[English] 日本語
 Yorodumi
Yorodumi- EMDB-16834: Structure of the mammalian Pol II-SPT6-Elongin complex, lacking E... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the mammalian Pol II-SPT6-Elongin complex, lacking ELOA latch (SPT6 focused map for composite map 2) | ||||||||||||
 Map data Map data | focused map for SPT6 in composite map 2 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Transcription elongation / Elongin / RNA polymerase II / Cryo-EM / TRANSCRIPTION | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | ||||||||||||
 Authors Authors | Chen Y / Kokic G / Dienemann C / Dybkov O / Urlaub H / Cramer P | ||||||||||||
| Funding support | European Union,  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structure of the transcribing RNA polymerase II-Elongin complex. Authors: Ying Chen / Goran Kokic / Christian Dienemann / Olexandr Dybkov / Henning Urlaub / Patrick Cramer /   Abstract: Elongin is a heterotrimeric elongation factor for RNA polymerase (Pol) II transcription that is conserved among metazoa. Here, we report three cryo-EM structures of human Elongin bound to ...Elongin is a heterotrimeric elongation factor for RNA polymerase (Pol) II transcription that is conserved among metazoa. Here, we report three cryo-EM structures of human Elongin bound to transcribing Pol II. The structures show that Elongin subunit ELOA binds the RPB2 side of Pol II and anchors the ELOB-ELOC subunit heterodimer. ELOA contains a 'latch' that binds between the end of the Pol II bridge helix and funnel helices, thereby inducing a conformational change near the polymerase active center. The latch is required for the elongation-stimulatory activity of Elongin, but not for Pol II binding, indicating that Elongin functions by allosterically regulating the conformational mobility of the polymerase active center. Elongin binding to Pol II is incompatible with association of the super elongation complex, PAF1 complex and RTF1, which also contain an elongation-stimulatory latch element. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16834.map.gz emd_16834.map.gz | 300.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16834-v30.xml emd-16834-v30.xml emd-16834.xml emd-16834.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16834_fsc.xml emd_16834_fsc.xml | 15.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_16834.png emd_16834.png | 58.8 KB | ||
| Masks |  emd_16834_msk_1.map emd_16834_msk_1.map | 325 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16834.cif.gz emd-16834.cif.gz | 4.1 KB | ||
| Others |  emd_16834_half_map_1.map.gz emd_16834_half_map_1.map.gz emd_16834_half_map_2.map.gz emd_16834_half_map_2.map.gz | 260.6 MB 261.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16834 http://ftp.pdbj.org/pub/emdb/structures/EMD-16834 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16834 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16834 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16834.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16834.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | focused map for SPT6 in composite map 2 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
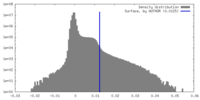
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16834_msk_1.map emd_16834_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
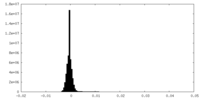
| Density Histograms |
-Half map: half map 2
| File | emd_16834_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
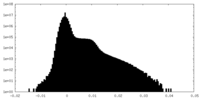
| Density Histograms |
-Half map: half map 1
| File | emd_16834_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
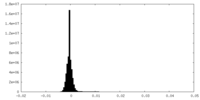
| Density Histograms |
- Sample components
Sample components
-Entire : Transcription elongation complex consisting of RNA Polymerase II,...
| Entire | Name: Transcription elongation complex consisting of RNA Polymerase II, SPT6 and Elongin |
|---|---|
| Components |
|
-Supramolecule #1: Transcription elongation complex consisting of RNA Polymerase II,...
| Supramolecule | Name: Transcription elongation complex consisting of RNA Polymerase II, SPT6 and Elongin type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R3.5/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2.1 |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.09 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 7.5 µm / Nominal defocus min: 0.35000000000000003 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)