[English] 日本語
 Yorodumi
Yorodumi- EMDB-20713: In vivo structure of the Legionella type II secretion system by e... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20713 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | In vivo structure of the Legionella type II secretion system by electron cryotomography (aligning the OM-complex). | |||||||||
 Map data Map data | Subtomogram average of the Legionella pneumophila type II secretion system (aligning the OM-complex). | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 40.0 Å | |||||||||
 Authors Authors | Ghosal D / Jensen GJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2019 Journal: Nat Microbiol / Year: 2019Title: In vivo structure of the Legionella type II secretion system by electron cryotomography. Authors: Debnath Ghosal / Ki Woo Kim / Huaixin Zheng / Mohammed Kaplan / Hilary K Truchan / Alberto E Lopez / Ian E McIntire / Joseph P Vogel / Nicholas P Cianciotto / Grant J Jensen /    Abstract: The type II secretion system (T2SS) is a multiprotein envelope-spanning assembly that translocates a wide range of virulence factors, enzymes and effectors through the outer membrane of many Gram- ...The type II secretion system (T2SS) is a multiprotein envelope-spanning assembly that translocates a wide range of virulence factors, enzymes and effectors through the outer membrane of many Gram-negative bacteria. Here, using electron cryotomography and subtomogram averaging methods, we reveal the in vivo structure of an intact T2SS imaged within the human pathogen Legionella pneumophila. Although the T2SS has only limited sequence and component homology with the evolutionarily related type IV pilus (T4P) system, we show that their overall architectures are remarkably similar. Despite similarities, there are also differences, including, for example, that the T2SS-ATPase complex is usually present but disengaged from the inner membrane, the T2SS has a much longer periplasmic vestibule and it has a short-lived flexible pseudopilus. Placing atomic models of the components into our electron cryotomography map produced a complete architectural model of the intact T2SS that provides insights into the structure and function of its components, its position within the cell envelope and the interactions between its different subcomplexes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20713.map.gz emd_20713.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20713-v30.xml emd-20713-v30.xml emd-20713.xml emd-20713.xml | 8.7 KB 8.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20713.png emd_20713.png | 68.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20713 http://ftp.pdbj.org/pub/emdb/structures/EMD-20713 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20713 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20713 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20713.map.gz / Format: CCP4 / Size: 2.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20713.map.gz / Format: CCP4 / Size: 2.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of the Legionella pneumophila type II secretion system (aligning the OM-complex). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 8.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
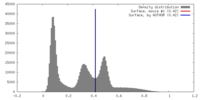
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Subtomogram average of the Legionella pneumophila type II secreti...
| Entire | Name: Subtomogram average of the Legionella pneumophila type II secretion system (aligning the OM-complex). |
|---|---|
| Components |
|
-Supramolecule #1: Subtomogram average of the Legionella pneumophila type II secreti...
| Supramolecule | Name: Subtomogram average of the Legionella pneumophila type II secretion system (aligning the OM-complex). type: complex / ID: 1 / Parent: 0 Details: We used electron cryotomography and subtomogram averaging to visualize L. pneumophila type II secretion system complex directly in intact, frozen-hydrated bacterial cells |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 6.9 |
|---|---|
| Grid | Material: COPPER |
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 40.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name:  IMOD / Number subtomograms used: 440 IMOD / Number subtomograms used: 440 |
|---|---|
| Extraction | Number tomograms: 1993 / Number images used: 440 / Software - Name: PEET |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)