[English] 日本語
 Yorodumi
Yorodumi- EMDB-14796: CryoEM structure of mitochondrial complex I from Chaetomium therm... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of mitochondrial complex I from Chaetomium thermophilum (state 2) - membrane arm | |||||||||
 Map data Map data | Density modified map (Terwilliger et al., 2020) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Proton transporter / Mitochondrial membrane protein / Complex / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology information5'-flap endonuclease activity / NADH dehydrogenase complex / replication fork reversal / mitochondrial [2Fe-2S] assembly complex / single-stranded DNA helicase activity / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / electron transport coupled proton transport / mitochondrial electron transport, NADH to ubiquinone / acyl binding ...5'-flap endonuclease activity / NADH dehydrogenase complex / replication fork reversal / mitochondrial [2Fe-2S] assembly complex / single-stranded DNA helicase activity / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / electron transport coupled proton transport / mitochondrial electron transport, NADH to ubiquinone / acyl binding / mitochondrial respiratory chain complex I assembly / NADH dehydrogenase activity / oxidoreductase activity, acting on NAD(P)H / respiratory chain complex I / NADH dehydrogenase (ubiquinone) activity / acyl carrier activity / ATP synthesis coupled electron transport / aerobic respiration / electron transport chain / mitochondrial intermembrane space / mitochondrial membrane / 4 iron, 4 sulfur cluster binding / DNA helicase / mitochondrial inner membrane / DNA repair / mitochondrion / DNA binding / ATP binding / membrane / metal ion binding / nucleus Similarity search - Function | |||||||||
| Biological species |  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.83 Å | |||||||||
 Authors Authors | Laube E / Kuehlbrandt W | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Conformational changes in mitochondrial complex I of the thermophilic eukaryote . Authors: Eike Laube / Jakob Meier-Credo / Julian D Langer / Werner Kühlbrandt /  Abstract: Mitochondrial complex I is a redox-driven proton pump that generates proton-motive force across the inner mitochondrial membrane, powering oxidative phosphorylation and ATP synthesis in eukaryotes. ...Mitochondrial complex I is a redox-driven proton pump that generates proton-motive force across the inner mitochondrial membrane, powering oxidative phosphorylation and ATP synthesis in eukaryotes. We report the structure of complex I from the thermophilic fungus , determined by cryoEM up to 2.4-Å resolution. We show that the complex undergoes a transition between two conformations, which we refer to as state 1 and state 2. The conformational switch is manifest in a twisting movement of the peripheral arm relative to the membrane arm, but most notably in substantial rearrangements of the Q-binding cavity and the E-channel, resulting in a continuous aqueous passage from the E-channel to subunit ND5 at the far end of the membrane arm. The conformational changes in the complex interior resemble those reported for mammalian complex I, suggesting a highly conserved, universal mechanism of coupling electron transport to proton pumping. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14796.map.gz emd_14796.map.gz | 43.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14796-v30.xml emd-14796-v30.xml emd-14796.xml emd-14796.xml | 53 KB 53 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14796_fsc.xml emd_14796_fsc.xml | 20.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_14796.png emd_14796.png | 109.5 KB | ||
| Filedesc metadata |  emd-14796.cif.gz emd-14796.cif.gz | 12 KB | ||
| Others |  emd_14796_additional_1.map.gz emd_14796_additional_1.map.gz emd_14796_half_map_1.map.gz emd_14796_half_map_1.map.gz emd_14796_half_map_2.map.gz emd_14796_half_map_2.map.gz | 388.4 MB 720.4 MB 720.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14796 http://ftp.pdbj.org/pub/emdb/structures/EMD-14796 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14796 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14796 | HTTPS FTP |
-Related structure data
| Related structure data |  7zmeMC  7zm7C  7zm8C  7zmbC  7zmgC  7zmhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14796.map.gz / Format: CCP4 / Size: 46.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14796.map.gz / Format: CCP4 / Size: 46.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Density modified map (Terwilliger et al., 2020) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.837 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: unmodified/unsharpened map
| File | emd_14796_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unmodified/unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
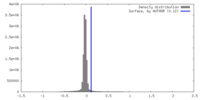
| Density Histograms |
-Half map: EM half map A
| File | emd_14796_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM_half_map_A | ||||||||||||
| Projections & Slices |
| ||||||||||||
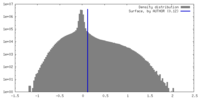
| Density Histograms |
-Half map: EM half map B
| File | emd_14796_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM_half_map_B | ||||||||||||
| Projections & Slices |
| ||||||||||||
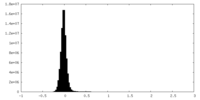
| Density Histograms |
- Sample components
Sample components
+Entire : Mitochondrial NADH:ubiquinone oxidoreductase in LMNG
+Supramolecule #1: Mitochondrial NADH:ubiquinone oxidoreductase in LMNG
+Macromolecule #1: NADH-ubiquinone oxidoreductase chain 1
+Macromolecule #2: NADH dehydrogenase subunit 2
+Macromolecule #3: NADH-ubiquinone oxidoreductase chain 3
+Macromolecule #4: NADH-ubiquinone oxidoreductase chain 4
+Macromolecule #5: NADH-ubiquinone oxidoreductase chain 5
+Macromolecule #6: NADH-ubiquinone oxidoreductase chain 6
+Macromolecule #7: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 7
+Macromolecule #8: Subunit NDUFS5 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #9: Subunit NDUFA1 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #10: NADH-ubiquinone oxidoreductase-like protein
+Macromolecule #11: NADH-ubiquinone oxidoreductase chain 4L
+Macromolecule #12: Acyl carrier protein
+Macromolecule #13: Complex I-B22
+Macromolecule #14: Complex I-ESSS
+Macromolecule #15: NADH-ubiquinone oxidoreductase
+Macromolecule #16: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 13
+Macromolecule #17: NADH-ubiquinone oxidoreductase-like protein
+Macromolecule #18: NADH dehydrogenase (Ubiquinone)-like protein
+Macromolecule #19: Subunit NDUFC2 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #20: Subunit NDUFB3 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #21: Subunit NDUFB10 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #22: Subunit NDUFB2 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #23: Subunit NDUFA3 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #24: Subunit NDUFB6 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #25: Subunit NDUFB4 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #26: Subunit NDUFB5 of NADH-ubiquinone oxidoreductase (Complex I)
+Macromolecule #27: 1,2-Distearoyl-sn-glycerophosphoethanolamine
+Macromolecule #28: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+Macromolecule #29: CARDIOLIPIN
+Macromolecule #30: Lauryl Maltose Neopentyl Glycol
+Macromolecule #31: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
+Macromolecule #32: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 30 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 6503 / Average exposure time: 2.11 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)