+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20532 | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Toll receptor EM map from Topaz picks | ||||||||||||||||||||||||||||||
 Map data Map data | Sharpened map after CryoSparc 2.9 non-uniform refinement using DoG picks | ||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||
| Biological species |  | ||||||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | ||||||||||||||||||||||||||||||
 Authors Authors | Rapp M / Bepler T / Morin A / Brasch J / Shapiro L / Noble AJ / Berger B | ||||||||||||||||||||||||||||||
| Funding support |  United States, 9 items United States, 9 items
| ||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2019 Journal: Nat Methods / Year: 2019Title: Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Authors: Tristan Bepler / Andrew Morin / Micah Rapp / Julia Brasch / Lawrence Shapiro / Alex J Noble / Bonnie Berger /  Abstract: Cryo-electron microscopy is a popular method for the determination of protein structures; however, identifying a sufficient number of particles for analysis can take months of manual effort. Current ...Cryo-electron microscopy is a popular method for the determination of protein structures; however, identifying a sufficient number of particles for analysis can take months of manual effort. Current computational approaches find many false positives and require ad hoc postprocessing, especially for unusually shaped particles. To address these shortcomings, we develop Topaz, an efficient and accurate particle-picking pipeline using neural networks trained with a general-purpose positive-unlabeled learning method. This framework enables particle detection models to be trained with few sparsely labeled particles and no labeled negatives. Topaz retrieves many more real particles than conventional picking methods while maintaining low false-positive rates, is capable of picking challenging unusually shaped proteins (for example, small, non-globular and asymmetric particles), produces more representative particle sets and does not require post hoc curation. We demonstrate the performance of Topaz on two difficult datasets and three conventional datasets. Topaz is modular, standalone, free and open source ( http://topaz.csail.mit.edu ). | ||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20532.map.gz emd_20532.map.gz | 117.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20532-v30.xml emd-20532-v30.xml emd-20532.xml emd-20532.xml | 19.3 KB 19.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20532.png emd_20532.png | 39.6 KB | ||
| Masks |  emd_20532_msk_1.map emd_20532_msk_1.map | 125 MB |  Mask map Mask map | |
| Others |  emd_20532_additional.map.gz emd_20532_additional.map.gz emd_20532_additional_1.map.gz emd_20532_additional_1.map.gz emd_20532_half_map_1.map.gz emd_20532_half_map_1.map.gz emd_20532_half_map_2.map.gz emd_20532_half_map_2.map.gz | 62 MB 62 MB 116.2 MB 116.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20532 http://ftp.pdbj.org/pub/emdb/structures/EMD-20532 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20532 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20532 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10215 (Title: Rabbit muscle aldolase single particle cryoEM / Data size: 1.4 TB EMPIAR-10215 (Title: Rabbit muscle aldolase single particle cryoEM / Data size: 1.4 TBData #1: Micrographs along with all other magnification images from the collection [micrographs - single frame] Data #2: Micrograph frames [micrographs - multiframe] Data #3: CryoEM structures in the paper [picked particles - single frame - unprocessed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20532.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20532.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map after CryoSparc 2.9 non-uniform refinement using DoG picks | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_20532_msk_1.map emd_20532_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
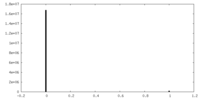
| Density Histograms |
-Additional map: Unsharpened map after CryoSparc 2.9 non-uniform refinement using...
| File | emd_20532_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map after CryoSparc 2.9 non-uniform refinement using DoG picks | ||||||||||||
| Projections & Slices |
| ||||||||||||
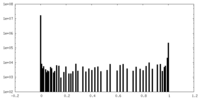
| Density Histograms |
-Additional map: Unsharpened map after CryoSparc 2.9 non-uniform refinement using...
| File | emd_20532_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map after CryoSparc 2.9 non-uniform refinement using DoG picks | ||||||||||||
| Projections & Slices |
| ||||||||||||
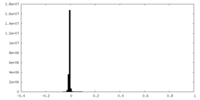
| Density Histograms |
-Half map: Half map after CryoSparc 2.9 non-uniform refinement using DoG picks
| File | emd_20532_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map after CryoSparc 2.9 non-uniform refinement using DoG picks | ||||||||||||
| Projections & Slices |
| ||||||||||||
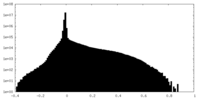
| Density Histograms |
-Half map: Half map after CryoSparc 2.9 non-uniform refinement using DoG picks
| File | emd_20532_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map after CryoSparc 2.9 non-uniform refinement using DoG picks | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Toll receptor
| Entire | Name: Toll receptor |
|---|---|
| Components |
|
-Supramolecule #1: Toll receptor
| Supramolecule | Name: Toll receptor / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 105 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Average electron dose: 73.48 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)