[English] 日本語
 Yorodumi
Yorodumi- EMDB-15864: Cryo-EM map of lincomycin bound to the Listeria monocytogenes 50S... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of lincomycin bound to the Listeria monocytogenes 50S ribosomal subunit. | |||||||||
 Map data Map data | post-processed masked map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribosome / Listeria monocytogenes / Lincomycin / 50S / antibiotic | |||||||||
| Biological species |  Listeria monocytogenes E (bacteria) Listeria monocytogenes E (bacteria) | |||||||||
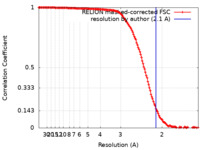
| Method | single particle reconstruction / cryo EM / Resolution: 2.1 Å | |||||||||
 Authors Authors | Koller TO / Crowe-McAuliffe C / Wilson DN | |||||||||
| Funding support |  Sweden, 2 items Sweden, 2 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2022 Journal: Nucleic Acids Res / Year: 2022Title: Structural basis for HflXr-mediated antibiotic resistance in Listeria monocytogenes. Authors: Timm O Koller / Kathryn J Turnbull / Karolis Vaitkevicius / Caillan Crowe-McAuliffe / Mohammad Roghanian / Ondřej Bulvas / Jose A Nakamoto / Tatsuaki Kurata / Christina Julius / Gemma C ...Authors: Timm O Koller / Kathryn J Turnbull / Karolis Vaitkevicius / Caillan Crowe-McAuliffe / Mohammad Roghanian / Ondřej Bulvas / Jose A Nakamoto / Tatsuaki Kurata / Christina Julius / Gemma C Atkinson / Jörgen Johansson / Vasili Hauryliuk / Daniel N Wilson /      Abstract: HflX is a ubiquitous bacterial GTPase that splits and recycles stressed ribosomes. In addition to HflX, Listeria monocytogenes contains a second HflX homolog, HflXr. Unlike HflX, HflXr confers ...HflX is a ubiquitous bacterial GTPase that splits and recycles stressed ribosomes. In addition to HflX, Listeria monocytogenes contains a second HflX homolog, HflXr. Unlike HflX, HflXr confers resistance to macrolide and lincosamide antibiotics by an experimentally unexplored mechanism. Here, we have determined cryo-EM structures of L. monocytogenes HflXr-50S and HflX-50S complexes as well as L. monocytogenes 70S ribosomes in the presence and absence of the lincosamide lincomycin. While the overall geometry of HflXr on the 50S subunit is similar to that of HflX, a loop within the N-terminal domain of HflXr, which is two amino acids longer than in HflX, reaches deeper into the peptidyltransferase center. Moreover, unlike HflX, the binding of HflXr induces conformational changes within adjacent rRNA nucleotides that would be incompatible with drug binding. These findings suggest that HflXr confers resistance using an allosteric ribosome protection mechanism, rather than by simply splitting and recycling antibiotic-stalled ribosomes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15864.map.gz emd_15864.map.gz | 33.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15864-v30.xml emd-15864-v30.xml emd-15864.xml emd-15864.xml | 26.1 KB 26.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15864_fsc.xml emd_15864_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_15864.png emd_15864.png | 89.6 KB | ||
| Filedesc metadata |  emd-15864.cif.gz emd-15864.cif.gz | 4.7 KB | ||
| Others |  emd_15864_additional_1.map.gz emd_15864_additional_1.map.gz emd_15864_additional_2.map.gz emd_15864_additional_2.map.gz emd_15864_half_map_1.map.gz emd_15864_half_map_1.map.gz emd_15864_half_map_2.map.gz emd_15864_half_map_2.map.gz | 193.2 MB 223.3 MB 193.6 MB 193.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15864 http://ftp.pdbj.org/pub/emdb/structures/EMD-15864 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15864 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15864 | HTTPS FTP |
-Validation report
| Summary document |  emd_15864_validation.pdf.gz emd_15864_validation.pdf.gz | 941.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15864_full_validation.pdf.gz emd_15864_full_validation.pdf.gz | 941.4 KB | Display | |
| Data in XML |  emd_15864_validation.xml.gz emd_15864_validation.xml.gz | 21.8 KB | Display | |
| Data in CIF |  emd_15864_validation.cif.gz emd_15864_validation.cif.gz | 28.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15864 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15864 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15864 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15864 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15864.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15864.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | post-processed masked map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.7725 Å | ||||||||||||||||||||||||||||||||||||
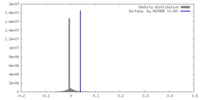
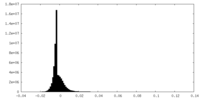
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: 3D refined map
| File | emd_15864_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D refined map | ||||||||||||
| Projections & Slices |
| ||||||||||||
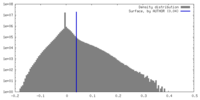
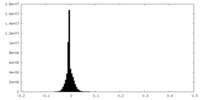
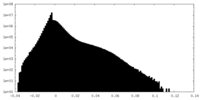
| Density Histograms |
-Additional map: post-processed map
| File | emd_15864_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | post-processed map | ||||||||||||
| Projections & Slices |
| ||||||||||||
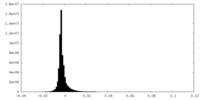
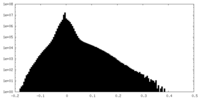
| Density Histograms |
-Half map: half-map1
| File | emd_15864_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
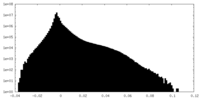
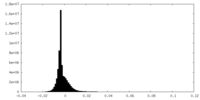
| Density Histograms |
-Half map: half-map2
| File | emd_15864_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
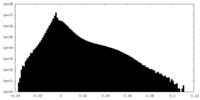
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM map of lincomycin bound to the Listeria monocytogenes 50S...
| Entire | Name: Cryo-EM map of lincomycin bound to the Listeria monocytogenes 50S ribosomal subunit. |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM map of lincomycin bound to the Listeria monocytogenes 50S...
| Supramolecule | Name: Cryo-EM map of lincomycin bound to the Listeria monocytogenes 50S ribosomal subunit. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#29 |
|---|---|
| Source (natural) | Organism:  Listeria monocytogenes E (bacteria) Listeria monocytogenes E (bacteria) |
| Molecular weight | Theoretical: 1 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
Details: 0.05% Nikkol | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 | |||||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||||||||||||||
| Details | 5 OD260/mL |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.4000000000000001 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 270000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)