[English] 日本語
 Yorodumi
Yorodumi- EMDB-16887: Cryo-EM structure of the undecorated barbed end of filamentous be... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the undecorated barbed end of filamentous beta/gamma actin | ||||||||||||
 Map data Map data | 3D-refined, sharpened cryo-EM density map of the undecorated barbed end of F-actin. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Actin filament / cytoskeletal protein / ATPase / STRUCTURAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationCell-extracellular matrix interactions / Formation of the canonical BAF (cBAF) complex / Formation of the polybromo-BAF (pBAF) complex / Formation of the embryonic stem cell BAF (esBAF) complex / Formation of the non-canonical BAF (ncBAF) complex / Formation of neuronal progenitor and neuronal BAF (npBAF and nBAF) / Adherens junctions interactions / Regulation of CDH1 Function / Formation of the dystrophin-glycoprotein complex (DGC) / B-WICH complex positively regulates rRNA expression ...Cell-extracellular matrix interactions / Formation of the canonical BAF (cBAF) complex / Formation of the polybromo-BAF (pBAF) complex / Formation of the embryonic stem cell BAF (esBAF) complex / Formation of the non-canonical BAF (ncBAF) complex / Formation of neuronal progenitor and neuronal BAF (npBAF and nBAF) / Adherens junctions interactions / Regulation of CDH1 Function / Formation of the dystrophin-glycoprotein complex (DGC) / B-WICH complex positively regulates rRNA expression / Gap junction degradation / Formation of annular gap junctions / RHOF GTPase cycle / EPHB-mediated forward signaling / MAP2K and MAPK activation / Regulation of actin dynamics for phagocytic cup formation / RHO GTPases Activate WASPs and WAVEs / RHO GTPases activate IQGAPs / RHO GTPases Activate Formins / DNA Damage Recognition in GG-NER / UCH proteinases / VEGFA-VEGFR2 Pathway / cellular response to cytochalasin B / Clathrin-mediated endocytosis / regulation of transepithelial transport / morphogenesis of a polarized epithelium / structural constituent of postsynaptic actin cytoskeleton / protein localization to adherens junction / dense body / Tat protein binding / postsynaptic actin cytoskeleton / adherens junction assembly / apical protein localization / tight junction / apical junction complex / regulation of norepinephrine uptake / transporter regulator activity / NuA4 histone acetyltransferase complex / establishment or maintenance of cell polarity / cortical cytoskeleton / nitric-oxide synthase binding / brush border / regulation of synaptic vesicle endocytosis / kinesin binding / regulation of protein localization to plasma membrane / positive regulation of double-strand break repair via homologous recombination / axonogenesis / calyx of Held / nitric-oxide synthase regulator activity / actin filament / adherens junction / cell motility / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / Schaffer collateral - CA1 synapse / cytoplasmic ribonucleoprotein granule / nucleosome / lamellipodium / actin cytoskeleton / cytoskeleton / regulation of cell cycle / ribonucleoprotein complex / axon / focal adhesion / synapse / protein kinase binding / glutamatergic synapse / ATP hydrolysis activity / protein-containing complex / ATP binding / membrane / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |   Amanita phalloides (death cap) Amanita phalloides (death cap) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.59 Å | ||||||||||||
 Authors Authors | Oosterheert W / Blanc FEC / Roy A / Belyy A / Hofnagel O / Hummer G / Bieling P / Raunser S | ||||||||||||
| Funding support |  Germany, European Union, 3 items Germany, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Molecular mechanisms of inorganic-phosphate release from the core and barbed end of actin filaments. Authors: Wout Oosterheert / Florian E C Blanc / Ankit Roy / Alexander Belyy / Micaela Boiero Sanders / Oliver Hofnagel / Gerhard Hummer / Peter Bieling / Stefan Raunser /  Abstract: The release of inorganic phosphate (P) from actin filaments constitutes a key step in their regulated turnover, which is fundamental to many cellular functions. The mechanisms underlying P release ...The release of inorganic phosphate (P) from actin filaments constitutes a key step in their regulated turnover, which is fundamental to many cellular functions. The mechanisms underlying P release from the core and barbed end of actin filaments remain unclear. Here, using human and bovine actin isoforms, we combine cryo-EM with molecular-dynamics simulations and in vitro reconstitution to demonstrate how actin releases P through a 'molecular backdoor'. While constantly open at the barbed end, the backdoor is predominantly closed in filament-core subunits and opens only transiently through concerted amino acid rearrangements. This explains why P escapes rapidly from the filament end but slowly from internal subunits. In a nemaline-myopathy-associated actin variant, the backdoor is predominantly open in filament-core subunits, resulting in accelerated P release and filaments with drastically shortened ADP-P caps. Our results provide the molecular basis for P release from actin and exemplify how a disease-linked mutation distorts the nucleotide-state distribution and atomic structure of the filament. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16887.map.gz emd_16887.map.gz | 203.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16887-v30.xml emd-16887-v30.xml emd-16887.xml emd-16887.xml | 24.2 KB 24.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16887_fsc.xml emd_16887_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_16887.png emd_16887.png | 90.9 KB | ||
| Masks |  emd_16887_msk_1.map emd_16887_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16887.cif.gz emd-16887.cif.gz | 7.2 KB | ||
| Others |  emd_16887_additional_1.map.gz emd_16887_additional_1.map.gz emd_16887_half_map_1.map.gz emd_16887_half_map_1.map.gz emd_16887_half_map_2.map.gz emd_16887_half_map_2.map.gz | 108.1 MB 200.4 MB 200.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16887 http://ftp.pdbj.org/pub/emdb/structures/EMD-16887 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16887 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16887 | HTTPS FTP |
-Related structure data
| Related structure data |  8oi6MC  8oi8C  8oidC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16887.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16887.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D-refined, sharpened cryo-EM density map of the undecorated barbed end of F-actin. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.21 Å | ||||||||||||||||||||||||||||||||||||
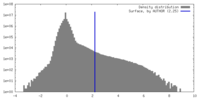
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16887_msk_1.map emd_16887_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: 3D-refined, unsharpened cryo-EM density map of the undecorated...
| File | emd_16887_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D-refined, unsharpened cryo-EM density map of the undecorated barbed end of F-actin. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2 of the refinement of the...
| File | emd_16887_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 of the refinement of the undecorated barbed end of F-actin. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1 of the refinement of the...
| File | emd_16887_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 of the refinement of the undecorated barbed end of F-actin. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : actin-phalloidin complex
| Entire | Name: actin-phalloidin complex |
|---|---|
| Components |
|
-Supramolecule #1: actin-phalloidin complex
| Supramolecule | Name: actin-phalloidin complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: Actin was purified as monomer from bovine thymus. Short filaments were reconstituted in vitro to obtain the barbed end structure. |
|---|
-Supramolecule #2: cytosolic beta-gamma actin
| Supramolecule | Name: cytosolic beta-gamma actin / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: phalloidin
| Supramolecule | Name: phalloidin / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Amanita phalloides (death cap) Amanita phalloides (death cap) |
-Macromolecule #1: Actin, cytoplasmic 1
| Macromolecule | Name: Actin, cytoplasmic 1 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.79568 KDa |
| Sequence | String: MDDDIAALVV DNGSGMCKAG FAGDDAPRAV FPSIVGRPRH QGVMVGMGQK DSYVGDEAQS KRGILTLKYP IE(HIC)GIV TNW DDMEKIWHHT FYNELRVAPE EHPVLLTEAP LNPKANREKM TQIMFETFNT PAMYVAIQAV LSLYASGRTT GIVMDSG DG VTHTVPIYEG ...String: MDDDIAALVV DNGSGMCKAG FAGDDAPRAV FPSIVGRPRH QGVMVGMGQK DSYVGDEAQS KRGILTLKYP IE(HIC)GIV TNW DDMEKIWHHT FYNELRVAPE EHPVLLTEAP LNPKANREKM TQIMFETFNT PAMYVAIQAV LSLYASGRTT GIVMDSG DG VTHTVPIYEG YALPHAILRL DLAGRDLTDY LMKILTERGY SFTTTAEREI VRDIKEKLCY VALDFEQEMA TAASSSSL E KSYELPDGQV ITIGNERFRC PEALFQPSFL GMESCGIHET TFNSIMKCDV DIRKDLYANT VLSGGTTMYP GIADRMQKE ITALAPSTMK IKIIAPPERK YSVWIGGSIL ASLSTFQQMW ISKQEYDESG PSIVHRKCF UniProtKB: Actin, cytoplasmic 1 |
-Macromolecule #2: PHALLOIDIN
| Macromolecule | Name: PHALLOIDIN / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Amanita phalloides (death cap) Amanita phalloides (death cap) |
| Molecular weight | Theoretical: 808.899 Da |
| Sequence | String: W(EEP)A(DTH)C(HYP)A |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.1 Component:
Details: 10 mM imidazole pH 7.1, 100 mM KCl, 2 mM MgCl2 and 1 mM EGTA | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 286 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | Sample was directly collected from size-exclusion chromatography fractions. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Details | Microsope alignment was performed using SHERPA (Thermo Fisher) |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Number grids imaged: 1 / Number real images: 1316 / Average exposure time: 4.0 sec. / Average electron dose: 56.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 120000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: c / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | Real space refinement in phenix. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-8oi6: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)