+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13120 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mature capsid of bacteriophage phiRSA1 | |||||||||
 Map data Map data | Mature capsid of bacteriophage phiRSA1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | bacteriophage / capsid / VIRUS | |||||||||
| Function / homology | Bacteriophage P2, capsid / Phage major capsid protein, P2 family / P2 family phage major capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Ralstonia virus RSA1 / Ralstonia virus RSA1 /  Ralstonia solanacearum (bacteria) Ralstonia solanacearum (bacteria) | |||||||||
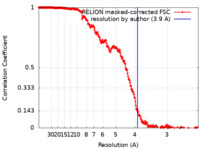
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Effantin G / Fujiwara A | |||||||||
 Citation Citation |  Journal: Int J Mol Sci / Year: 2021 Journal: Int J Mol Sci / Year: 2021Title: High Resolution Structure of the Mature Capsid of Bacteriophage ϕRSA1 by Cryo-Electron Microscopy. Authors: Grégory Effantin / Akiko Fujiwara / Takeru Kawasaki / Takashi Yamada / Guy Schoehn /   Abstract: The ϕRSA1 bacteriophage has been isolated from , a gram negative bacteria having a significant economic impact on many important crops. We solved the three-dimensional structure of the ϕRSA1 mature ...The ϕRSA1 bacteriophage has been isolated from , a gram negative bacteria having a significant economic impact on many important crops. We solved the three-dimensional structure of the ϕRSA1 mature capsid to 3.9 Å resolution by cryo-electron microscopy. The capsid shell, that contains the 39 kbp of dsDNA genome, has an icosahedral symmetry characterized by an unusual triangulation number of T = 7, . The ϕRSA1 capsid is composed solely of the polymerization of the major capsid protein, gp8, which exhibits the typical "Johnson" fold first characterized in bacteriophage HK97. As opposed to the latter, the ϕRSA1 mature capsid is not stabilized by covalent crosslinking between its subunits, nor by the addition of a decoration protein. We further describe the molecular interactions occurring between the subunits of the ϕRSA1 capsid and their relationships with the other known bacteriophages. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13120.map.gz emd_13120.map.gz | 1012.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13120-v30.xml emd-13120-v30.xml emd-13120.xml emd-13120.xml | 8.9 KB 8.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13120_fsc.xml emd_13120_fsc.xml | 23.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_13120.png emd_13120.png | 185.6 KB | ||
| Filedesc metadata |  emd-13120.cif.gz emd-13120.cif.gz | 4.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13120 http://ftp.pdbj.org/pub/emdb/structures/EMD-13120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13120 | HTTPS FTP |
-Related structure data
| Related structure data |  7oz4MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13120.map.gz / Format: CCP4 / Size: 1.1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13120.map.gz / Format: CCP4 / Size: 1.1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mature capsid of bacteriophage phiRSA1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.21 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ralstonia solanacearum
| Entire | Name:  Ralstonia solanacearum (bacteria) Ralstonia solanacearum (bacteria) |
|---|---|
| Components |
|
-Supramolecule #1: Ralstonia solanacearum
| Supramolecule | Name: Ralstonia solanacearum / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 305 / Sci species name: Ralstonia solanacearum / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Virus shell | Shell ID: 1 / Name: gp8 / Diameter: 640.0 Å / T number (triangulation number): 7 |
-Macromolecule #1: p2 family phage major capsid protein
| Macromolecule | Name: p2 family phage major capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ralstonia virus RSA1 Ralstonia virus RSA1 |
| Molecular weight | Theoretical: 37.831531 KDa |
| Sequence | String: MRNDTRRLFA AYKAAIAKLN GVERVDEKFS VAPSVQQKLE TKVQESSDFL KSINFYGVPE QEGEKIGLGV SGPVASTTDT TQQDRETSD ISTMDGRRYR CEQTNSDTHI TYQKLDAWAK FADFQTRIRD AIIKRQALDR IMIGFNGVSR AATSDRVANP M LQDVNKGW ...String: MRNDTRRLFA AYKAAIAKLN GVERVDEKFS VAPSVQQKLE TKVQESSDFL KSINFYGVPE QEGEKIGLGV SGPVASTTDT TQQDRETSD ISTMDGRRYR CEQTNSDTHI TYQKLDAWAK FADFQTRIRD AIIKRQALDR IMIGFNGVSR AATSDRVANP M LQDVNKGW LQNLREQAPQ RVMKEGKAAA GKITVGGAGA DYGNLDALVY DITNHLVEPW YAEDPDLVVV CGRNLLSDKY FP LVNRDRD PVQQIAADLI ISQKRIGNLP AIRVPYFPAN GLLVTRLDNL SIYYQEGGRR RTILDNAKRD RIENYESSND AYV IEDLAC AAMAENIALA AAA UniProtKB: P2 family phage major capsid protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)