[English] 日本語
 Yorodumi
Yorodumi- EMDB-17041: Cryo-EM structure of P5A-ATPase CtSpf1 (E1P-ADP state with membra... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of P5A-ATPase CtSpf1 (E1P-ADP state with membranous-feature bound) | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | translocase / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationP-type ion transporter activity / ATPase-coupled monoatomic cation transmembrane transporter activity / intracellular calcium ion homeostasis / endoplasmic reticulum membrane / ATP hydrolysis activity / ATP binding Similarity search - Function | |||||||||||||||
| Biological species |  Thermochaetoides thermophila (fungus) Thermochaetoides thermophila (fungus) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.47 Å | |||||||||||||||
 Authors Authors | Li P / Gourdon P | |||||||||||||||
| Funding support |  Sweden, Sweden,  Denmark, 4 items Denmark, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: The structure and function of P5A-ATPases. Authors: Ping Li / Viktoria Bågenholm / Per Hägglund / Karin Lindkvist-Petersson / Kaituo Wang / Pontus Gourdon /    Abstract: Endoplasmic reticulum (ER) membrane resident P5A-ATPases broadly affect protein biogenesis and quality control, and yet their molecular function remains debated. Here, we report cryo-EM structures of ...Endoplasmic reticulum (ER) membrane resident P5A-ATPases broadly affect protein biogenesis and quality control, and yet their molecular function remains debated. Here, we report cryo-EM structures of a P5A-ATPase, CtSpf1, covering multiple transport intermediates of the E1 → E1-ATP → E1P-ADP → E1P → E2P → E2.P → E2 → E1 cycle. In the E2P and E2.P states a cleft spans the entire membrane, holding a polypeptide cargo molecule. The cargo includes an ER luminal extension, pinpointed as the C-terminus in the E2.P state, which reenters the membrane in E2P. The E1 structure harbors a cytosol-facing cavity that is blocked by an insertion we refer to as the Plug-domain. The Plug-domain is nestled to key ATPase features and is displaced in the E1P-ADP and E1P states. Collectively, our findings are compatible with a broad range of proteins as cargo, with the P5A-ATPases serving a role in membrane removal of helices, although insertion/secretion cannot be excluded, as well as with a mechanistic role of the Plug-domain. #1:  Journal: Acta Crystallogr., Sect. D: Biol. Crystallogr. / Year: 2018 Journal: Acta Crystallogr., Sect. D: Biol. Crystallogr. / Year: 2018Title: Real-space refinement in PHENIX for cryo-EM and crystallography Authors: Li P / Gourdon P | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17041.map.gz emd_17041.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17041-v30.xml emd-17041-v30.xml emd-17041.xml emd-17041.xml | 17.4 KB 17.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17041_fsc.xml emd_17041_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_17041.png emd_17041.png | 89.4 KB | ||
| Filedesc metadata |  emd-17041.cif.gz emd-17041.cif.gz | 6.3 KB | ||
| Others |  emd_17041_half_map_1.map.gz emd_17041_half_map_1.map.gz emd_17041_half_map_2.map.gz emd_17041_half_map_2.map.gz | 116 MB 116 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17041 http://ftp.pdbj.org/pub/emdb/structures/EMD-17041 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17041 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17041 | HTTPS FTP |
-Validation report
| Summary document |  emd_17041_validation.pdf.gz emd_17041_validation.pdf.gz | 953.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17041_full_validation.pdf.gz emd_17041_full_validation.pdf.gz | 953.1 KB | Display | |
| Data in XML |  emd_17041_validation.xml.gz emd_17041_validation.xml.gz | 19.2 KB | Display | |
| Data in CIF |  emd_17041_validation.cif.gz emd_17041_validation.cif.gz | 24.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17041 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17041 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17041 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17041 | HTTPS FTP |
-Related structure data
| Related structure data |  8op5MC  8op3C  8op4C  8op6C  8op7C  8op8C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17041.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17041.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.846 Å | ||||||||||||||||||||||||||||||||||||
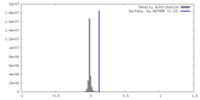
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_17041_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17041_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CtSpf1 with AlF4-ADP bound
| Entire | Name: CtSpf1 with AlF4-ADP bound |
|---|---|
| Components |
|
-Supramolecule #1: CtSpf1 with AlF4-ADP bound
| Supramolecule | Name: CtSpf1 with AlF4-ADP bound / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila (fungus) Thermochaetoides thermophila (fungus) |
-Macromolecule #1: Cation-transporting ATPase-like protein
| Macromolecule | Name: Cation-transporting ATPase-like protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila (fungus) Thermochaetoides thermophila (fungus) |
| Molecular weight | Theoretical: 149.103156 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAPLVDNPQI KSAELLRPLP LYQHAYVWPY VIVWPVFLRV YLTQELYDKY IGAQEWTFVW IISIVTFQTL TWLCTHWSVN LNALFTAKK ASSIEDAQLI KVIPVANAGA ADICKLVRDK VGDNKTNISF LFQKRRFLWY PERKAFSTLE FDIDAEPKPT L SKFQLSRG ...String: MAPLVDNPQI KSAELLRPLP LYQHAYVWPY VIVWPVFLRV YLTQELYDKY IGAQEWTFVW IISIVTFQTL TWLCTHWSVN LNALFTAKK ASSIEDAQLI KVIPVANAGA ADICKLVRDK VGDNKTNISF LFQKRRFLWY PERKAFSTLE FDIDAEPKPT L SKFQLSRG IESEDELKRL EQHYGTNTFD IPVPTFTELF KEHAVAPFFV FQVFCVGLWL LDEYWYYSLF TLVMLVVFES TV VWQRQRT LTEFRSMSIK PYPIYVYRLG KWTEIQSDKL LPGDLVSVTR TKEDSGVACD MILVEGTAIV NEAMLSGEST PLL KDSIQL RPGDAVLEVD GLDKNSLLWG GTKVLQITHG TAEEERPKPA SGIPPPPDNG AMAVVTKTGF ETSQGSLVRT MIYS TERVS ANNTEALLFI LFLLVFALAA SWYVWDEGVR KDRKRSKLLL DCILIITSVV PPELPMELSL AVNTSLSALA KFAIF CTEP FRIPFAGRID VACFDKTGTL TGEDLVVEGI AGLGLGHSGT DTPKEADGAH TRMVSVHDAG METTLVLATA HALVKL DEG EIVGDPMEKA TLNALGWVLG KNDTLTSKPG NAASSGILGT VQIKRRFQFS SALKRQSSVA TITATEVKTG RKLRGSF VG VKGAPETIMK MLVTVPEHYE ETYKYFTRRG SRVLALAYKQ LTTEGELGAN KINDLKRESV EADLHFAGFL VLQCPLKE D AKQAVRMLNE SSHRVVMITG DNPLTAVHVA KEVEIVDRDV LILDAPEHSV YGEESLVWRS VDDKIRIDVD PTKPIDPEI LKTKDLCVTG YALNKFKGQV GWKSLLRYTW VYARVSPKQK EDILLGLKDM GYYTLMAGDG TNDVGALKQA HVGVALLNGT QEDLNRIAE HTRNQKMKEL YQKQVDLMAR WGQPPPPVPA MIAHLYPPGP SNPHYQKAME REAQKRGVTV EQLAKVNGTN V TSNPAGVQ QQSGQDAKKA KQVEAAKKAA NFADKLTSSL MEAEMDDEPP TLKLGDASVA APFTSKLRNV MAIPNILRQG RC TLVATIQ MYKILALNCL ISAYSLSVLY LEGIKFGDGQ ITISGMLMSV CFLSISRARS VEGLSKERPQ PNIFNFYIIG SIL GQFAVH VATLIYIAQL CDQIEPRTEV IDLEAEFKPS LLNSAVYLLQ LIQQISTFAV NYQGRPFRES LSENKGMFYG IVGV TAIAF ACSTEMLPEL NEAMKLVPFN ENFKTIMTTV MIIDFVACYV IEWVLKKLFS DLRARDIAER RPDQLERERV RKEKE AREK EEEEERKERE RIEAFERRLE EKRTRLVEAA AQREQQQQQW AQRR UniProtKB: Cation-transporting ATPase-like protein |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 1 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #3: TETRAFLUOROALUMINATE ION
| Macromolecule | Name: TETRAFLUOROALUMINATE ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: ALF |
|---|---|
| Molecular weight | Theoretical: 102.975 Da |
| Chemical component information |  ChemComp-ALF: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 2 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.7000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)