[English] 日本語
 Yorodumi
Yorodumi- EMDB-1890: EcoR124 Type I DNA restriction-modification enzyme complex in clo... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1890 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | EcoR124 Type I DNA restriction-modification enzyme complex in closed state with bound 30bp cognate DNA fragment. 3D reconstruction by single particle analysis from negative stain EM. | |||||||||
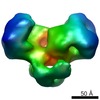
 Map data Map data | EcoR124I R2 M2 S1 complex with 30bp DNA fragment bound (closed state). DNA is not visible in the negative stain images or map. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | EcoR124 / endonuclease / methyltransferase / type I restriction / DNA / HsdS / HsdM / HsdR / electron microscopy / negative stain / translocase / DEAD-box / ATPase | |||||||||
| Function / homology | Restriction endonuclease, type I, HsdR / Type I restriction modification DNA specificity domain / DNA methylase, adenine-specific / protein binding / DNA restriction-modification system Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 21.0 Å | |||||||||
 Authors Authors | Kennaway CK / Taylor JE / Song CF / Potrzebowski W / White JH / Swiderska A / Obarska-Kosinska A / Callow P / Cooper LP / Roberts GA ...Kennaway CK / Taylor JE / Song CF / Potrzebowski W / White JH / Swiderska A / Obarska-Kosinska A / Callow P / Cooper LP / Roberts GA / Bujnicki JM / Trinick J / Kneale GG / Dryden DTF | |||||||||
 Citation Citation |  Journal: Genes Dev / Year: 2012 Journal: Genes Dev / Year: 2012Title: Structure and operation of the DNA-translocating type I DNA restriction enzymes. Authors: Christopher K Kennaway / James E N Taylor / Chun Feng Song / Wojciech Potrzebowski / William Nicholson / John H White / Anna Swiderska / Agnieszka Obarska-Kosinska / Philip Callow / Laurie P ...Authors: Christopher K Kennaway / James E N Taylor / Chun Feng Song / Wojciech Potrzebowski / William Nicholson / John H White / Anna Swiderska / Agnieszka Obarska-Kosinska / Philip Callow / Laurie P Cooper / Gareth A Roberts / Jean-Baptiste Artero / Janusz M Bujnicki / John Trinick / G Geoff Kneale / David T F Dryden /    Abstract: Type I DNA restriction/modification (RM) enzymes are molecular machines found in the majority of bacterial species. Their early discovery paved the way for the development of genetic engineering. ...Type I DNA restriction/modification (RM) enzymes are molecular machines found in the majority of bacterial species. Their early discovery paved the way for the development of genetic engineering. They control (restrict) the influx of foreign DNA via horizontal gene transfer into the bacterium while maintaining sequence-specific methylation (modification) of host DNA. The endonuclease reaction of these enzymes on unmethylated DNA is preceded by bidirectional translocation of thousands of base pairs of DNA toward the enzyme. We present the structures of two type I RM enzymes, EcoKI and EcoR124I, derived using electron microscopy (EM), small-angle scattering (neutron and X-ray), and detailed molecular modeling. DNA binding triggers a large contraction of the open form of the enzyme to a compact form. The path followed by DNA through the complexes is revealed by using a DNA mimic anti-restriction protein. The structures reveal an evolutionary link between type I RM enzymes and type II RM enzymes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1890.map.gz emd_1890.map.gz | 214.9 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1890-v30.xml emd-1890-v30.xml emd-1890.xml emd-1890.xml | 14.4 KB 14.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_1890.png emd_1890.png | 155.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1890 http://ftp.pdbj.org/pub/emdb/structures/EMD-1890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1890 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1890.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1890.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EcoR124I R2 M2 S1 complex with 30bp DNA fragment bound (closed state). DNA is not visible in the negative stain images or map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.96 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : EcoR124I R2 M2 S1 complex with 30 bp cognate dsDNA fragment
| Entire | Name: EcoR124I R2 M2 S1 complex with 30 bp cognate dsDNA fragment |
|---|---|
| Components |
|
-Supramolecule #1000: EcoR124I R2 M2 S1 complex with 30 bp cognate dsDNA fragment
| Supramolecule | Name: EcoR124I R2 M2 S1 complex with 30 bp cognate dsDNA fragment type: sample / ID: 1000 / Details: Stained with uranyl acetate / Oligomeric state: 1x HsdS, 2x HsdM, 2x HsdR, 1x dsDNA / Number unique components: 4 |
|---|---|
| Molecular weight | Experimental: 384 KDa / Theoretical: 415 KDa / Method: Small angle neutron scattering (SANS) |
-Macromolecule #1: EcoR124I HsdS specificity subunit
| Macromolecule | Name: EcoR124I HsdS specificity subunit / type: protein_or_peptide / ID: 1 / Name.synonym: HsdS / Number of copies: 1 / Oligomeric state: Monomer / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 50 KDa |
| Sequence | GO: protein binding InterPro: Type I restriction modification DNA specificity domain |
-Macromolecule #2: EcoR124I HsdM methyltransferase subunit
| Macromolecule | Name: EcoR124I HsdM methyltransferase subunit / type: protein_or_peptide / ID: 2 / Name.synonym: HsdM / Number of copies: 2 / Oligomeric state: Dimer / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 59 KDa |
| Sequence | GO: protein binding / InterPro: DNA methylase, adenine-specific |
-Macromolecule #3: EcoR124I HsdR endonuclease subunit
| Macromolecule | Name: EcoR124I HsdR endonuclease subunit / type: protein_or_peptide / ID: 3 / Name.synonym: HsdR / Number of copies: 2 / Oligomeric state: monomer / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 120 KDa |
| Sequence | GO: DNA restriction-modification system / InterPro: Restriction endonuclease, type I, HsdR |
-Macromolecule #4: Deoxyribonucleic acid
| Macromolecule | Name: Deoxyribonucleic acid / type: dna / ID: 4 / Name.synonym: DNA / Classification: DNA / Structure: DOUBLE HELIX / Synthetic?: Yes |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 15 KDa |
| Sequence | String: CCGTGCAGAA TTCGAGGTCG ACGGATCCGG |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.050 mg/mL |
|---|---|
| Buffer | pH: 4.7 / Details: 20mM Tris-Cl, 100 mM NaCl, |
| Staining | Type: NEGATIVE Details: Protein was adsorbed onto UV treated carbon for 1 minute, blotted, then 1% uranyl acetate solution was applied for 1 min then blotted, three times. |
| Grid | Details: 400 mesh copper |
| Vitrification | Cryogen name: NONE / Instrument: OTHER / Details: Negative stain |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 1200EXII |
|---|---|
| Temperature | Average: 294 K |
| Alignment procedure | Legacy - Astigmatism: Corrected at 80,000x |
| Details | Customised JEOL 1200 EX microscope, low dose mode. |
| Date | Sep 1, 2010 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: OTHER / Digitization - Sampling interval: 15 µm / Number real images: 30 / Average electron dose: 40 e/Å2 / Details: Scanned on Imacon scanner / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 80 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 0.991 µm / Nominal defocus min: 0.405 µm / Nominal magnification: 40000 |
| Sample stage | Specimen holder: Side entry / Specimen holder model: JEOL |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | PDBEntryID_givenInChain. Protocol: rigid body. HsdR (2W00) was fitted into the density after the core methylase (HsdS and 2x HsdM) was fitted. |
| Refinement | Space: RECIPROCAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















