+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-14170 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Low resolution negative stain map of SslE | |||||||||
 Map data Map data | Low resolution map of SslE | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | metalloprotease / biofilm formation / MEMBRANE PROTEIN | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 22.0 Å | |||||||||
 Authors Authors | Fenn K / Darbari VC / Garnett JA / Corsini PM | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: NPJ Biofilms Microbiomes / Year: 2022 Journal: NPJ Biofilms Microbiomes / Year: 2022Title: Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation. Authors: Paula M Corsini / Sunjun Wang / Saima Rehman / Katherine Fenn / Amin Sagar / Slobodan Sirovica / Leanne Cleaver / Charlotte J C Edwards-Gayle / Giulia Mastroianni / Ben Dorgan / Lee M Sewell ...Authors: Paula M Corsini / Sunjun Wang / Saima Rehman / Katherine Fenn / Amin Sagar / Slobodan Sirovica / Leanne Cleaver / Charlotte J C Edwards-Gayle / Giulia Mastroianni / Ben Dorgan / Lee M Sewell / Steven Lynham / Dinu Iuga / W Trent Franks / James Jarvis / Guy H Carpenter / Michael A Curtis / Pau Bernadó / Vidya C Darbari / James A Garnett /   Abstract: Escherichia coli is a Gram-negative bacterium that colonises the human intestine and virulent strains can cause severe diarrhoeal and extraintestinal diseases. The protein SslE is secreted by a range ...Escherichia coli is a Gram-negative bacterium that colonises the human intestine and virulent strains can cause severe diarrhoeal and extraintestinal diseases. The protein SslE is secreted by a range of pathogenic and commensal E. coli strains. It can degrade mucins in the intestine, promotes biofilm maturation and it is a major determinant of infection in virulent strains, although how it carries out these functions is not well understood. Here, we examine SslE from the commensal E. coli Waksman and BL21 (DE3) strains and the enterotoxigenic H10407 and enteropathogenic E2348/69 strains. We reveal that SslE has a unique and dynamic structure in solution and in response to acidification within mature biofilms it can form a unique aggregate with amyloid-like properties. Furthermore, we show that both SslE monomers and aggregates bind DNA in vitro and co-localise with extracellular DNA (eDNA) in mature biofilms, and SslE aggregates may also associate with cellulose under certain conditions. Our results suggest that interactions between SslE and eDNA are important for biofilm maturation in many E. coli strains and SslE may also be a factor that drives biofilm formation in other SslE-secreting bacteria. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14170.map.gz emd_14170.map.gz | 292.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14170-v30.xml emd-14170-v30.xml emd-14170.xml emd-14170.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14170.png emd_14170.png | 14.5 KB | ||
| Filedesc metadata |  emd-14170.cif.gz emd-14170.cif.gz | 5.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14170 http://ftp.pdbj.org/pub/emdb/structures/EMD-14170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14170 | HTTPS FTP |
-Validation report
| Summary document |  emd_14170_validation.pdf.gz emd_14170_validation.pdf.gz | 296.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14170_full_validation.pdf.gz emd_14170_full_validation.pdf.gz | 296.2 KB | Display | |
| Data in XML |  emd_14170_validation.xml.gz emd_14170_validation.xml.gz | 5.3 KB | Display | |
| Data in CIF |  emd_14170_validation.cif.gz emd_14170_validation.cif.gz | 6.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14170 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14170 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14170 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14170 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14170.map.gz / Format: CCP4 / Size: 12.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14170.map.gz / Format: CCP4 / Size: 12.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Low resolution map of SslE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
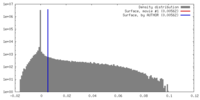
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SslE
| Entire | Name: SslE |
|---|---|
| Components |
|
-Supramolecule #1: SslE
| Supramolecule | Name: SslE / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 160 KDa |
-Macromolecule #1: SslE
| Macromolecule | Name: SslE / type: protein_or_peptide / ID: 1 / Enantiomer: DEXTRO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: CDGGGSGSSS DTPPVDSGTG SLPEVKPDPT PNPEPTPEPT PDPEPTPEPT PDPEPTPEPE PEPVPTKTGY LTLGGSLRVT GDITCNDESS DGFTFTPGDK VTCVAGNNTT IATFDTQSEA ARSLRAVEKV SFSLEDAQEL AGSDNKKSNA LSLVTSMNSC PANTEQVCLE ...String: CDGGGSGSSS DTPPVDSGTG SLPEVKPDPT PNPEPTPEPT PDPEPTPEPT PDPEPTPEPE PEPVPTKTGY LTLGGSLRVT GDITCNDESS DGFTFTPGDK VTCVAGNNTT IATFDTQSEA ARSLRAVEKV SFSLEDAQEL AGSDNKKSNA LSLVTSMNSC PANTEQVCLE FSSVIESKRF DSLYKQIDLA PEEFKKLVNE EVENNAATDK APSTHTSPVV PATTPGTKPD LNASFVSANA EQFYQYQPTE IILSEGRLVD SQGDGVVGVN YYTNSGRGVT GENGEFSFSW GETISFGIDT FELGSVRGNK STIALTELGD EVRGANIDQL IHRYSKAGQN HTRVVPDEVR KVFAEYPNVI NEIINLSLSN GATLGEGEQV VNLPNEFIEQ FKTGQAKEID TAICAKTDGC NEARWFSLTT RNVNDGQIQG VINKLWGVDT NYKSVSKFHV FHDSTNFYGS TGNARGQAVV NISNAAFPIL MARNDKNYWL AFGEKRAWDK NELAYITEAP SIVRPENVTR ETATFNLPFI SLGQVGDGKL MVIGNPHYNS ILRCPNGYSW NGGVNKDGQC TLNSDPDDMK NFMENVLRYL SNDRWLPDAK SSMTVGTNLD TVYFKKHGQV LGNSAPFAFH KDFTGITVKP MTSYGNLNPD EVPLLILNGF EYVTQWGSDP YSIPLRADTS KPKLTQQDVT DLIAYMNKGG SVLIMENVMS NLKEESASGF VRLLDAAGLS MALNKSVVNN DPQGYPDRVR QRRSTPIWVY ERYPAVDGKP PYTIDDTTKE VIWKYQQENK PDDKPKLEVA SWQEEVEGKQ VTQFAFIDEA DHKTPESLAA AKQRILDAFP GLEVCKDSDY HYEVNCLEYR PGTGVPVTGG MYVPQYTQLD LGADTAKAML QAADLGTNIQ RLYQHELYFR TNGRQGERLN SVDLERLYQN MSVWLWNETK YRYEEGKEDE LGFKTFTEFL NCYTNNAYVG TQCSAELKKS LIDNKMIYGE ESSKAGMMNP SYPLNYMEKP LTRLMLGRSW WDLNIKVDVE KYPGAVSEEG QNVTETISLY SNPTKWFAGN MQSTGLWAPA QKEVTIKSNA NVPVTVTVAL ADDLTGREKH EVALNRPPRV TKTYSLDASG TVKFKVPYGG LIYIKGDSKD NESASFTFTG VVKAPFYKDG AWKNDLNSPA PLGELESDAF VYTAPKKNLN ASNYTGGLKQ FANDLDTFAS SMNDFYGRNE EDGKHRMFTY KNLTGHKHRF ANDVQISIGD AHSGYPVMNS SFSTNSTTLP TTPLNDWLIW HEVGHNAAET PLTVPGATEV ANNVLALYMQ DRYLGKMNRV ADDITVAPEY LEESNGQAWA RGGAGDRLLM YAQLKEWAEK NFDIKKWYPE GELPKFFSDR EGMKGWNLFQ LMHRKARGDD VGDKTFGGKN YCAESNGNAA DTLMLCASWV AQTDLSEFFK KWNPGANAYQ LPGASEMSFE GGVSQSAYNT LASLKLPKPE QGPETINKVT EHKMSVE |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 8 / Component - Concentration: 50.0 mM / Component - Formula: Tris-HCl / Component - Name: Tris |
| Staining | Type: NEGATIVE / Material: Uranyl acetate Details: 2% (v/v) uranyl acetate was applied for staining for 1 min. The excess liquid was blotted |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2100 |
|---|---|
| Image recording | Film or detector model: OTHER / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 50000 |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 22.0 Å / Resolution method: OTHER / Number images used: 11000 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)