[English] 日本語
 Yorodumi
Yorodumi- EMDB-17713: Catalytic module of human CTLH E3 ligase bound to multiphosphoryl... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Catalytic module of human CTLH E3 ligase bound to multiphosphorylated UBE2H~ubiquitin | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | E3 ubiquitin ligase / E2 ubiquitin-conjugating enzyme / phosphorylation / CTLH / GID / UBE2H / LIGASE | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationGID complex / actomyosin contractile ring / enucleate erythrocyte development / negative regulation of myeloid cell apoptotic process / (E3-independent) E2 ubiquitin-conjugating enzyme / protein K11-linked ubiquitination / E2 ubiquitin-conjugating enzyme / erythrocyte maturation / ubiquitin conjugating enzyme activity / ubiquitin ligase complex ...GID complex / actomyosin contractile ring / enucleate erythrocyte development / negative regulation of myeloid cell apoptotic process / (E3-independent) E2 ubiquitin-conjugating enzyme / protein K11-linked ubiquitination / E2 ubiquitin-conjugating enzyme / erythrocyte maturation / ubiquitin conjugating enzyme activity / ubiquitin ligase complex / protein K48-linked ubiquitination / Maturation of protein E / cytoskeleton organization / Maturation of protein E / ER Quality Control Compartment (ERQC) / Myoclonic epilepsy of Lafora / FLT3 signaling by CBL mutants / IRAK2 mediated activation of TAK1 complex / Prevention of phagosomal-lysosomal fusion / Alpha-protein kinase 1 signaling pathway / Glycogen synthesis / IRAK1 recruits IKK complex / IRAK1 recruits IKK complex upon TLR7/8 or 9 stimulation / Endosomal Sorting Complex Required For Transport (ESCRT) / Membrane binding and targetting of GAG proteins / Negative regulation of FLT3 / Regulation of TBK1, IKKε (IKBKE)-mediated activation of IRF3, IRF7 / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / Regulation of TBK1, IKKε-mediated activation of IRF3, IRF7 upon TLR3 ligation / IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation / Constitutive Signaling by NOTCH1 HD Domain Mutants / NOTCH2 Activation and Transmission of Signal to the Nucleus / TICAM1,TRAF6-dependent induction of TAK1 complex / TICAM1-dependent activation of IRF3/IRF7 / regulation of mitotic cell cycle / APC/C:Cdc20 mediated degradation of Cyclin B / Regulation of FZD by ubiquitination / Downregulation of ERBB4 signaling / APC-Cdc20 mediated degradation of Nek2A / p75NTR recruits signalling complexes / InlA-mediated entry of Listeria monocytogenes into host cells / TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling / NF-kB is activated and signals survival / TRAF6-mediated induction of TAK1 complex within TLR4 complex / Regulation of pyruvate metabolism / Pexophagy / Downregulation of ERBB2:ERBB3 signaling / Regulation of innate immune responses to cytosolic DNA / NRIF signals cell death from the nucleus / Regulation of PTEN localization / VLDLR internalisation and degradation / Activated NOTCH1 Transmits Signal to the Nucleus / Synthesis of active ubiquitin: roles of E1 and E2 enzymes / Translesion synthesis by REV1 / TICAM1, RIP1-mediated IKK complex recruitment / Regulation of BACH1 activity / Translesion synthesis by POLK / InlB-mediated entry of Listeria monocytogenes into host cell / JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 / Activation of IRF3, IRF7 mediated by TBK1, IKKε (IKBKE) / MAP3K8 (TPL2)-dependent MAPK1/3 activation / Downregulation of TGF-beta receptor signaling / Translesion synthesis by POLI / Josephin domain DUBs / IKK complex recruitment mediated by RIP1 / Gap-filling DNA repair synthesis and ligation in GG-NER / PINK1-PRKN Mediated Mitophagy / TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) / Regulation of activated PAK-2p34 by proteasome mediated degradation / TNFR1-induced NF-kappa-B signaling pathway / TCF dependent signaling in response to WNT / Regulation of NF-kappa B signaling / activated TAK1 mediates p38 MAPK activation / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / Asymmetric localization of PCP proteins / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / NOTCH3 Activation and Transmission of Signal to the Nucleus / Negative regulators of DDX58/IFIH1 signaling / Ubiquitin-dependent degradation of Cyclin D / Regulation of signaling by CBL / Fanconi Anemia Pathway / Negative regulation of FGFR3 signaling / Peroxisomal protein import / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / AUF1 (hnRNP D0) binds and destabilizes mRNA / Deactivation of the beta-catenin transactivating complex / TNFR2 non-canonical NF-kB pathway / Stabilization of p53 / Negative regulation of FGFR2 signaling / Assembly of the pre-replicative complex / Negative regulation of FGFR4 signaling / Vpu mediated degradation of CD4 / Downregulation of SMAD2/3:SMAD4 transcriptional activity / Negative regulation of FGFR1 signaling / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / Termination of translesion DNA synthesis / EGFR downregulation / Regulation of TNFR1 signaling Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
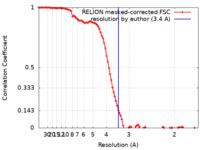
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||
 Authors Authors | Chrustowicz J / Sherpa D / Prabu RJ / Schulman BA | ||||||||||||
| Funding support |  Germany, European Union, 3 items Germany, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2024 Journal: Mol Cell / Year: 2024Title: Multisite phosphorylation dictates selective E2-E3 pairing as revealed by Ubc8/UBE2H-GID/CTLH assemblies. Authors: Jakub Chrustowicz / Dawafuti Sherpa / Jerry Li / Christine R Langlois / Eleftheria C Papadopoulou / D Tung Vu / Laura A Hehl / Özge Karayel / Viola Beier / Susanne von Gronau / Judith ...Authors: Jakub Chrustowicz / Dawafuti Sherpa / Jerry Li / Christine R Langlois / Eleftheria C Papadopoulou / D Tung Vu / Laura A Hehl / Özge Karayel / Viola Beier / Susanne von Gronau / Judith Müller / J Rajan Prabu / Matthias Mann / Gary Kleiger / Arno F Alpi / Brenda A Schulman /   Abstract: Ubiquitylation is catalyzed by coordinated actions of E3 and E2 enzymes. Molecular principles governing many important E3-E2 partnerships remain unknown, including those for RING-family GID/CTLH E3 ...Ubiquitylation is catalyzed by coordinated actions of E3 and E2 enzymes. Molecular principles governing many important E3-E2 partnerships remain unknown, including those for RING-family GID/CTLH E3 ubiquitin ligases and their dedicated E2, Ubc8/UBE2H (yeast/human nomenclature). GID/CTLH-Ubc8/UBE2H-mediated ubiquitylation regulates biological processes ranging from yeast metabolic signaling to human development. Here, cryoelectron microscopy (cryo-EM), biochemistry, and cell biology reveal this exquisitely specific E3-E2 pairing through an unconventional catalytic assembly and auxiliary interactions 70-100 Å away, mediated by E2 multisite phosphorylation. Rather than dynamic polyelectrostatic interactions reported for other ubiquitylation complexes, multiple Ubc8/UBE2H phosphorylation sites within acidic CK2-targeted sequences specifically anchor the E2 C termini to E3 basic patches. Positions of phospho-dependent interactions relative to the catalytic domains correlate across evolution. Overall, our data show that phosphorylation-dependent multivalency establishes a specific E3-E2 partnership, is antagonistic with dephosphorylation, rigidifies the catalytic centers within a flexing GID E3-substrate assembly, and facilitates substrate collision with ubiquitylation active sites. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17713.map.gz emd_17713.map.gz | 6.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17713-v30.xml emd-17713-v30.xml emd-17713.xml emd-17713.xml | 27.4 KB 27.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17713_fsc.xml emd_17713_fsc.xml | 9.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_17713.png emd_17713.png | 106.6 KB | ||
| Masks |  emd_17713_msk_1.map emd_17713_msk_1.map | 76.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17713.cif.gz emd-17713.cif.gz | 7.8 KB | ||
| Others |  emd_17713_additional_1.map.gz emd_17713_additional_1.map.gz emd_17713_half_map_1.map.gz emd_17713_half_map_1.map.gz emd_17713_half_map_2.map.gz emd_17713_half_map_2.map.gz | 66.3 MB 59.9 MB 59.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17713 http://ftp.pdbj.org/pub/emdb/structures/EMD-17713 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17713 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17713 | HTTPS FTP |
-Related structure data
| Related structure data |  8pjnMC  8pmqC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17713.map.gz / Format: CCP4 / Size: 76.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17713.map.gz / Format: CCP4 / Size: 76.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.851 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17713_msk_1.map emd_17713_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_17713_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17713_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17713_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of human CTLHSR4 E3 ligase bound to engineered ARMC8-spec...
| Entire | Name: Complex of human CTLHSR4 E3 ligase bound to engineered ARMC8-specific VH and multiphosphorylated UBE2H~ubiquitin |
|---|---|
| Components |
|
-Supramolecule #1: Complex of human CTLHSR4 E3 ligase bound to engineered ARMC8-spec...
| Supramolecule | Name: Complex of human CTLHSR4 E3 ligase bound to engineered ARMC8-specific VH and multiphosphorylated UBE2H~ubiquitin type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: Map obtained by focus refinement over the catalytic module (RMND5A, MAEA) and UBE2H~ubiquitin |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 600 KDa |
-Macromolecule #1: E3 ubiquitin-protein transferase RMND5A
| Macromolecule | Name: E3 ubiquitin-protein transferase RMND5A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: RING-type E3 ubiquitin transferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.043449 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MDQCVTVERE LEKVLHKFSG YGQLCERGLE ELIDYTGGLK HEILQSHGQD AELSGTLSLV LTQCCKRIKD TVQKLASDHK DIHSSVSRV GKAIDKNFDS DISSVGIDGC WQADSQRLLN EVMVEHFFRQ GMLDVAEELC QESGLSVDPS QKEPFVELNR I LEALKVRV ...String: MDQCVTVERE LEKVLHKFSG YGQLCERGLE ELIDYTGGLK HEILQSHGQD AELSGTLSLV LTQCCKRIKD TVQKLASDHK DIHSSVSRV GKAIDKNFDS DISSVGIDGC WQADSQRLLN EVMVEHFFRQ GMLDVAEELC QESGLSVDPS QKEPFVELNR I LEALKVRV LRPALEWAVS NREMLIAQNS SLEFKLHRLY FISLLMGGTT NQREALQYAK NFQPFALNHQ KDIQVLMGSL VY LRQGIEN SPYVHLLDAN QWADICDIFT RDACALLGLS VESPLSVSFS AGCVALPALI NIKAVIEQRQ CTGVWNQKDE LPI EVDLGK KCWYHSIFAC PILRQQTTDN NPPMKLVCGH IISRDALNKM FNGSKLKCPY CPMEQSPGDA KQIFF UniProtKB: E3 ubiquitin-protein transferase RMND5A |
-Macromolecule #2: Ubiquitin-conjugating enzyme E2 H
| Macromolecule | Name: Ubiquitin-conjugating enzyme E2 H / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: E2 ubiquitin-conjugating enzyme |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 21.260986 KDa |
| Recombinant expression | Organism:  Trichoplusia (butterflies/moths) Trichoplusia (butterflies/moths) |
| Sequence | String: MSSPSPGKRR MDTDVVKLIE SKHEVTILGG LNEFVVKFYG PQGTPYEGGV WKVRVDLPDK YPFKSPSIGF MNKIFHPNID EASGTVKLD VINQTWTALY DLTNIFESFL PQLLAYPNPI DPLNGDAAAM YLHRPEEYKQ KIKEYIQKYA TEEALKEQEE G TGD(SEP)(SEP) ...String: MSSPSPGKRR MDTDVVKLIE SKHEVTILGG LNEFVVKFYG PQGTPYEGGV WKVRVDLPDK YPFKSPSIGF MNKIFHPNID EASGTVKLD VINQTWTALY DLTNIFESFL PQLLAYPNPI DPLNGDAAAM YLHRPEEYKQ KIKEYIQKYA TEEALKEQEE G TGD(SEP)(SEP)(SEP)E(SEP) (SEP)M(SEP)DF(SEP)EDEA QDMEL UniProtKB: Ubiquitin-conjugating enzyme E2 H |
-Macromolecule #3: E3 ubiquitin-protein transferase MAEA
| Macromolecule | Name: E3 ubiquitin-protein transferase MAEA / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: RING-type E3 ubiquitin transferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.356332 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MAVQESAAQL SMTLKVQEYP TLKVPYETLN KRFRAAQKNI DRETSHVTMV VAELEKTLSG CPAVDSVVSL LDGVVEKLSV LKRKAVESI QAEDESAKLC KRRIEHLKEH SSDQPAAASV WKRKRMDRMM VEHLLRCGYY NTAVKLARQS GIEDLVNIEM F LTAKEVEE ...String: MAVQESAAQL SMTLKVQEYP TLKVPYETLN KRFRAAQKNI DRETSHVTMV VAELEKTLSG CPAVDSVVSL LDGVVEKLSV LKRKAVESI QAEDESAKLC KRRIEHLKEH SSDQPAAASV WKRKRMDRMM VEHLLRCGYY NTAVKLARQS GIEDLVNIEM F LTAKEVEE SLERRETATC LAWCHDNKSR LRKMKSCLEF SLRIQEFIEL IRQNKRLDAV RHARKHFSQA EGSQLDEVRQ AM GMLAFPP DTHISPYKDL LDPARWRMLI QQFRYDNYRL HQLGNNSVFT LTLQAGLSAI KTPQCYKEDG SSKSPDCPVC SRS LNKLAQ PLPMAHCANS RLVCKISGDV MNENNPPMML PNGYVYGYNS LLSIRQDDKV VCPRTKEVFH FSQAEKVYIM UniProtKB: E3 ubiquitin-protein transferase MAEA |
-Macromolecule #4: Ubiquitin
| Macromolecule | Name: Ubiquitin / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.576831 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQIFVKTLTG KTITLEVEPS DTIENVKAKI QDKEGIPPDQ QRLIFAGKQL EDGRTLSDYN IQKESTLHLV LRLRGG UniProtKB: Polyubiquitin-C |
-Macromolecule #5: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 68.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)