNK SFGENTLIV HLDGNLHTDD RVALKSITVQ MQLENAADGK VFGSFAENLA FLLQCLKAGG KHSKSVIFIL EEFDLFCAHH N QTLLYNLF ...String: SNAMPEADRE LVSIRRFLKE RLQRDYTTLR GYAKERSNVR LLLQRTAEMG ESNSLLLLGP RGSGKTTLIN SVLADLLPNK SFGENTLIV HLDGNLHTDD RVALKSITVQ MQLENAADGK VFGSFAENLA FLLQCLKAGG KHSKSVIFIL EEFDLFCAHH N QTLLYNLF DVSQSAQAPI CVLGVTCRLD VIELLEKRVK SRFSHRQVFL FPSLRRFEDY VDLCRDLLSL PTGNSLLLAA EK IYNLQNI QSGALYFSRN HFDPGEYGFS PRLRDAWNKQ ICKVLATQQA RSTLQALHDF DISEAYLKNF LFRLVAHLRP QSP HITAEK MAAVGSQFEG DDKIELLCGL SVLELCLIIA IKHHSQIYDR DSFNFEIIYA RFSKFAKVST TMQAVERSIV LKAF EHLRI AELIMPLTGG AGGGVGKVQK EFEMHKLALT YSQIHHCMQR YQALPTEVAQ WAQSSLI UniProtKB: Origin recognition complex subunit 4 |
-Macromolecule #2: Origin recognition complex subunit 5
| Macromolecule | Name: Origin recognition complex subunit 5 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 52.175066 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MEAICSSLEP LFPCREAAIE TLGELIGDSS ETYPSAIYLF GHSGTGKTAL TRAFLKECGK RQNVRTAHLN AIECYTTKIM LEILLDSLA PDQGDALKVD NMLDFVEQLR RQAATRVEDQ GFLIAVDNAE RLRDMDANVL PVLLRLQELT NLNLCVILLS Q LPFEKFYN ...String: MEAICSSLEP LFPCREAAIE TLGELIGDSS ETYPSAIYLF GHSGTGKTAL TRAFLKECGK RQNVRTAHLN AIECYTTKIM LEILLDSLA PDQGDALKVD NMLDFVEQLR RQAATRVEDQ GFLIAVDNAE RLRDMDANVL PVLLRLQELT NLNLCVILLS Q LPFEKFYN KTGLSEIVCL HLAQYNKAET QRILGSDFQQ VRNQLLEQFA QDKKRLEICQ EAVTEDFYNN YLNLFLSVFY KA CRDVPEL QLTARKCLST YLEPVLDGTV DATDISRLWR HIAGPLRSAL TQIYMRIEKP AEEVEDFTAI EDQSVRKLAQ SLE LPYYAK FLLIAAFLAS HNAAKQDKRL FVKHHGKQRK RMQTVNARAK TTEKMSTTLG PKSFSIDRLL AIFYAILEEK VGLT CNLLS QISTLVHLNL LSFVSGEQNI MEGSARLQCT IGLEFVLQIG KVVGFNVRQY LCDFM UniProtKB: Origin recognition complex subunit 5 |
-Macromolecule #3: Origin recognition complex subunit 1
| Macromolecule | Name: Origin recognition complex subunit 1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 54.485633 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: SNAPRRSIHL SNIVEQRVFE DDEIISTPKR GRSKKTVQDN DEDYSPKKSV QKTPTRTRRS STTTKTATTP SKGITTATAT PMTPSQKMK KIRAGELSPS MQQRTDLPAK DSSKSELQLA REQLHVSVVP KSLPCREREF ENIYAFLEGK IQDQCGGCMY V SGVPGTGK ...String: SNAPRRSIHL SNIVEQRVFE DDEIISTPKR GRSKKTVQDN DEDYSPKKSV QKTPTRTRRS STTTKTATTP SKGITTATAT PMTPSQKMK KIRAGELSPS MQQRTDLPAK DSSKSELQLA REQLHVSVVP KSLPCREREF ENIYAFLEGK IQDQCGGCMY V SGVPGTGK TATVTGVIRT LQRMAKQNEL PAFEYLEING MRLTEPRQAY VQIYKQLTGK TVSWEQAHAL LEKRFTTPAP RR VTTVLLV DELDILCNRR QDVVYNLLDW PTKSAAKLVV VTIANTMDLP ERLLMGKVTS RLGLTRLTFQ PYSHKQLQEI VTA RLGGSE TFKGEAVQLV ARKVAAVSGD ARRALDICRR ATEIADTAAV KCVTMLHVQQ ALAEMIASAK VQAIRNCSRM EQIF LQAIA AEVTRTGVEE TTFMGVYQQV ETIAAFMGVT FPPPGRALRL CSKLGAERLI ISEHSRNDLF QKILLNVSAD DIHYA LRVE EMVN UniProtKB: Origin recognition complex subunit 1 |
-Macromolecule #4: Origin recognition complex subunit 2
| Macromolecule | Name: Origin recognition complex subunit 2 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 69.086445 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSASNKGGYK TPRKENLMSI ENLTNSEEES EDLNTAMVGN AVESQPKVTS RRSTRRPSPT KKYQAYQKES NGKGQEERIV VNYVEMSDE RSSDAEDQEE EESIEESENA ARPAAKDLHL IQSEYNVAGT SMFGFNTPKK RDAMALAALN ATPCTPKTPK T PRLGVKTP ...String: MSASNKGGYK TPRKENLMSI ENLTNSEEES EDLNTAMVGN AVESQPKVTS RRSTRRPSPT KKYQAYQKES NGKGQEERIV VNYVEMSDE RSSDAEDQEE EESIEESENA ARPAAKDLHL IQSEYNVAGT SMFGFNTPKK RDAMALAALN ATPCTPKTPK T PRLGVKTP DTKRKKSMDQ PKTPAHVRTR VKKQIAKIVA DSDEDFSGDE SDFRPSDEES SSSSSSSDAG NSSDNDAADD EP KTPSRAR RAIVVPVLPK TPSAARLRQS ARAKKSNEFV PESDGYFHSH ASSKILTSDH TLDRLKNPRL AADRVFSLLS EIK TSAEHE GSINAIMEEY RSYFPKWMCI LNEGFNILLY GLGSKHQLLQ SFHREVLHKQ TVLVVNGFFP SLTIKDMLDS ITSD ILDAG ISPANPHEAV DMIEEEFALI PETHLFLIVH NLDGAMLRNV KAQAILSRLA RIPNIHLLAS IDHINTPLLW DQGKL CSFN FSWWDCTTML PYTNETAFEN SLLVQNSGEL ALSSMRSVFS SLTTNSRGIY MLIVKYQLKN KGNATYQGMP FRDLYS SCR EAFLVSSDLA LRAQLTEFLD HKLVKSKRSV DGSEQLTIPI DGALLQQFLE EQEKK UniProtKB: Origin recognition complex subunit 2 |
-Macromolecule #5: Origin recognition complex subunit 3
| Macromolecule | Name: Origin recognition complex subunit 3 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 82.364523 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MDPTISVSKG CFVYKNGATR AGKKAASKRK RPAAESSSLL GKEVVQQPFY EEYRKAWNQI NDHIADLQHR SYARTLEQLV DFVVGQAER DTPDEVLPTA ALLTGINQPD HLSQFTALTQ RLHAQRAAMV CVLQSRDCAT LKAAVETLVF GLVEDNAEVE Q MEDEDEDE ...String: MDPTISVSKG CFVYKNGATR AGKKAASKRK RPAAESSSLL GKEVVQQPFY EEYRKAWNQI NDHIADLQHR SYARTLEQLV DFVVGQAER DTPDEVLPTA ALLTGINQPD HLSQFTALTQ RLHAQRAAMV CVLQSRDCAT LKAAVETLVF GLVEDNAEVE Q MEDEDEDE DGAERDRKRL RRSQCTMKQL KSWYTNNFDS EQKRRQLVVI LPDFECFNAS VLQDLILILS AHCGSLPFVL VL GVATAMT AVHGTLPYHV SSKIRLRVFQ TQAAPTGLNE VLDKVLLSPK YAFHLSGKTF KFLTHIFLYY DFSIHGFIQG FKY CLMEHF FGGNAFALCT DYSKALGRIK QLTHEDMETI RRLPSFRPYV EQINDCKRII AVLTDDDYLK KKLPQLLRDC LLHF LLFRC SLEFLTELVG DLPRCPLGKL RRELYVNCLN RAIISTPEYK ECLQMLSFLS KDEFVAKVNR ALERTEQFLV EEIAP LELG EACTAVLRPK LEAIRLAVDE VVKATMATIT TTSPNETRQA TDHLTPVASR QELKDQLLQR SKEDKMRHQL NTPTTQ FGR ALQKTLQLIE TQIVQDHLRA LQDAPPIHEL FVFSDIATVR RNIIGAPRAA LHTALNNPHF YMQCKCCELQ DQSLLVG TL PDLSVVYKLH LECGRMINLF DWLQAFRSVV SDSDHEEVAQ EQIDPQIQAR FTRAVAELQF LGYIKMSKRK TDHATRLT W UniProtKB: Origin recognition complex subunit 3 |
-Macromolecule #6: Origin recognition complex subunit 6
| Macromolecule | Name: Origin recognition complex subunit 6 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 29.284129 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MTTLIEQLIT KMGLREEPNV LEKTTELVRL LELRSTNVPL QINEYGKIVL CADLASCMIG IAFDKEQALK LSGLRKSQYL NNKRMFEKL LDLNKLASVN DICVQLGLNE VARKAEELMT LFKGVAATED MGTDTSHPQY ATMAVFQACR LLKKKVSKSK L MPFSNLRP ...String: MTTLIEQLIT KMGLREEPNV LEKTTELVRL LELRSTNVPL QINEYGKIVL CADLASCMIG IAFDKEQALK LSGLRKSQYL NNKRMFEKL LDLNKLASVN DICVQLGLNE VARKAEELMT LFKGVAATED MGTDTSHPQY ATMAVFQACR LLKKKVSKSK L MPFSNLRP SQFQLLEQQW ERMIAKHHKE SKVPSSTDME GKLKENQNEN IKGHEAKKAH KPPPEDYEIW KARMLAKAQA KL KELEASQ SHMDSQLLEA UniProtKB: Origin recognition complex subunit 6 |
-Macromolecule #7: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 7 / Number of copies: 3 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #8: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 8 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil, UltrAuFoil, R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 10 sec. / Pretreatment - Atmosphere: OTHER | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | The complex used for grid preparation was reconstituted with DNA. This structure was obtained from a subset of particles not bound to DNA. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)