+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21005 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Negative stain EM map of CODH/ACS in the closed conformation | |||||||||||||||
 Map data Map data | Negative stain map of CODH/ACS in the closed conformation | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |  Moorella thermoacetica (bacteria) Moorella thermoacetica (bacteria) | |||||||||||||||
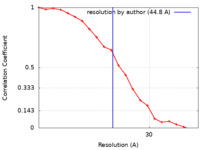
| Method | single particle reconstruction / negative staining / Resolution: 44.8 Å | |||||||||||||||
 Authors Authors | Cohen SE / Brignole EJ / Drennan CL | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Structure / Year: 2021 Journal: Structure / Year: 2021Title: Negative-Stain Electron Microscopy Reveals Dramatic Structural Rearrangements in Ni-Fe-S-Dependent Carbon Monoxide Dehydrogenase/Acetyl-CoA Synthase. Authors: Steven E Cohen / Edward J Brignole / Elizabeth C Wittenborn / Mehmet Can / Samuel Thompson / Stephen W Ragsdale / Catherine L Drennan /  Abstract: The Ni-Fe-S-containing A-cluster of acetyl-coenzyme A (CoA) synthase (ACS) assembles acetyl-CoA from carbon monoxide (CO), a methyl group (CH), and CoA. To accomplish this feat, ACS must bind CoA and ...The Ni-Fe-S-containing A-cluster of acetyl-coenzyme A (CoA) synthase (ACS) assembles acetyl-CoA from carbon monoxide (CO), a methyl group (CH), and CoA. To accomplish this feat, ACS must bind CoA and interact with two other proteins that contribute the CO and CH, respectively: CO dehydrogenase (CODH) and corrinoid Fe-S protein (CFeSP). Previous structural data show that, in the model acetogen Moorella thermoacetica, domain 1 of ACS binds to CODH such that a 70-Å-long internal channel is created that allows CO to travel from CODH to the A-cluster. The A-cluster is largely buried and is inaccessible to CFeSP for methylation. Here we use electron microscopy to capture multiple snapshots of ACS that reveal previously uncharacterized domain motion, forming extended and hyperextended structural states. In these structural states, the A-cluster is accessible for methylation by CFeSP. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21005.map.gz emd_21005.map.gz | 953 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21005-v30.xml emd-21005-v30.xml emd-21005.xml emd-21005.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21005_fsc.xml emd_21005_fsc.xml | 1.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_21005.png emd_21005.png | 29.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21005 http://ftp.pdbj.org/pub/emdb/structures/EMD-21005 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21005 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21005 | HTTPS FTP |
-Validation report
| Summary document |  emd_21005_validation.pdf.gz emd_21005_validation.pdf.gz | 77.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21005_full_validation.pdf.gz emd_21005_full_validation.pdf.gz | 76.9 KB | Display | |
| Data in XML |  emd_21005_validation.xml.gz emd_21005_validation.xml.gz | 495 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21005 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21005 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21005 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21005 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21005.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21005.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Negative stain map of CODH/ACS in the closed conformation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : carbon monoxide dheydrogenase/acetyl-CoA synthase tetramer
| Entire | Name: carbon monoxide dheydrogenase/acetyl-CoA synthase tetramer |
|---|---|
| Components |
|
-Supramolecule #1: carbon monoxide dheydrogenase/acetyl-CoA synthase tetramer
| Supramolecule | Name: carbon monoxide dheydrogenase/acetyl-CoA synthase tetramer type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Moorella thermoacetica (bacteria) Moorella thermoacetica (bacteria) |
-Macromolecule #1: Carbon monoxide dehdyrogenase
| Macromolecule | Name: Carbon monoxide dehdyrogenase / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO / EC number: anaerobic carbon monoxide dehydrogenase |
|---|---|
| Source (natural) | Organism:  Moorella thermoacetica (bacteria) Moorella thermoacetica (bacteria) |
| Sequence | String: MPRFRDLSHN CRPSEAPRVM EPKNRDRTVD PAVLEMLVKS KDDKVITAFD RFVAQQPQCK IGYEGICCR FCMAGPCRIK ATDGPGSRGI CGASAWTIVA RNVGLMILTG AAAHCEHGNH I AHALVEMA EGKAPDYSVK DEAKLKEVCR RVGIEVEGKS VLELAQEVGE ...String: MPRFRDLSHN CRPSEAPRVM EPKNRDRTVD PAVLEMLVKS KDDKVITAFD RFVAQQPQCK IGYEGICCR FCMAGPCRIK ATDGPGSRGI CGASAWTIVA RNVGLMILTG AAAHCEHGNH I AHALVEMA EGKAPDYSVK DEAKLKEVCR RVGIEVEGKS VLELAQEVGE KALEDFRRLK GE GEATWLM TTINEGRKEK FRTHNVVPFG IHASISELVN QAHMGMDNDP VNLVFSAIRV ALA DYTGEH IATDFSDILF GTPQPVVSEA NMGVLDPDQV NFVLHGHNPL LSEIIVQAAR EMEG EAKAA GAKGINLVGI CCTGNEVLMR QGIPLVTSFA SQELAICTGA IDAMCVDVQC IMPSI SAVA ECYHTRIITT ADNAKIPGAY HIDYQTATAI ESAKTAIRMA IEAFKERKES NRPVYI PQI KNRVVAGWSL EALTKLLATQ NAQNPIRVLN QAILDGELAG VALICGCNNL KGFQDNS HL TVMKELLKNN VFVVATGCSA QAAGKLGLLD PANVETYCGD GLKGFLKRLG EGANIEIG L PPVFHMGSCV DNSRAVDLLM AMANDLGVDT PKVPFVASAP EAMSGKAAAI GTWWVSLGV PTHVGTMPPV EGSDLIYSIL TQIASDVYGG YFIFEMDPQV AARKILDALE YRTWKLGVHK EVAERYETK LCQGY |
-Macromolecule #2: Acetyl-CoA synthase
| Macromolecule | Name: Acetyl-CoA synthase / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO / EC number: CO-methylating acetyl-CoA synthase |
|---|---|
| Source (natural) | Organism:  Moorella thermoacetica (bacteria) Moorella thermoacetica (bacteria) |
| Sequence | String: MTDFDKIFEG AIPEGKEPVA LFREVYHGAI TATSYAEILL NQAIRTYGPD HPVGYPDTAY YLPVIRCFS GEEVKKLGDL PPILNRKRAQ VSPVLNFENA RLAGEATWYA AEIIEALRYL K YKPDEPLL PPPWTGFIGD PVVRRFGIKM VDWTIPGEAI ILGRAKDSKA ...String: MTDFDKIFEG AIPEGKEPVA LFREVYHGAI TATSYAEILL NQAIRTYGPD HPVGYPDTAY YLPVIRCFS GEEVKKLGDL PPILNRKRAQ VSPVLNFENA RLAGEATWYA AEIIEALRYL K YKPDEPLL PPPWTGFIGD PVVRRFGIKM VDWTIPGEAI ILGRAKDSKA LAKIVKELMG MG FMLFICD EAVEQLLEEN VKLGIDYIAY PLGNFTQIVH AANYALRAGM MFGGVTPGAR EEQ RDYQRR RIRAFVLYLG EHDMVKTAAA FGAIFTGFPV ITDQPLPEDK QIPDWFFSVE DYDK IVQIA METRGIKLTK IKLDLPINFG PAFEGESIRK GDMYVEMGGN RTPAFELVRT VSESE ITDG KIEVIGPDID QIPEGSKLPL GILVDIYGRK MQADFEGVLE RRIHDFINYG EGLWHT GQR NINWLRVSKD AVAKGFRFKN YGEILVAKMK EEFPAIVDRV QVTIFTDEAK VKEYMEV AR EKYKERDDRM RGLTDETVDT FYSCVLCQSF APNHVCIVTP ERVGLCGAVS WLDAKASY E INHAGPNQPI PKEGEIDPIK GIWKSVNDYL YTASNRNLEQ VCLYTLMENP MTSCGCFEA IMAILPECNG IMITTRDHAG MTPSGMTFST LAGMIGGGTQ TPGFMGIGRT YIVSKKFISA DGGIARIVW MPKSLKDFLH DEFVRRSVEE GLGEDFIDKI ADETIGTTVD EILPYLEEKG H PALTMDPI M |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | .017 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.6 Component:
| |||||||||
| Staining | Type: NEGATIVE / Material: uranyl acetate Details: Samples were prepared by blotting protein from EM grid using filter paper, followed by multiple rounds of staining and blotting. | |||||||||
| Grid | Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.038 kPa / Details: -15 mA for 60 seconds | |||||||||
| Details | The sample was predominantly monodisperse, as seen by negative-stain TEM. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number grids imaged: 1 / Average exposure time: 1.0 sec. / Average electron dose: 23.0 e/Å2 Details: Images were collected as tilt pairs at 0 and 55 degrees. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal magnification: 62000 |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)