[English] 日本語
 Yorodumi
Yorodumi- EMDB-18767: K48-linked ubiquitin chain formation with a cullin-RING E3 ligase... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB~acceptor UB-BRD4 | |||||||||
 Map data Map data | DeepEMhancer sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CUL2 / VHL / ELOBC / BRD4 / MZ1 / PROTAC / Ubiquitin / Ubiquitin Ligase / Ubiquitin chain formation / Poliubiquitylation / LIGASE / NEDD8 | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
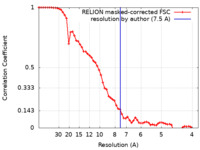
| Method | single particle reconstruction / cryo EM / Resolution: 7.5 Å | |||||||||
 Authors Authors | Liwocha J / Prabu JR / Kleiger G / Schulman BA | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Mechanism of millisecond Lys48-linked poly-ubiquitin chain formation by cullin-RING ligases. Authors: Joanna Liwocha / Jerry Li / Nicholas Purser / Chutima Rattanasopa / Samuel Maiwald / David T Krist / Daniel C Scott / Barbara Steigenberger / J Rajan Prabu / Brenda A Schulman / Gary Kleiger /   Abstract: E3 ubiquitin ligases, in collaboration with E2 ubiquitin-conjugating enzymes, modify proteins with poly-ubiquitin chains. Cullin-RING ligase (CRL) E3s use Cdc34/UBE2R-family E2s to build Lys48-linked ...E3 ubiquitin ligases, in collaboration with E2 ubiquitin-conjugating enzymes, modify proteins with poly-ubiquitin chains. Cullin-RING ligase (CRL) E3s use Cdc34/UBE2R-family E2s to build Lys48-linked poly-ubiquitin chains to control an enormous swath of eukaryotic biology. Yet the molecular mechanisms underlying this exceptional linkage specificity and millisecond kinetics of poly-ubiquitylation remain unclear. Here we obtain cryogenic-electron microscopy (cryo-EM) structures that provide pertinent insight into how such poly-ubiquitin chains are forged. The CRL RING domain not only activates the E2-bound ubiquitin but also shapes the conformation of a distinctive UBE2R2 loop, positioning both the ubiquitin to be transferred and the substrate-linked acceptor ubiquitin within the active site. The structures also reveal how the ubiquitin-like protein NEDD8 uniquely activates CRLs during chain formation. NEDD8 releases the RING domain from the CRL, but unlike previous CRL-E2 structures, does not contact UBE2R2. These findings suggest how poly-ubiquitylation may be accomplished by many E2s and E3s. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18767.map.gz emd_18767.map.gz | 16.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18767-v30.xml emd-18767-v30.xml emd-18767.xml emd-18767.xml | 19.6 KB 19.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18767_fsc.xml emd_18767_fsc.xml | 6.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_18767.png emd_18767.png | 58.3 KB | ||
| Masks |  emd_18767_msk_1.map emd_18767_msk_1.map | 18.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18767.cif.gz emd-18767.cif.gz | 4.4 KB | ||
| Others |  emd_18767_additional_1.map.gz emd_18767_additional_1.map.gz emd_18767_half_map_1.map.gz emd_18767_half_map_1.map.gz emd_18767_half_map_2.map.gz emd_18767_half_map_2.map.gz | 1.7 MB 13.8 MB 13.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18767 http://ftp.pdbj.org/pub/emdb/structures/EMD-18767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18767 | HTTPS FTP |
-Validation report
| Summary document |  emd_18767_validation.pdf.gz emd_18767_validation.pdf.gz | 671.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18767_full_validation.pdf.gz emd_18767_full_validation.pdf.gz | 671.4 KB | Display | |
| Data in XML |  emd_18767_validation.xml.gz emd_18767_validation.xml.gz | 11.2 KB | Display | |
| Data in CIF |  emd_18767_validation.cif.gz emd_18767_validation.cif.gz | 15.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18767 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18767 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18767 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18767 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18767.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18767.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DeepEMhancer sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.925 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18767_msk_1.map emd_18767_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Postprocessed map
| File | emd_18767_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_18767_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_18767_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : K48-linked ubiquitin chain formation with a cullin-RING E3 ligase...
| Entire | Name: K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB~acceptor UB-BRD4 |
|---|---|
| Components |
|
-Supramolecule #1: K48-linked ubiquitin chain formation with a cullin-RING E3 ligase...
| Supramolecule | Name: K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB~acceptor UB-BRD4 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Molecular weight | Theoretical: 190 KDa |
-Supramolecule #2: RING E3 ligase (RBX1) and Cullin-2 (CUL2)
| Supramolecule | Name: RING E3 ligase (RBX1) and Cullin-2 (CUL2) / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: NEDD8, ELOB, ELOC, VHL-MZ1, UBE2R2-donor UB-acceptor UB-BRD4 and ...
| Supramolecule | Name: NEDD8, ELOB, ELOC, VHL-MZ1, UBE2R2-donor UB-acceptor UB-BRD4 and K48-linked ubiquitin chain type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1-#2, #5-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 66.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)