+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1849 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cognate 70S-TC complex | ||||||||||||
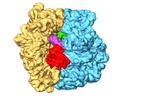
 Map data Map data | Surface rendered view of Cognate 70S-TC complex | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | ribosome / ternary complex / cognate | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationguanyl-nucleotide exchange factor complex / protein-synthesizing GTPase / guanosine tetraphosphate binding / stringent response / negative regulation of cytoplasmic translational initiation / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / translational elongation / misfolded RNA binding / Group I intron splicing ...guanyl-nucleotide exchange factor complex / protein-synthesizing GTPase / guanosine tetraphosphate binding / stringent response / negative regulation of cytoplasmic translational initiation / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / translational elongation / misfolded RNA binding / Group I intron splicing / RNA folding / translation elongation factor activity / transcriptional attenuation / translational termination / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / four-way junction DNA binding / negative regulation of cytoplasmic translation / DnaA-L2 complex / regulation of mRNA stability / translation repressor activity / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / mRNA regulatory element binding translation repressor activity / positive regulation of RNA splicing / regulation of DNA-templated transcription elongation / transcription elongation factor complex / response to reactive oxygen species / cytosolic ribosome assembly / ribosome assembly / assembly of large subunit precursor of preribosome / transcription antitermination / DNA endonuclease activity / translational initiation / regulation of cell growth / DNA-templated transcription termination / response to radiation / maintenance of translational fidelity / mRNA 5'-UTR binding / regulation of translation / large ribosomal subunit / ribosomal small subunit assembly / transferase activity / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / hydrolase activity / mRNA binding / GTPase activity / GTP binding / magnesium ion binding / DNA binding / RNA binding / zinc ion binding / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.25 Å | ||||||||||||
 Authors Authors | Agirrezabala X / Schreiner E / Trabuco LG / Lei J / Ortiz-Meoz RF / Schulten K / Green R / Frank J | ||||||||||||
 Citation Citation |  Journal: EMBO J / Year: 2011 Journal: EMBO J / Year: 2011Title: Structural insights into cognate versus near-cognate discrimination during decoding. Authors: Xabier Agirrezabala / Eduard Schreiner / Leonardo G Trabuco / Jianlin Lei / Rodrigo F Ortiz-Meoz / Klaus Schulten / Rachel Green / Joachim Frank /  Abstract: The structural basis of the tRNA selection process is investigated by cryo-electron microscopy of ribosomes programmed with UGA codons and incubated with ternary complex (TC) containing the near- ...The structural basis of the tRNA selection process is investigated by cryo-electron microscopy of ribosomes programmed with UGA codons and incubated with ternary complex (TC) containing the near-cognate Trp-tRNA(Trp) in the presence of kirromycin. Going through more than 350 000 images and employing image classification procedures, we find ∼8% in which the TC is bound to the ribosome. The reconstructed 3D map provides a means to characterize the arrangement of the near-cognate aa-tRNA with respect to elongation factor Tu (EF-Tu) and the ribosome, as well as the domain movements of the ribosome. One of the interesting findings is that near-cognate tRNA's acceptor stem region is flexible and CCA end becomes disordered. The data bring direct structural insights into the induced-fit mechanism of decoding by the ribosome, as the analysis of the interactions between small and large ribosomal subunit, aa-tRNA and EF-Tu and comparison with the cognate case (UGG codon) offers clues on how the conformational signals conveyed to the GTPase differ in the two cases. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1849.map.gz emd_1849.map.gz | 54.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1849-v30.xml emd-1849-v30.xml emd-1849.xml emd-1849.xml | 11.8 KB 11.8 KB | Display Display |  EMDB header EMDB header |
| Images |  1849.tif 1849.tif | 1.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1849 http://ftp.pdbj.org/pub/emdb/structures/EMD-1849 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1849 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1849 | HTTPS FTP |
-Related structure data
| Related structure data |  4v6kMC  1850C  4v6lC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1849.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1849.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Surface rendered view of Cognate 70S-TC complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Cognate 70S-TC complex
| Entire | Name: Cognate 70S-TC complex |
|---|---|
| Components |
|
-Supramolecule #1000: Cognate 70S-TC complex
| Supramolecule | Name: Cognate 70S-TC complex / type: sample / ID: 1000 Details: 70S.fMet-tRNAfMet.EF-Tu.Trp-tRNATrp on cognate codon Oligomeric state: Monomeric / Number unique components: 4 |
|---|---|
| Molecular weight | Theoretical: 2.8 MDa / Method: Sedimentation |
-Supramolecule #1: 70S Ribosome
| Supramolecule | Name: 70S Ribosome / type: complex / ID: 1 / Recombinant expression: No / Ribosome-details: ribosome-prokaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: fMet-tRNAfMet
| Macromolecule | Name: fMet-tRNAfMet / type: rna / ID: 1 / Name.synonym: fMet-tRNAfMet / Classification: TRANSFER / Structure: SINGLE STRANDED / Synthetic?: No |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: CGCGGGGTGG AGCAGCCTGG TAGCTCGTCG GGCTCATAAC CCGAAGATCG TCGGTTCAAA TCCGGCCCCC GCAACCA |
-Macromolecule #3: Trp-tRNATrp
| Macromolecule | Name: Trp-tRNATrp / type: rna / ID: 3 / Classification: TRANSFER / Structure: SINGLE STRANDED / Synthetic?: No |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: AGGGGCGTAG TTCAATTGGT AGAGCACCGG TCTCCAAAAC CGGGTGTTGG GAGTTCGAGT CTCTCCGCCC CTGCCA |
-Macromolecule #2: EF-Tu
| Macromolecule | Name: EF-Tu / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.04 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: HiFi (50 mM Tris-HCl pH 7.5, 70mM NH4Cl, 30 mM KCl, 3.5 mM MgCl2, 0.5 mM spermidine, 8mM putrescine, 2 mM DTT) |
| Vitrification | Cryogen name: NITROGEN / Chamber humidity: 90 % / Chamber temperature: 80 K / Instrument: OTHER / Details: Vitrification instrument: Vitrobot / Method: Blot for 6 seconds |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective corrected at 100,000 times magnification |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F415 (4k x 4k) / Average electron dose: 20 e/Å2 Details: Automated data collection system AutoEMation (CCD mag. 100000x) TVIPS TemCam-F415 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 58269 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: Defocus groups, Wiener filter |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 8.25 Å / Resolution method: OTHER / Software - Name: Spider / Number images used: 359223 |
-Atomic model buiding 1
| Initial model | PDB ID:  2i2v |
|---|---|
| Software | Name: MDFF |
| Details | Protocol: MD-based flexible fitting |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: RMSD, cross correlation |
| Output model |  PDB-4v6k: |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















