[English] 日本語
 Yorodumi
Yorodumi- EMDB-15613: RNA polymerase bound to purified in vitro transcribed regulatory ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | RNA polymerase bound to purified in vitro transcribed regulatory RNA putL - inactive, open clamp state | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | RNA polymerase / transcriptional pausing / transcription termination / regulatory RNA / TRANSCRIPTION | |||||||||||||||
| Biological species |  | |||||||||||||||
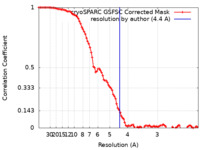
| Method | single particle reconstruction / cryo EM / Resolution: 4.4 Å | |||||||||||||||
 Authors Authors | Dey S / Weixlbaumer A | |||||||||||||||
| Funding support | European Union,  France, 4 items France, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2022 Journal: Mol Cell / Year: 2022Title: Structural insights into RNA-mediated transcription regulation in bacteria. Authors: Sanjay Dey / Claire Batisse / Jinal Shukla / Michael W Webster / Maria Takacs / Charlotte Saint-André / Albert Weixlbaumer /  Abstract: RNA can regulate its own synthesis without auxiliary proteins. For example, U-rich RNA sequences signal RNA polymerase (RNAP) to pause transcription and are required for transcript release at ...RNA can regulate its own synthesis without auxiliary proteins. For example, U-rich RNA sequences signal RNA polymerase (RNAP) to pause transcription and are required for transcript release at intrinsic terminators in all kingdoms of life. In contrast, the regulatory RNA putL suppresses pausing and termination in cis. However, how nascent RNA modulates its own synthesis remains largely unknown. We present cryo-EM reconstructions of RNAP captured during transcription of putL variants or an unrelated sequence at a U-rich pause site. Our results suggest how putL suppresses pausing and promotes its synthesis. We demonstrate that transcribing a U-rich sequence, a ubiquitous trigger of intrinsic termination, promotes widening of the RNAP nucleic-acid-binding channel. Widening destabilizes RNAP interactions with DNA and RNA to facilitate transcript dissociation reminiscent of intrinsic transcription termination. Surprisingly, RNAP remains bound to DNA after transcript release. Our results provide the structural framework to understand RNA-mediated intrinsic transcription termination. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15613.map.gz emd_15613.map.gz | 79.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15613-v30.xml emd-15613-v30.xml emd-15613.xml emd-15613.xml | 16.4 KB 16.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15613_fsc.xml emd_15613_fsc.xml | 9.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_15613.png emd_15613.png | 41.1 KB | ||
| Filedesc metadata |  emd-15613.cif.gz emd-15613.cif.gz | 4.6 KB | ||
| Others |  emd_15613_additional_1.map.gz emd_15613_additional_1.map.gz emd_15613_half_map_1.map.gz emd_15613_half_map_1.map.gz emd_15613_half_map_2.map.gz emd_15613_half_map_2.map.gz | 41.7 MB 77.7 MB 77.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15613 http://ftp.pdbj.org/pub/emdb/structures/EMD-15613 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15613 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15613 | HTTPS FTP |
-Validation report
| Summary document |  emd_15613_validation.pdf.gz emd_15613_validation.pdf.gz | 852.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15613_full_validation.pdf.gz emd_15613_full_validation.pdf.gz | 851.7 KB | Display | |
| Data in XML |  emd_15613_validation.xml.gz emd_15613_validation.xml.gz | 17.6 KB | Display | |
| Data in CIF |  emd_15613_validation.cif.gz emd_15613_validation.cif.gz | 22.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15613 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15613 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15613 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15613 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15613.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15613.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_15613_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
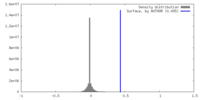
| Density Histograms |
-Half map: #1
| File | emd_15613_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
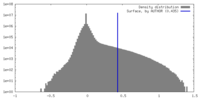
| Density Histograms |
-Half map: #2
| File | emd_15613_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
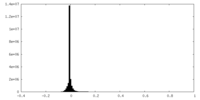
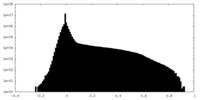
| Density Histograms |
- Sample components
Sample components
-Entire : Escherichia coli RNA polymerase and in vitro transcribed and puri...
| Entire | Name: Escherichia coli RNA polymerase and in vitro transcribed and purified putL RNA complex - inactive, open clamp state |
|---|---|
| Components |
|
-Supramolecule #1: Escherichia coli RNA polymerase and in vitro transcribed and puri...
| Supramolecule | Name: Escherichia coli RNA polymerase and in vitro transcribed and purified putL RNA complex - inactive, open clamp state type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 435 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 12 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM Tris-glutamate pH 8.0, 50 mM K-glutamate, 10 mM Mg-glutamate, 0.001 mM ZnCl2, 2mM DTT |
| Grid | Model: C-flat-1.2/1.3 / Material: GOLD / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV Details: Quantifoil UltrAuFoil R1.2/1.3 300 mesh holey gold grids were plasma cleaned on a Model 1070 (Fischione Instruments) for 30 sec at 70% power and with an 80% Argon and 20% Oxygen mixture ...Details: Quantifoil UltrAuFoil R1.2/1.3 300 mesh holey gold grids were plasma cleaned on a Model 1070 (Fischione Instruments) for 30 sec at 70% power and with an 80% Argon and 20% Oxygen mixture prior to the application of 0.004 ml of sample. Grids were plunge frozen into liquid ethane using a Vitrobot mark IV (FEI) with 95% chamber humidity at 283K.. |
| Details | The complex for cryo-EM analysis was prepared in vitro by mixing E. coli RNAP with DNA oligonucleotides and in vitro transcribed and purified putL RNA to mimic an RNAP EC halted at the U-rich pause (G93). The complex was prepared in 20 mM Tris-glutamate pH 8.0, 50 mM K-glutamate, 10 mM Mg-glutamate, 0.001 mM ZnCl2, and 2mM DTT. The final complex was purified on a Superose 6 Increase 3.2/300 gel filtration column equilibrated in Glutamate buffer. Samples were concentrated to 10-12 mg/mL using an Amicon Ultra 0.5mL centrifugal filter unit (30 KDa MWCO). Before grid freezing, 8 mM of CHAPSO was added to freshly prepared sample to overcome preferred particle orientation. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Average exposure time: 2.2 sec. / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)