[English] 日本語
 Yorodumi
Yorodumi- EMDB-2157: Helical structures of WHAMM around MTs revealed by Electron cryo-... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2157 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Helical structures of WHAMM around MTs revealed by Electron cryo-microscopy | |||||||||
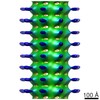
 Map data Map data | Reconstruction of WHAMM with N-terminal MBP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | actin nucleation / membrane tubulation / microtubules / WHAMM | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 18.0 Å | |||||||||
 Authors Authors | Shen QT / Hsiue PP / Sindelar CV / Welch MD / Campellone KG / Wang HW | |||||||||
 Citation Citation |  Journal: J Cell Biol / Year: 2012 Journal: J Cell Biol / Year: 2012Title: Structural insights into WHAMM-mediated cytoskeletal coordination during membrane remodeling. Authors: Qing-Tao Shen / Peter P Hsiue / Charles V Sindelar / Matthew D Welch / Kenneth G Campellone / Hong-Wei Wang /  Abstract: The microtubule (MT) and actin cytoskeletons drive many essential cellular processes, yet fairly little is known about how their functions are coordinated. One factor that mediates important cross ...The microtubule (MT) and actin cytoskeletons drive many essential cellular processes, yet fairly little is known about how their functions are coordinated. One factor that mediates important cross talk between these two systems is WHAMM, a Golgi-associated protein that utilizes MT binding and actin nucleation activities to promote membrane tubulation during intracellular transport. Using cryoelectron microscopy and other biophysical and biochemical approaches, we unveil the underlying mechanisms for how these activities are coordinated. We find that WHAMM bound to the outer surface of MT protofilaments via a novel interaction between its central coiled-coil region and tubulin heterodimers. Upon the assembly of WHAMM onto MTs, its N-terminal membrane-binding domain was exposed at the MT periphery, where it can recruit vesicles and remodel them into tubular structures. In contrast, MT binding masked the C-terminal portion of WHAMM and prevented it from promoting actin nucleation. These results give rise to a model whereby distinct MT-bound and actin-nucleating populations of WHAMM collaborate during membrane tubulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2157.map.gz emd_2157.map.gz | 26.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2157-v30.xml emd-2157-v30.xml emd-2157.xml emd-2157.xml | 9.5 KB 9.5 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2157.png EMD-2157.png | 178.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2157 http://ftp.pdbj.org/pub/emdb/structures/EMD-2157 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2157 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2157 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2157.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2157.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of WHAMM with N-terminal MBP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : WHAMM around 13-pf MTs
| Entire | Name: WHAMM around 13-pf MTs |
|---|---|
| Components |
|
-Supramolecule #1000: WHAMM around 13-pf MTs
| Supramolecule | Name: WHAMM around 13-pf MTs / type: sample / ID: 1000 / Number unique components: 117 |
|---|
-Macromolecule #1: WHAMM around MTs
| Macromolecule | Name: WHAMM around MTs / type: protein_or_peptide / ID: 1 / Number of copies: 17 / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Strain: Hela / synonym: Human Homo sapiens (human) / Strain: Hela / synonym: Human |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 / Details: 80mM PIPES pH 6.8, 1mM MgCl2, 1mM EGTA, 1mM DTT |
|---|---|
| Grid | Details: 300 mesh holly carbon grid, glow discharged. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 120 K / Instrument: GATAN CRYOPLUNGE 3 / Method: Blot for 2.5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Min: 88 K / Max: 101 K / Average: 95 K |
| Date | Nov 16, 2009 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 77 / Average electron dose: 15 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 34000 / Illumination mode: OTHER / Imaging mode: DIFFRACTION / Cs: 2.2 mm / Nominal defocus max: 0.003 µm / Nominal defocus min: 0.0015 µm / Nominal magnification: 29000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 18.0 Å / Resolution method: OTHER / Software - Name: Spider, EMAN |
|---|---|
| CTF correction | Details: Each particle |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - #0 - Chain ID: A / Chain - #1 - Chain ID: B |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | Protocol: Rigid body |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















