[English] 日本語
 Yorodumi
Yorodumi- EMDB-14625: Subtomogram averaging of Rubisco from Cyanobium carboxysome (oute... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram averaging of Rubisco from Cyanobium carboxysome (outerlayer) | ||||||||||||
 Map data Map data | main map from cisTEM reconstruction within emClarity package, postprocessed with relion | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |  Cyanobium sp. PCC 7001 (bacteria) Cyanobium sp. PCC 7001 (bacteria) | ||||||||||||
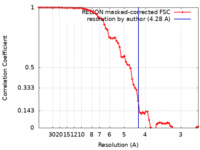
| Method | subtomogram averaging / cryo EM / Resolution: 4.28 Å | ||||||||||||
 Authors Authors | Ni T / Zhu Y / Seaton-Burn W / Zhang P | ||||||||||||
| Funding support |  United Kingdom, European Union, 3 items United Kingdom, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structure and assembly of cargo Rubisco in two native α-carboxysomes. Authors: Tao Ni / Yaqi Sun / Will Burn / Monsour M J Al-Hazeem / Yanan Zhu / Xiulian Yu / Lu-Ning Liu / Peijun Zhang /   Abstract: Carboxysomes are a family of bacterial microcompartments in cyanobacteria and chemoautotrophs. They encapsulate Ribulose 1,5-bisphosphate carboxylase/oxygenase (Rubisco) and carbonic anhydrase ...Carboxysomes are a family of bacterial microcompartments in cyanobacteria and chemoautotrophs. They encapsulate Ribulose 1,5-bisphosphate carboxylase/oxygenase (Rubisco) and carbonic anhydrase catalyzing carbon fixation inside a proteinaceous shell. How Rubisco complexes pack within the carboxysomes is unknown. Using cryo-electron tomography, we determine the distinct 3D organization of Rubisco inside two distant α-carboxysomes from a marine α-cyanobacterium Cyanobium sp. PCC 7001 where Rubiscos are organized in three concentric layers, and from a chemoautotrophic bacterium Halothiobacillus neapolitanus where they form intertwining spirals. We further resolve the structures of native Rubisco as well as its higher-order assembly at near-atomic resolutions by subtomogram averaging. The structures surprisingly reveal that the authentic intrinsically disordered linker protein CsoS2 interacts with Rubiscos in native carboxysomes but functions distinctively in the two α-carboxysomes. In contrast to the uniform Rubisco-CsoS2 association in the Cyanobium α-carboxysome, CsoS2 binds only to the Rubiscos close to the shell in the Halo α-carboxysome. Our findings provide critical knowledge of the assembly principles of α-carboxysomes, which may aid in the rational design and repurposing of carboxysome structures for new functions. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14625.map.gz emd_14625.map.gz | 31 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14625-v30.xml emd-14625-v30.xml emd-14625.xml emd-14625.xml | 14.2 KB 14.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14625_fsc.xml emd_14625_fsc.xml | 7.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_14625.png emd_14625.png | 99.4 KB | ||
| Others |  emd_14625_half_map_1.map.gz emd_14625_half_map_1.map.gz emd_14625_half_map_2.map.gz emd_14625_half_map_2.map.gz | 20.2 MB 20.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14625 http://ftp.pdbj.org/pub/emdb/structures/EMD-14625 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14625 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14625 | HTTPS FTP |
-Validation report
| Summary document |  emd_14625_validation.pdf.gz emd_14625_validation.pdf.gz | 779.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14625_full_validation.pdf.gz emd_14625_full_validation.pdf.gz | 779.4 KB | Display | |
| Data in XML |  emd_14625_validation.xml.gz emd_14625_validation.xml.gz | 13.2 KB | Display | |
| Data in CIF |  emd_14625_validation.cif.gz emd_14625_validation.cif.gz | 18.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14625 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14625 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14625 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14625 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14625.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14625.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map from cisTEM reconstruction within emClarity package, postprocessed with relion | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: emClarity half map 1 from cisTEM reconstruction
| File | emd_14625_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | emClarity half map 1 from cisTEM reconstruction | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: emClarity half map 1 from cisTEM reconstruction
| File | emd_14625_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | emClarity half map 1 from cisTEM reconstruction | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of Rubisco within Cyanobium carboxysome
| Entire | Name: Structure of Rubisco within Cyanobium carboxysome |
|---|---|
| Components |
|
-Supramolecule #1: Structure of Rubisco within Cyanobium carboxysome
| Supramolecule | Name: Structure of Rubisco within Cyanobium carboxysome / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Cyanobium sp. PCC 7001 (bacteria) Cyanobium sp. PCC 7001 (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 3.65 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 2.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)