+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20393 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of wild type mediator with MED25-FLAG | |||||||||
 Map data Map data | em-volume_P1 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.49 Å | |||||||||
 Authors Authors | Asturias F / Zhao H | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2019 Journal: Cell / Year: 2019Title: A Pliable Mediator Acts as a Functional Rather Than an Architectural Bridge between Promoters and Enhancers. Authors: Laila El Khattabi / Haiyan Zhao / Jens Kalchschmidt / Natalie Young / Seolkyoung Jung / Peter Van Blerkom / Philippe Kieffer-Kwon / Kyong-Rim Kieffer-Kwon / Solji Park / Xiang Wang / Jordan ...Authors: Laila El Khattabi / Haiyan Zhao / Jens Kalchschmidt / Natalie Young / Seolkyoung Jung / Peter Van Blerkom / Philippe Kieffer-Kwon / Kyong-Rim Kieffer-Kwon / Solji Park / Xiang Wang / Jordan Krebs / Subhash Tripathi / Noboru Sakabe / Débora R Sobreira / Su-Chen Huang / Suhas S P Rao / Nathanael Pruett / Daniel Chauss / Erica Sadler / Andrea Lopez / Marcelo A Nóbrega / Erez Lieberman Aiden / Francisco J Asturias / Rafael Casellas /   Abstract: While Mediator plays a key role in eukaryotic transcription, little is known about its mechanism of action. This study combines CRISPR-Cas9 genetic screens, degron assays, Hi-C, and cryoelectron ...While Mediator plays a key role in eukaryotic transcription, little is known about its mechanism of action. This study combines CRISPR-Cas9 genetic screens, degron assays, Hi-C, and cryoelectron microscopy (cryo-EM) to dissect the function and structure of mammalian Mediator (mMED). Deletion analyses in B, T, and embryonic stem cells (ESC) identified a core of essential subunits required for Pol II recruitment genome-wide. Conversely, loss of non-essential subunits mostly affects promoters linked to multiple enhancers. Contrary to current models, however, mMED and Pol II are dispensable to physically tether regulatory DNA, a topological activity requiring architectural proteins. Cryo-EM analysis revealed a conserved core, with non-essential subunits increasing structural complexity of the tail module, a primary transcription factor target. Changes in tail structure markedly increase Pol II and kinase module interactions. We propose that Mediator's structural pliability enables it to integrate and transmit regulatory signals and act as a functional, rather than an architectural bridge, between promoters and enhancers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20393.map.gz emd_20393.map.gz | 632.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20393-v30.xml emd-20393-v30.xml emd-20393.xml emd-20393.xml | 15.2 KB 15.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20393_fsc.xml emd_20393_fsc.xml | 20 KB | Display |  FSC data file FSC data file |
| Images |  emd_20393.png emd_20393.png | 73.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20393 http://ftp.pdbj.org/pub/emdb/structures/EMD-20393 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20393 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20393 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20393.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20393.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | em-volume_P1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
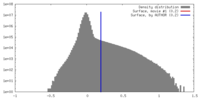
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 30 proteins complex
| Entire | Name: 30 proteins complex |
|---|---|
| Components |
|
-Supramolecule #1: 30 proteins complex
| Supramolecule | Name: 30 proteins complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 1.0 MDa |
-Macromolecule #1: protein complex
| Macromolecule | Name: protein complex / type: other / ID: 1 / Classification: other |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MAPVQLDNHQ LIPPGGGGGS SGGGGSSSGS ASAPAPPPPA AAVAAAAAAA ASPGYRLSTL IEFLLHRAY SELMVLTDLL PRKSDVERKI EIVQFASRTR QLFVRLLALV KWANDAGKVE K CAMISSFL DQQAILFVDT ADRLASLARD ALVHARLPSF AIPYAIDVLT ...String: MAPVQLDNHQ LIPPGGGGGS SGGGGSSSGS ASAPAPPPPA AAVAAAAAAA ASPGYRLSTL IEFLLHRAY SELMVLTDLL PRKSDVERKI EIVQFASRTR QLFVRLLALV KWANDAGKVE K CAMISSFL DQQAILFVDT ADRLASLARD ALVHARLPSF AIPYAIDVLT TGSYPRLPTC IR DKIIPPD PITKIEKQAT LHQLNQILRH RLVTTDLPPQ LANLTVANGR VKFRVEGEFE ATL TVMGDD PEVPWRLLKL EILVEDKETG DGRALVHSMQ IDFIHQLVQS RLFADEKPLQ DMYN CLHCF CLSLQLEVLH SQTLMLIRER WGDLVQVERY HAGKSLSLSV WNQQVLGRKT GTASV HKVT IKIDENDVSK PLQIFHDPPL PASDSKLVER AMKIDHLSIE KLLIDSVHAR AHQRLQ ELK AILRSFNANE SSSIETALPA LIVPILEPCG NSECLHIFVD LHSGMFQLML YGLDPAT LE DMEKSLNDDM KRIIPWIQQL KFWLGQQRCK QSIKHLPTIT TETLQLANYS THPIGSLS K NKLFIKLTRL PQYYIVVEML EVPNKPTQLS YNYYFMSVST ADREDSPVMA LLLQQFKDN IQDLMSYTKT GKQTRTGTKH KLSDDPCPID SKKAKRSGEM CAFNKVLAHF VAMCDTNMPF VGLRLELSN LEIPHQGVQV EGDGFNHAIR LLKIPPCKGI SEETQKALDR SLLDCTFRLQ G RNNRTWVA ELVFANCPLN GTSTREQGPS RHVYLTYENL LSEPVGGRKV VEMFLNDWSS IA RLYECVL EFARSLPEIP AHLNIFSEVR VYNYRKLILC YGTTKGSSIS IQWNSIHQKF HIA LGTVGP NSGCSNCHNT ILHQLQEMFN KTPNVVQLLQ VLFDTQAPLN AINKLPTVPM LGLT QRTNT AYQCFSILPQ SSTHIRLAFR NMYCIDIYCR SRGVVAIRDG AYSLFDNSKL VEGFY PAPG LKTFLNMFVD SNQDARRRSV NEDDNPPSPI GGDMMDSLIS QLQPPQQQPF PKQPGT SGA YPLTSPPTSY HSTVNQSPSM MHTQSPGNLH AASSPSGALR APSPASFVPT PPPSSHG IS IGPGASFASP HGTLDPSSPY TMVSPSGRAG NWPGSPQVSG PSPATRLPGM SPANPSLH S PVPDVSHSPR AGTSSQTMPT NMPPPRKLPQ RSWAASIPTI LTHSALNILL LPSPTPGLV PGLAGSYLCS PLERFLGSVI MRRHLQRIIQ QETLQLINSN EPGVIMFKTD ALKCRVALSP KTNQTLQLK VTPENAGQWK PDELQVLEKF FETRVAGPPF KANTLIAFTK LLGAPTHILR D CVHIMKLE LFPDQATQLK WNVQFCLTIP PSAPPIAPPG TPAVVLKSKM LFFLQLTQKT SV PPQEPVS IIVPIIYDMA SGTTQQADIP RQQNSSVAAP MMVSNILKRF AEMNPPRQGE CTI FAAVRD LMANLTLPPG GRP |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.9 / Component - Concentration: 50.0 mM / Component - Formula: C8H18N2O4S / Component - Name: HEPES |
| Grid | Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 3.0 nm / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER / Pretreatment - Pressure: 101.325 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 273 K / Instrument: FEI VITROBOT MARK II |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Digitization - Frames/image: 32-40 / Number grids imaged: 1 / Number real images: 3854 / Average exposure time: 3.0 sec. / Average electron dose: 53.9 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 5.0 µm / Calibrated defocus min: 1.2 µm / Calibrated magnification: 22000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 22000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)