+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13190 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | P5C3 is a potent fab neutralizer | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | spike / antibody / VIRAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of viral entry into host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / host cell surface receptor binding / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / host cell plasma membrane / virion membrane ...positive regulation of viral entry into host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / host cell surface receptor binding / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / host cell plasma membrane / virion membrane / membrane / identical protein binding Similarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
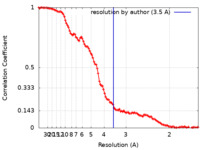
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||
 Authors Authors | perez L | ||||||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2021 Journal: Cell Rep / Year: 2021Title: A highly potent antibody effective against SARS-CoV-2 variants of concern. Authors: Craig Fenwick / Priscilla Turelli / Laurent Perez / Céline Pellaton / Line Esteves-Leuenberger / Alex Farina / Jérémy Campos / Erica Lana / Flurin Fiscalini / Charlène Raclot / Florence ...Authors: Craig Fenwick / Priscilla Turelli / Laurent Perez / Céline Pellaton / Line Esteves-Leuenberger / Alex Farina / Jérémy Campos / Erica Lana / Flurin Fiscalini / Charlène Raclot / Florence Pojer / Kelvin Lau / Davide Demurtas / Marc Descatoire / Victor S Joo / Mathilde Foglierini / Alessandra Noto / Rana Abdelnabi / Caroline S Foo / Laura Vangeel / Johan Neyts / Wenjuan Du / Berend-Jan Bosch / Geertruida Veldman / Pieter Leyssen / Volker Thiel / Roger LeGrand / Yves Lévy / Didier Trono / Giuseppe Pantaleo /      Abstract: Control of the ongoing SARS-CoV-2 pandemic is endangered by the emergence of viral variants with increased transmission efficiency, resistance to marketed therapeutic antibodies, and reduced ...Control of the ongoing SARS-CoV-2 pandemic is endangered by the emergence of viral variants with increased transmission efficiency, resistance to marketed therapeutic antibodies, and reduced sensitivity to vaccine-induced immunity. Here, we screen B cells from COVID-19 donors and identify P5C3, a highly potent and broadly neutralizing monoclonal antibody with picomolar neutralizing activity against all SARS-CoV-2 variants of concern (VOCs) identified to date. Structural characterization of P5C3 Fab in complex with the spike demonstrates a neutralizing activity defined by a large buried surface area, highly overlapping with the receptor-binding domain (RBD) surface necessary for ACE2 interaction. We further demonstrate that P5C3 shows complete prophylactic protection in the SARS-CoV-2-infected hamster challenge model. These results indicate that P5C3 opens exciting perspectives either as a prophylactic agent in immunocompromised individuals with poor response to vaccination or as combination therapy in SARS-CoV-2-infected individuals. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13190.map.gz emd_13190.map.gz | 93.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13190-v30.xml emd-13190-v30.xml emd-13190.xml emd-13190.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13190_fsc.xml emd_13190_fsc.xml | 13.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_13190.png emd_13190.png | 39.6 KB | ||
| Filedesc metadata |  emd-13190.cif.gz emd-13190.cif.gz | 7.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13190 http://ftp.pdbj.org/pub/emdb/structures/EMD-13190 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13190 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13190 | HTTPS FTP |
-Related structure data
| Related structure data |  7p40MC  7phgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13190.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13190.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.64 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
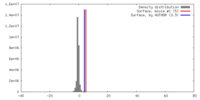
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Severe acute respiratory syndrome coronavirus 2
| Entire | Name:  |
|---|---|
| Components |
|
-Supramolecule #1: Severe acute respiratory syndrome coronavirus 2
| Supramolecule | Name: Severe acute respiratory syndrome coronavirus 2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: Spike glycoprotein
| Supramolecule | Name: Spike glycoprotein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Fab
| Supramolecule | Name: Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 Details: ...Details: MGILPSPGMPALLSLVSLLSVLLMGCVAETGTQCVNLTTRTQLPPAYTNSFTRGVYYPDKVFRSSVLHSTQDLFLPFFSNVTWFHAIHVSGTNGTKRFDNPVLPFNDGVYFASTEKSNIIRGWIFGTTLDSKTQSLLIVNNATNVVIKVCEFQFCNDPFLGVYYHKNNKSWMESEFRVYSSANNCTFEYVSQPFLMDLEGKQGNFKNLREFVFKNIDGYFKIYSKHTPINLVRDLPQGFSALEPLVDLPIGINITRFQTLLALHRSYLTPGDSSSGWTAGAAAYYVGYLQPRTFLLKYNENGTITDAVDCALDPLSETKCTLKSFTVEKGIYQTSNFRVQPTESIVRFPNITNLCPFGEVFNATRFASVYAWNRKRISNCVADYSVLYNSASFSTFKCYGVSPTKLNDLCFTNVYADSFVIRGDEVRQIAPGQTGKIADYNYKLPDDFTGCVIAWNSNNLDSKVGGNYNYLYRLFRKSNLKPFERDISTEIYQAGSTPCNGVEGFNCYFPLQSYGFQPTNGVGYQPYRVVVLSFELLHAPATVCGPKKSTNLVKNKCVNFNFNGLTGTGVLTESNKKFLPFQQFGRDIADTTDAVRDPQTLEILDITPCSFGGVSVITPGTNTSNQVAVLYQDVNCTEVPVAIHADQLTPTWRVYSTGSNVFQTRAGCLIGAEHVNNSYECDIPIGAGICASYQTQTNSPSGAGSVASQSIIAYTMSLGAENSVAYSNNSIAIPTNFTISVTTEILPVSMTKTSVDCTMYICGDSTECSNLLLQYGSFCTQLNRALTGIAVEQDKNTQEVFAQVKQIYKTPPIKDFGGFNFSQILPDPSKPSKRSFIEDLLFNKVTLADAGFIKQYGDCLGDIAARDLICAQKFNGLTVLPPLLTDEMIAQYTSALLAGTITSGWTFGAGAALQIPFAMQMAYRFNGIGVTQNVLYENQKLIANQFNSAIGKIQDSLSSTASALGKLQDVVNQNAQALNTLVKQLSSNFGAISSVLNDILSRLDPPEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRASANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQSAPHGVVFLHVTYVPAQEKNFTTAPAICHDGKAHFPREGVFVSNGTHWFVTQRNFYEPQIITTDNTFVSGNCDVVIGIVNNTVYDPLQPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRLNEVA KNLNESLIDLQELGKYEQYIKGSGRENLYFQGGGGSGYIPEAPRDGQAYVRKDGEWVLLSTFLGHHHHHHHH Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 141.560859 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GTQCVNLTTR TQLPPAYTNS FTRGVYYPDK VFRSSVLHST QDLFLPFFSN VTWFHAIHV SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV CEFQFCNDPF L GVYYHKNN ...String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GTQCVNLTTR TQLPPAYTNS FTRGVYYPDK VFRSSVLHST QDLFLPFFSN VTWFHAIHV SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV CEFQFCNDPF L GVYYHKNN KSWMESEFRV YSSANNCTFE YVSQPFLMDL EGKQGNFKNL REFVFKNIDG YFKIYSKHTP INLVRDLPQG FS ALEPLVD LPIGINITRF QTLLALHRSY LTPGDSSSGW TAGAAAYYVG YLQPRTFLLK YNENGTITDA VDCALDPLSE TKC TLKSFT VEKGIYQTSN FRVQPTESIV RFPNITNLCP FGEVFNATRF ASVYAWNRKR ISNCVADYSV LYNSASFSTF KCYG VSPTK LNDLCFTNVY ADSFVIRGDE VRQIAPGQTG KIADYNYKLP DDFTGCVIAW NSNNLDSKVG GNYNYLYRLF RKSNL KPFE RDISTEIYQA GSTPCNGVEG FNCYFPLQSY GFQPTNGVGY QPYRVVVLSF ELLHAPATVC GPKKSTNLVK NKCVNF NFN GLTGTGVLTE SNKKFLPFQQ FGRDIADTTD AVRDPQTLEI LDITPCSFGG VSVITPGTNT SNQVAVLYQD VNCTEVP VA IHADQLTPTW RVYSTGSNVF QTRAGCLIGA EHVNNSYECD IPIGAGICAS YQTQTNSPSG AGSVASQSII AYTMSLGA E NSVAYSNNSI AIPTNFTISV TTEILPVSMT KTSVDCTMYI CGDSTECSNL LLQYGSFCTQ LNRALTGIAV EQDKNTQEV FAQVKQIYKT PPIKDFGGFN FSQILPDPSK PSKRSPIEDL LFNKVTLADA GFIKQYGDCL GDIAARDLIC AQKFNGLTVL PPLLTDEMI AQYTSALLAG TITSGWTFGA GPALQIPFPM QMAYRFNGIG VTQNVLYENQ KLIANQFNSA IGKIQDSLSS T PSALGKLQ DVVNQNAQAL NTLVKQLSSN FGAISSVLND ILSRLDPPEA EVQIDRLITG RLQSLQTYVT QQLIRAAEIR AS ANLAATK MSECVLGQSK RVDFCGKGYH LMSFPQSAPH GVVFLHVTYV PAQEKNFTTA PAICHDGKAH FPREGVFVSN GTH WFVTQR NFYEPQIITT DNTFVSGNCD VVIGIVNNTV YDPLQPELDS FKEELDKYFK NHTSPDVDLG DISGINASVV NIQK EIDRL NEVAKNLNES LIDLQELGKY EQYIKGSGRE NLYFQGGGGS GYIPEAPRDG QAYVRKDGEW VLLSTFLGHH HHHHH H UniProtKB: Spike glycoprotein |
-Macromolecule #2: Variable Light Chain P5C3 (VL)
| Macromolecule | Name: Variable Light Chain P5C3 (VL) / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.388875 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EIVLTQSPGT LSLSPGERAT LSCRGSQSVR SSYLGWYQQK PGQAPRLLIY GASSRATGIP DRFSGSGSGT DFTLTISRLE PEDFAVYYC QQYGSSPWTF GQGTKVEIKG TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: EIVLTQSPGT LSLSPGERAT LSCRGSQSVR SSYLGWYQQK PGQAPRLLIY GASSRATGIP DRFSGSGSGT DFTLTISRLE PEDFAVYYC QQYGSSPWTF GQGTKVEIKG TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC |
-Macromolecule #3: Variable Heavy Chain P5C3 (VH)
| Macromolecule | Name: Variable Heavy Chain P5C3 (VH) / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.362133 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QMQLVQSGPE VKKPGTSVKV SCKASGFTFT SSAVQWVRQA RGQRLEWIGW IVVGSGNTDY AQQFQERVTI TRDMSTSTAY MELSSLGSE DTAVYYCAAP NCSGGSCYDG FDLWGQGTMV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA ...String: QMQLVQSGPE VKKPGTSVKV SCKASGFTFT SSAVQWVRQA RGQRLEWIGW IVVGSGNTDY AQQFQERVTI TRDMSTSTAY MELSSLGSE DTAVYYCAAP NCSGGSCYDG FDLWGQGTMV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA LTSGVHTFPA VLQSSGLYSL SSVVTVPSSS LGTQTYICNV NHKPSNTKVD KKV |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.3 / Component - Concentration: 150.0 mM / Component - Formula: PBS / Component - Name: PBS |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: OTHER / Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 12303 / Average electron dose: 38.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 2.5 µm / Calibrated defocus min: 0.8 µm / Calibrated magnification: 10500 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.27 mm / Nominal magnification: 10500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: HELIUM |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



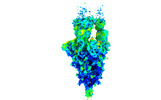










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)