+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ab typeII filament from Guam ALS/PDC | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ab typeII filaments / Guam ALS/PDC / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationamyloid-beta complex / growth cone lamellipodium / cellular response to norepinephrine stimulus / collateral sprouting in absence of injury / growth cone filopodium / microglia development / Formyl peptide receptors bind formyl peptides and many other ligands / axo-dendritic transport / regulation of Wnt signaling pathway / axon midline choice point recognition ...amyloid-beta complex / growth cone lamellipodium / cellular response to norepinephrine stimulus / collateral sprouting in absence of injury / growth cone filopodium / microglia development / Formyl peptide receptors bind formyl peptides and many other ligands / axo-dendritic transport / regulation of Wnt signaling pathway / axon midline choice point recognition / regulation of synapse structure or activity / hippocampal neuron apoptotic process / astrocyte activation involved in immune response / NMDA selective glutamate receptor signaling pathway / regulation of spontaneous synaptic transmission / mating behavior / growth factor receptor binding / peptidase activator activity / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / PTB domain binding / positive regulation of amyloid fibril formation / Golgi-associated vesicle / astrocyte projection / Lysosome Vesicle Biogenesis / neuron remodeling / Deregulated CDK5 triggers multiple neurodegenerative pathways in Alzheimer's disease models / nuclear envelope lumen / dendrite development / TRAF6 mediated NF-kB activation / positive regulation of protein metabolic process / signaling receptor activator activity / negative regulation of long-term synaptic potentiation / Advanced glycosylation endproduct receptor signaling / transition metal ion binding / The NLRP3 inflammasome / main axon / intracellular copper ion homeostasis / modulation of excitatory postsynaptic potential / regulation of multicellular organism growth / ECM proteoglycans / response to insulin-like growth factor stimulus / regulation of presynapse assembly / positive regulation of T cell migration / neuronal dense core vesicle / Purinergic signaling in leishmaniasis infection / cellular response to manganese ion / positive regulation of chemokine production / Notch signaling pathway / swimming behavior / neuron projection maintenance / extracellular matrix organization / clathrin-coated pit / positive regulation of mitotic cell cycle / axonogenesis / Mitochondrial protein degradation / positive regulation of calcium-mediated signaling / ionotropic glutamate receptor signaling pathway / astrocyte activation / platelet alpha granule lumen / response to interleukin-1 / regulation of neuron apoptotic process / cellular response to copper ion / cellular response to cAMP / positive regulation of glycolytic process / endosome lumen / trans-Golgi network membrane / dendritic shaft / positive regulation of interleukin-1 beta production / protein serine/threonine kinase binding / positive regulation of long-term synaptic potentiation / learning / central nervous system development / adult locomotory behavior / Post-translational protein phosphorylation / serine-type endopeptidase inhibitor activity / locomotory behavior / microglial cell activation / cellular response to nerve growth factor stimulus / TAK1-dependent IKK and NF-kappa-B activation / positive regulation of non-canonical NF-kappaB signal transduction / synapse organization / visual learning / recycling endosome / regulation of long-term neuronal synaptic plasticity / Golgi lumen / positive regulation of interleukin-6 production / positive regulation of JNK cascade / response to lead ion / cognition / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / cellular response to amyloid-beta / endocytosis / neuron projection development / positive regulation of tumor necrosis factor production / positive regulation of inflammatory response / calcium ion transport / Platelet degranulation / regulation of translation / heparin binding / regulation of gene expression Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
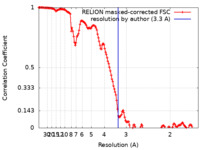
| Method | helical reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Qi C / Yang S / Scheres SHW / Goedert M | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Tau filaments from amyotrophic lateral sclerosis/parkinsonism-dementia complex adopt the CTE fold. Authors: Chao Qi / Bert M Verheijen / Yasumasa Kokubo / Yang Shi / Stephan Tetter / Alexey G Murzin / Asa Nakahara / Satoru Morimoto / Marc Vermulst / Ryogen Sasaki / Eleonora Aronica / Yoshifumi ...Authors: Chao Qi / Bert M Verheijen / Yasumasa Kokubo / Yang Shi / Stephan Tetter / Alexey G Murzin / Asa Nakahara / Satoru Morimoto / Marc Vermulst / Ryogen Sasaki / Eleonora Aronica / Yoshifumi Hirokawa / Kiyomitsu Oyanagi / Akiyoshi Kakita / Benjamin Ryskeldi-Falcon / Mari Yoshida / Masato Hasegawa / Sjors H W Scheres / Michel Goedert /     Abstract: The amyotrophic lateral sclerosis/parkinsonism-dementia complex (ALS/PDC) of the island of Guam and the Kii peninsula of Japan is a fatal neurodegenerative disease of unknown cause that is ...The amyotrophic lateral sclerosis/parkinsonism-dementia complex (ALS/PDC) of the island of Guam and the Kii peninsula of Japan is a fatal neurodegenerative disease of unknown cause that is characterized by the presence of abundant filamentous tau inclusions in brains and spinal cords. Here, we used electron cryo-microscopy to determine the structures of tau filaments from the cerebral cortex of three cases of ALS/PDC from Guam and eight cases from Kii, as well as from the spinal cord of two of the Guam cases. Tau filaments had the chronic traumatic encephalopathy (CTE) fold, with variable amounts of Type I and Type II filaments. Paired helical tau filaments were also found in three Kii cases and tau filaments with the corticobasal degeneration fold in one Kii case. We identified a new Type III CTE tau filament, where protofilaments pack against each other in an antiparallel fashion. ALS/PDC is the third known tauopathy with CTE-type filaments and abundant tau inclusions in cortical layers II/III, the others being CTE and subacute sclerosing panencephalitis. Because these tauopathies are believed to have environmental causes, our findings support the hypothesis that ALS/PDC is caused by exogenous factors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17177.map.gz emd_17177.map.gz | 58.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17177-v30.xml emd-17177-v30.xml emd-17177.xml emd-17177.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17177_fsc.xml emd_17177_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_17177.png emd_17177.png | 45.6 KB | ||
| Masks |  emd_17177_msk_1.map emd_17177_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17177.cif.gz emd-17177.cif.gz | 5.6 KB | ||
| Others |  emd_17177_half_map_1.map.gz emd_17177_half_map_1.map.gz emd_17177_half_map_2.map.gz emd_17177_half_map_2.map.gz | 192.5 MB 192.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17177 http://ftp.pdbj.org/pub/emdb/structures/EMD-17177 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17177 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17177 | HTTPS FTP |
-Related structure data
| Related structure data |  8otfMC  8ot6C  8ot9C  8otcC  8otdC  8oteC  8otgC  8othC  8otiC  8otjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17177.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17177.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.824 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17177_msk_1.map emd_17177_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17177_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17177_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ab typeII filaments from Guam ALS/PDC
| Entire | Name: Ab typeII filaments from Guam ALS/PDC |
|---|---|
| Components |
|
-Supramolecule #1: Ab typeII filaments from Guam ALS/PDC
| Supramolecule | Name: Ab typeII filaments from Guam ALS/PDC / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Amyloid-beta precursor protein
| Macromolecule | Name: Amyloid-beta precursor protein / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 87.046219 KDa |
| Sequence | String: MLPGLALLLL AAWTARALEV PTDGNAGLLA EPQIAMFCGR LNMHMNVQNG KWDSDPSGTK TCIDTKEGIL QYCQEVYPEL QITNVVEAN QPVTIQNWCK RGRKQCKTHP HFVIPYRCLV GEFVSDALLV PDKCKFLHQE RMDVCETHLH WHTVAKETCS E KSTNLHDY ...String: MLPGLALLLL AAWTARALEV PTDGNAGLLA EPQIAMFCGR LNMHMNVQNG KWDSDPSGTK TCIDTKEGIL QYCQEVYPEL QITNVVEAN QPVTIQNWCK RGRKQCKTHP HFVIPYRCLV GEFVSDALLV PDKCKFLHQE RMDVCETHLH WHTVAKETCS E KSTNLHDY GMLLPCGIDK FRGVEFVCCP LAEESDNVDS ADAEEDDSDV WWGGADTDYA DGSEDKVVEV AEEEEVAEVE EE EADDDED DEDGDEVEEE AEEPYEEATE RTTSIATTTT TTTESVEEVV REVCSEQAET GPCRAMISRW YFDVTEGKCA PFF YGGCGG NRNNFDTEEY CMAVCGSAMS QSLLKTTQEP LARDPVKLPT TAASTPDAVD KYLETPGDEN EHAHFQKAKE RLEA KHRER MSQVMREWEE AERQAKNLPK ADKKAVIQHF QEKVESLEQE AANERQQLVE THMARVEAML NDRRRLALEN YITAL QAVP PRPRHVFNML KKYVRAEQKD RQHTLKHFEH VRMVDPKKAA QIRSQVMTHL RVIYERMNQS LSLLYNVPAV AEEIQD EVD ELLQKEQNYS DDVLANMISE PRISYGNDAL MPSLTETKTT VELLPVNGEF SLDDLQPWHS FGADSVPANT ENEVEPV DA RPAADRGLTT RPGSGLTNIK TEEISEVKMD AEFRHDSGYE VHHQKLVFFA EDVGSNKGAI IGLMVGGVVI ATVIVITL V MLKKKQYTSI HHGVVEVDAA VTPEERHLSK MQQNGYENPT YKFFEQMQN UniProtKB: Amyloid-beta precursor protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)