+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Three-dimensional Structure of Tropism-Switching Bordetella Bacteriophage | |||||||||
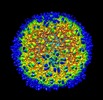
 マップデータ マップデータ | 2f 3D map of phage | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | cryoEM / cryo-electron microscopy / BPP / Bordetella plus phage | |||||||||
| 生物種 |  Bordetella phage BPP-1 (ファージ) Bordetella phage BPP-1 (ファージ) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 7.0 Å | |||||||||
 データ登録者 データ登録者 | Dai W / Hodes A / Hui WH / Gingery M / Miller JF / Zhou ZH | |||||||||
 引用 引用 |  ジャーナル: Proc Natl Acad Sci U S A / 年: 2010 ジャーナル: Proc Natl Acad Sci U S A / 年: 2010タイトル: Three-dimensional structure of tropism-switching Bordetella bacteriophage. 著者: Wei Dai / Asher Hodes / Wong H Hui / Mari Gingery / Jeff F Miller / Z Hong Zhou /  要旨: Bacteriophage BPP-1, which infects Bordetella species, can switch its specificity by mutations to the ligand-binding surface of its major tropism-determinant protein, Mtd. This targeted mutagenesis ...Bacteriophage BPP-1, which infects Bordetella species, can switch its specificity by mutations to the ligand-binding surface of its major tropism-determinant protein, Mtd. This targeted mutagenesis results from the activity of a phage-encoded diversity-generating retroelement. Purified Mtd binds its receptor with low affinity, yet BPP-1 binding and infection of Bordettella cells are efficient because of high-avidity binding between phage-associated Mtd and its receptor. Here, using an integrative approach of three-dimensional (3D) structural analyses of the entire phage by cryo-electron tomography and single-prticle cryo-electron microscopy, we provide direct localization of Mtd in the phage and the structural basis of the high-avidity binding of the BPP-1 phage. Our structure shows that each BPP-1 particle has a T = 7 icosahedral head and an unusual tail apparatus consisting of a short central tail "hub," six short tail spikes, and six extended tail fibers. Subtomographic averaging of the tail fiber maps revealed a two-lobed globular structure at the distal end of each long tail fiber. Tomographic reconstructions of immuno-gold-labeled BPP-1 directly localized Mtd to these globular structures. Finally, our icosahedral reconstruction of the BPP-1 head at 7A resolution reveals an HK97-like major capsid protein stabilized by a smaller cementing protein. Our structure represents a unique bacteriophage reconstruction with its tail fibers and ligand-binding domains shown in relation to its tail apparatus. The localization of Mtd at the distal ends of the six tail fibers explains the high avidity binding of Mtd molecules to cell surfaces for initiation of infection. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|---|
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_1620.map.gz emd_1620.map.gz | 450.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-1620-v30.xml emd-1620-v30.xml emd-1620.xml emd-1620.xml | 8.3 KB 8.3 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  1620.png 1620.png | 337.5 KB | ||
| その他 |  emd_1620_additional_1.map.gz emd_1620_additional_1.map.gz | 450.5 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1620 http://ftp.pdbj.org/pub/emdb/structures/EMD-1620 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1620 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1620 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_1620.map.gz / 形式: CCP4 / 大きさ: 881.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_1620.map.gz / 形式: CCP4 / 大きさ: 881.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | 2f 3D map of phage | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 これらの図は立方格子座標系で作成されたものです | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X: 0.97163 Å / Y: 0.97163 Å / Z: 0.9742 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-添付マップデータ: emd 1620 additional 1.map
| ファイル | emd_1620_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Bordetella Bacteriophage
| 全体 | 名称: Bordetella Bacteriophage |
|---|---|
| 要素 |
|
-超分子 #1000: Bordetella Bacteriophage
| 超分子 | 名称: Bordetella Bacteriophage / タイプ: sample / ID: 1000 / 集合状態: icosahedral particle of whole virus / Number unique components: 1 |
|---|
-超分子 #1: Bordetella phage BPP-1
| 超分子 | 名称: Bordetella phage BPP-1 / タイプ: virus / ID: 1 / Name.synonym: Bordetella Bacteriophage / NCBI-ID: 194699 / 生物種: Bordetella phage BPP-1 / ウイルスタイプ: VIRION / ウイルス・単離状態: STRAIN / ウイルス・エンベロープ: No / ウイルス・中空状態: No / Syn species name: Bordetella Bacteriophage |
|---|---|
| 宿主 | 生物種:  Bordetella (バクテリア) / 別称: BACTERIA(EUBACTERIA) Bordetella (バクテリア) / 別称: BACTERIA(EUBACTERIA) |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 凍結 | 凍結剤: ETHANE / チャンバー内温度: 100 K / 装置: HOMEMADE PLUNGER 詳細: Vitrification instrument: lab-made plunger. Vitrification was carried out at room temperature. CPV were embedded in a thin layer of vitreous ice suspended across the holes of holey carbon ...詳細: Vitrification instrument: lab-made plunger. Vitrification was carried out at room temperature. CPV were embedded in a thin layer of vitreous ice suspended across the holes of holey carbon films for cryoEM imaging. 手法: blot for 3 seconds with filter paper before plunging |
|---|
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI POLARA 300 |
|---|---|
| 撮影 | カテゴリ: CCD / フィルム・検出器のモデル: GENERIC TVIPS / 平均電子線量: 20 e/Å2 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 倍率(補正後): 154380 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.0 mm / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 1.0 µm / 倍率(公称値): 154380 |
| 試料ステージ | 試料ホルダー: Eucentric / 試料ホルダーモデル: GATAN LIQUID NITROGEN |
| 実験機器 |  モデル: Tecnai Polara / 画像提供: FEI Company |
- 画像解析
画像解析
| CTF補正 | 詳細: each particle |
|---|---|
| 最終 再構成 | 想定した対称性 - 点群: I (正20面体型対称) / アルゴリズム: OTHER / 解像度のタイプ: BY AUTHOR / 解像度: 7.0 Å / 解像度の算出法: FSC 0.5 CUT-OFF / ソフトウェア - 名称: IMIRS 詳細: Determination of particle orientation and center parameters and subsequent 3D reconstruction were carried out using programs in the IMIRS software package, which are based on Fourier common ...詳細: Determination of particle orientation and center parameters and subsequent 3D reconstruction were carried out using programs in the IMIRS software package, which are based on Fourier common lines and Fourier-Bessel synthesis methods. 使用した粒子像数: 9000 |
 ムービー
ムービー コントローラー
コントローラー











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)