[English] 日本語
 Yorodumi
Yorodumi- EMDB-1307: Electron cryotomography reveals the portal in the herpesvirus capsid. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1307 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
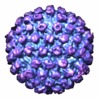
| Title | Electron cryotomography reveals the portal in the herpesvirus capsid. | |||||||||
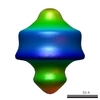
 Map data Map data | This is the 5-fold-averaged map. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Human herpesvirus 1 (Herpes simplex virus type 1) Human herpesvirus 1 (Herpes simplex virus type 1) | |||||||||
| Method | subtomogram averaging / cryo EM | |||||||||
 Authors Authors | Chang JT / Schmid MF / Rixon FJ / Chiu W | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2007 Journal: J Virol / Year: 2007Title: Electron cryotomography reveals the portal in the herpesvirus capsid. Authors: Juan T Chang / Michael F Schmid / Frazer J Rixon / Wah Chiu /  Abstract: Herpes simplex virus type 1 is a human pathogen responsible for a range of illnesses from cold sores to encephalitis. The icosahedral capsid has a portal at one fivefold vertex which, by analogy to ...Herpes simplex virus type 1 is a human pathogen responsible for a range of illnesses from cold sores to encephalitis. The icosahedral capsid has a portal at one fivefold vertex which, by analogy to portal-containing phages, is believed to mediate genome entry and exit. We used electron cryotomography to determine the structure of capsids lacking pentons. The portal vertex appears different from pentons, being located partially inside the capsid shell, a position equivalent to that of bacteriophage portals. Such similarity in portal organization supports the idea of the evolutionary relatedness of these viruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1307.map.gz emd_1307.map.gz | 52.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1307-v30.xml emd-1307-v30.xml emd-1307.xml emd-1307.xml | 8.9 KB 8.9 KB | Display Display |  EMDB header EMDB header |
| Images |  1307.gif 1307.gif | 97.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1307 http://ftp.pdbj.org/pub/emdb/structures/EMD-1307 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1307 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1307 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1307.map.gz / Format: CCP4 / Size: 89 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1307.map.gz / Format: CCP4 / Size: 89 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is the 5-fold-averaged map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.43 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Herpes Simplex Virus 1
| Entire | Name: Herpes Simplex Virus 1 |
|---|---|
| Components |
|
-Supramolecule #1000: Herpes Simplex Virus 1
| Supramolecule | Name: Herpes Simplex Virus 1 / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Human herpesvirus 1
| Supramolecule | Name: Human herpesvirus 1 / type: virus / ID: 1 / Name.synonym: Herpes Simplex / NCBI-ID: 10298 / Sci species name: Human herpesvirus 1 / Database: NCBI / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: Yes / Syn species name: Herpes Simplex |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: INVERTEBRATES Homo sapiens (human) / synonym: INVERTEBRATES |
| Virus shell | Shell ID: 1 / Name: Capsid / Diameter: 1250 Å / T number (triangulation number): 16 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 Details: 170mM NaCl, 3.4mM KCl, 10mM Na2HPO4, 1.8mM KH2PO4, 6.8mM CaCl2, 4.9mM MgCl2 |
|---|---|
| Grid | Details: 200 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: OTHER / Details: Vitrification instrument: Vitrobot |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010F |
|---|---|
| Temperature | Min: 93 K / Max: 93 K / Average: 93 K |
| Details | Tilted from -70 to +70 in 2 degree increments using Mr T software (unpublished). |
| Image recording | Category: CCD / Film or detector model: GENERIC GATAN / Average electron dose: 50 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 27680 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 3.0 µm / Nominal magnification: 20000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen cooled / Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: 70 ° / Tilt series - Axis1 - Max angle: 70 ° |
- Image processing
Image processing
| Details | Averaged 13 particles extracted from the reconstruction. Average number of tilts used in the 3D reconstructions: 71. Average tomographic tilt angle increment: 2. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Software - Name:  IMOD / Details: 5-fold-averaged IMOD / Details: 5-fold-averaged |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















