[English] 日本語
 Yorodumi
Yorodumi- EMDB-20134: Structure of a drug-like molecule stalled PCSK9 ribosome nascent ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20134 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
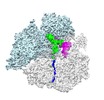
| Title | Structure of a drug-like molecule stalled PCSK9 ribosome nascent chain complex (PCSK9-RNC) with AP tRNA and PE tRNA (sample prepared with a short incubation time) | |||||||||
 Map data Map data | Structure of a drug-like molecule stalled PCSK9 ribosome nascent chain complex (PCSK9-RNC) with AP tRNA and PE tRNA (sample prepared with a short incubation time) | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.7 Å | |||||||||
 Authors Authors | Li W / Cate JHD | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2019 Journal: Nat Struct Mol Biol / Year: 2019Title: Structural basis for selective stalling of human ribosome nascent chain complexes by a drug-like molecule. Authors: Wenfei Li / Fred R Ward / Kim F McClure / Stacey Tsai-Lan Chang / Elizabeth Montabana / Spiros Liras / Robert G Dullea / Jamie H D Cate /  Abstract: The drug-like molecule PF-06446846 (PF846) binds the human ribosome and selectively blocks the translation of a small number of proteins by an unknown mechanism. In structures of PF846-stalled human ...The drug-like molecule PF-06446846 (PF846) binds the human ribosome and selectively blocks the translation of a small number of proteins by an unknown mechanism. In structures of PF846-stalled human ribosome nascent chain complexes, PF846 binds in the ribosome exit tunnel in a eukaryotic-specific pocket formed by 28S ribosomal RNA, and alters the path of the nascent polypeptide chain. PF846 arrests the translating ribosome in the rotated state of translocation, in which the peptidyl-transfer RNA 3'-CCA end is improperly docked in the peptidyl transferase center. Selections of messenger RNAs from mRNA libraries using translation extracts reveal that PF846 can stall translation elongation, arrest termination or even enhance translation, depending on nascent chain sequence context. These results illuminate how a small molecule selectively targets translation by the human ribosome, and provides a foundation for developing small molecules that modulate the production of proteins of therapeutic interest. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20134.map.gz emd_20134.map.gz | 210 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20134-v30.xml emd-20134-v30.xml emd-20134.xml emd-20134.xml | 9.2 KB 9.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20134.png emd_20134.png | 183.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20134 http://ftp.pdbj.org/pub/emdb/structures/EMD-20134 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20134 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20134 | HTTPS FTP |
-Related structure data
| Related structure data |  0526C  0596C  0597C  0598C  0599C  0600C  0601C  6oleC  6olfC  6olgC  6oliC  6olzC  6om0C  6om7C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20134.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20134.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of a drug-like molecule stalled PCSK9 ribosome nascent chain complex (PCSK9-RNC) with AP tRNA and PE tRNA (sample prepared with a short incubation time) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.22 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Structure of a drug-like molecule stalled PCSK9 ribosome nascent ...
| Entire | Name: Structure of a drug-like molecule stalled PCSK9 ribosome nascent chain complex (PCSK9-RNC) with AP tRNA and PE tRNA (sample prepared with a short incubation time) |
|---|---|
| Components |
|
-Supramolecule #1: Structure of a drug-like molecule stalled PCSK9 ribosome nascent ...
| Supramolecule | Name: Structure of a drug-like molecule stalled PCSK9 ribosome nascent chain complex (PCSK9-RNC) with AP tRNA and PE tRNA (sample prepared with a short incubation time) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HeLa Homo sapiens (human) / Recombinant cell: HeLa |
| Molecular weight | Theoretical: 4.3 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 4 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.7 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 8433 |
|---|---|
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
-Atomic model buiding 1
| Refinement | Space: REAL / Overall B value: 75 |
|---|
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















